Abstract
The innate immune system is a powerful barrier against invading pathogens. Interferons (IFNs) are a major part of the cytokine-mediated anti-viral innate immune response. After recognition of a pathogen by immune sensors, signaling cascades are activated that culminate in the release of IFNs. These activate cells in an autocrine or paracrine fashion eventually setting cells in an anti-viral state via upregulation of hundreds of interferon-stimulated genes (ISGs). To evade the anti-viral effect of the IFN system, successful viruses like the pandemic severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) evolved strategies to counteract both IFN induction and signaling. In fact, more than half of the about 30 proteins encoded by SARS-CoV-2 target the IFN system at multiple levels to escape IFN-mediated restriction. Here, we review recent insights into the molecular mechanisms used by SARS-CoV-2 proteins to suppress IFN production and the establishment of an anti-viral state.
Similar content being viewed by others
Introduction
Invading viruses are detected by pattern recognition receptors (PRRs) like RIG-I-like receptors (RLRs) and Toll-like receptors (TLRs) recognizing viral pathogen-associated molecular patterns (PAMPs) [1]. For example, infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the pathogen that causes the current coronavirus disease 2019 (COVID-19) pandemic, is recognized by various PRRs, most prominently the RLR melanoma differentiation-associated protein 5 (MDA5) and TLRs, such as TLR2 [2], 3 [3] and 4 [4]. The exact nature of the SARS-CoV-2 PAMP(s) is currently unknown. However, it was suggested that the Spike protein may mediate TLR2 and TLR4 activation [2, 5, 6]. After recognition of a pathogen by PRRs, downstream signaling cascades are induced that ultimately lead to the activation of a kinase called Tank-binding kinase 1 (TBK1) that mediates phosphorylation of a set of transcription factors called interferon regulatory factors (IRFs), among them IRF3 and IRF7. Dimerization and translocation of IRFs to the nucleus eventually induce the expression and subsequent release of various (pro-)inflammatory cytokines, most prominently interferons (IFNs). There are three major types of IFNs, type I, type II, and type III, classified by their receptor usage. While type I and III IFNs are released by most nucleated cell types upon PRR activation, type II IFNs are mainly produced by activated T cells and natural killer cells [7]. Type I IFNs include primarily IFN-α (comprising 13 different subtypes) and IFN-β, but also more recently IFN-ε/κ, IFN-ω, and IFN-ν [8]. The only cytokine classified as type II IFN is IFN-γ. Three type III IFNs, IFN-λ1, IFN-λ2, and IFN-λ3, were formerly known as Interleukins (IL) IL29, IL28A, and IL28B with overlapping but distinct functions compared to type I IFNs [9]. Notably, receptors for type III IFNs are more specifically expressed on epithelial cells and some types of immune cells, such as dendritic cells and neutrophils, whereas type I and II IFN-receptors are present on almost all nucleated cells [10, 11]. Binding of IFNs in either a paracrine or autocrine fashion to their respective receptors results in the activation of signal transduction kinases, among them janus kinases (JAK) and tyrosine kinases (TYK) [12, 13]. These kinases in turn activate members of the signal transducer and activator of transcription (STAT) protein family, that drive the expression of hundreds of interferon-stimulated genes (ISGs) [14], many of which are known to restrict viral replication and the spread of viruses [12, 13, 15]. Thus, an anti-viral state in both virus-infected cells and uninfected bystander cells [8, 13, 16] is induced. In addition to activating the innate immune defenses, IFNs also play an important role in recruiting and activating cells of the adaptive immune system. However, successful viruses like SARS-CoV-2 have evolved strategies to evade or counteract the induction, signaling, and anti-viral effects of IFNs [17,18,19]. In this review, we provide a brief overview of how SARS-CoV-2 uses many of its proteins to counteract IFN induction and signaling, thus preventing effective innate immune activation.
Main text
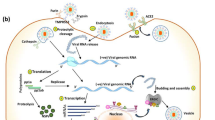
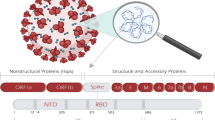
The 30 kb positive-sense single-strand RNA genome of SARS-CoV-2 encodes approximately 30 proteins [20]. Sixteen non-structural proteins (Nsp1-16) are produced by (auto-)proteolytic processing of two large precursor polyproteins open reading frame 1a (ORF1a) and ORF1ab, which are both translated from the full-length SARS-CoV-2 RNA. ORF1ab arises due to a ribosomal frameshift, allowing the translation to continue beyond the ORF1a stop codon. From subgenomic mRNAs, SARS-CoV-2 expresses four structural proteins, namely spike (S), envelop (E), membrane (M), and nucleocapsid (N), and several accessory factors. Among the accessory proteins, are ORF3a, ORF3b, ORF6, ORF7a, ORF7b, ORF8, ORF9b, and ORF10. Notably, some genes may encode for several proteins, such as the ORF3 locus which codes for ORF3a, b and possibly other products, like ORF3c [21]. Classically, viruses use their accessory proteins to counteract innate immune activation or effectors. However, recent reports have established that SARS-CoV-2 uses its non-structural, structural, as well as accessory proteins to counteract IFN induction and signaling [17,18,19].
Non-structural proteins
The 16 non-structural proteins encoded by SARS-CoV-2 are not part of the virion but are essential for viral replication and transcription. For example, they are involved in the formation of replication complexes and play a prominent role in IFN escape, with several directly targeting key players of the IFN signaling cascade (Table 1, Fig. 1). All Nsps, except Nsp2, 4, 7–11, and 16, were reported to diminish IFN induction and/or signaling [17, 22, 23]. Notably, Nsps often have enzymatic function, such as the two main proteases of SARS-CoV-2 Nsp3 and Nsp5.
Counteraction of the IFN system by SARS-CoV-2 proteins. Schematic depiction of the antagonism of the interferon (IFN) system by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) proteins. Incoming or replicating virus is recognized by Toll-like receptors (TLRs) or RIG-I-like receptors (RLRs) which eventually activates interferon regulatory factor 3 (IRF3) either through TANK binding kinase protein 1 (TBK1) or via mitochondrial antiviral-signaling protein (MAVS). Activated IRF3 dimerizes and translocates to the nucleus, where it induces the production of IFNs. IFNs bind to their respective receptors (e.g., interferon alpha and beta receptor subunit 1, IFNAR) to induce janus kinase (JAK) and tyrosine kinase (TYK) mediated activation of signal transducer and activator of transcription (STATs). Activated STAT complexes (ISGF3) translocate to the nucleus, where they induce transcription of interferon-stimulated genes (ISGs). Induction of ISGs sets the cell in an antiviral state that restricts infection and replication of the virus. SARS-CoV-2 interferes with signal transduction at multiple levels, as indicated by red highlights. Nsp non-structural protein, N nucleocapsid protein, M matrix protein, ORF open reading frame
Nsp1 is a 180-amino acid protein, consisting of a globular domain and a C-terminal helix–turn–helix motif. It binds to the cellular ribosome plugging the mRNA entry channel with its helix–turn–helix motif thereby preventing translation of mRNAs [24, 25]. Consequently, Nsp1 drastically reduces the expression of all types of IFNs and ISGs [24]. Notably, SARS-CoV-2 replicon systems lacking functional Nsp1 are more susceptible toward type I IFNs [26].
With a size of more than 200 kDa, Nsp3 is the largest protein among the Nsps. It is a key component of the viral replication and transcription complex that assembles on host-cell membranes. Nsp3 has papain-like protease (PLpro) activity, which is required for Nsp1-4 processing, and in addition functions as a deubiquitinase and deISGylase [27, 28]. Recent studies have shown that Nsp3 antagonizes ISGylation of MDA5 thereby blocking the activation of this PRR [17, 29]. In addition, Nsp3 downregulates signal transduction by deISGylating the transcription factor IRF3 [30].
The main protease of SARS-CoV-2, Nsp5 was reported to suppress induction of type I IFN expression and signaling induced by all types of IFNs [17, 31]. Mechanistically, it has been suggested that Nsp5 cleaves retinoic acid-inducible gene I (RIG-I) rendering it inactive and promotes proteasome-mediated destruction of the signaling adaptor mitochondrial anti-viral-signaling protein (MAVS) [32]. Thus, inhibition of Nsp5 protease activity could at least partially alleviate its impact on type I IFN induction.
Nsp6 (together with Nsp3 and 4) is involved in the formation of the SARS-CoV-2 replication compartment made up of double-membrane vesicles. In addition, it was reported that Nsp6 interacts with TBK1 to inhibit activation of IRF3, in turn restricting type I IFN induction [18].
The active subunit of the SARS-CoV-2 RNA-dependent RNA polymerase Nsp12 has been suggested to suppress type I IFN induction by inhibiting nuclear translocation of the transcription factor IRF3 [33]. However, several studies by other labs did not confirm this, as Nsp12 was not picked up in various screening approaches or did explicitly not affect endogenous type I IFN induction [17, 18, 34].
The SARS-CoV-2 encoded helicase Nsp13 inhibits type I IFN production and signaling by preventing the activation of STAT1 and STAT2, which are crucial for IFN signaling [18, 19, 35]. Furthermore, immunoprecipitation experiments combined with proteomic approaches suggest that Nsp13 may bind to TBK1 to inhibit its phosphorylation and thus activation of IRF3 [18, 22, 36].
Nsp14 is a guanine-N7-methyltransferase required for efficient SARS-CoV-2 transcription. It facilitates the formation of the mRNA cap, thus preventing the recognition of SARS-CoV-2 mRNAs by PRRs [37]. In addition, Nsp14 significantly reduced IFN-β promoter-driven luciferase activities and induced degradation of endogenous interferon alpha and beta receptor subunit 1 (IFNAR1) at the protein level, thus inhibiting type I IFN binding to cells and subsequent signaling [17, 19, 35].
Nsp15 is an uridine-specific endoribonuclease [38], which was suggested to remove and/or process viral RNA that would otherwise trigger detection of the virus and thus IFN induction. Furthermore, it was reported to reduce type I IFN production by interaction with ring finger protein 41 (RNF41), an E3 ligase associated with the activation of IRF3 [22].
While many molecular mechanisms of SARS-CoV-2 Nsps have been identified, future studies are needed to evaluate the various roles of many Nsps in viral replication and IFN antagonism. Furthermore, many Nsps function in complexes, which were not analyzed in the context of IFN induction/signaling antagonism yet. Importantly, some Nsps require enzymatic function to antagonize the IFN system and may be targeted, e.g., by small molecules. For example, orally available inhibitors of Nsp5 are already examined in clinical trials [39, 40]. These inhibitors may not only restrict SARS-CoV-2 via prevention of the cleavage of its protein products but also by increasing anti-viral immune responses.
Structural proteins
SARS-CoV-2 encodes four structural proteins on subgenomic mRNAs that are part of the assembled and infectious virion: S, N, M, and E. Recent studies reported that even the structural proteins of SARS-CoV-2 may play a role in antagonizing IFN production and signaling (Table 1). It has been shown that the N protein inhibits both type I IFN production and signaling by targeting RLRs [41] and decreasing the phosphorylation of STAT1 and STAT2 [39], respectively. In addition, the N protein wraps the genomic RNA, shielding it from recognition by PRRs. The M protein was found to suppress type I IFN induction upon Sendai virus (SeV) stimulation, or stimulation via expression of MDA5 or RIG-I [17, 19].
In addition to their structural functions as part of the virion, N and M play active roles in inhibiting the type I IFN system. Thus, future studies could investigate whether incoming virions may be sufficient to reduce type I IFN induction and signaling immediately after infection.
Accessory proteins
Besides non-structural proteins and structural proteins coronaviruses such as SARS-CoV-2 encode for accessory proteins. Although the roles of these proteins are not fully understood, it is known that they often play crucial roles in virus–host interactions, specifically with the innate immune system (Table 1).
The ORF3 locus encodes multiple proteins, ORF3a being the longest with smaller products, like ORF3b and ORF3c, produced from downstream start codons. ORF3a was suggested to interfere with proper activation of JAKs by inducing Suppressor of Cytokine Signalling 1 (SOCS1), thus attenuating IFN signaling [43]. ORF3b was reported to inhibit IFN production through its C-terminus [44]. Interestingly, some naturally occurring SARS-CoV-2 variants encode a longer version of ORF3b with increased activity, while other strains express a truncated and presumably inactive variant of this protein [45].
Despite its small size of only 7 kDa, ORF6 was consistently identified as a very potent inhibitor of type I and/or III IFN induction and signaling [17,18,19, 35]. Mechanistically, it was reported that ORF6 binds to nucleoporin 98 (Nup98) dislocating it from the nuclear pore [46], thus preventing efficient nuclear import of STATs [47] and IRF3 [19, 48], in turn inhibiting both type I IFN induction and signaling.
Two proteins are expressed from the ORF7 locus: ORF7a and ORF7b. While ORF7a is a 121 aa-long transmembrane protein, ORF7b encompasses only 43 aa. Recent evidence indicates that both ORF7a and ORF7b block STAT2 phosphorylation, thereby inhibiting type I IFN signaling [18]. Curiously, the modification of ORF7a by covalently conjugated ubiquitin promotes its function as an antagonist of type I IFN responses [49].
ORF9b is a small 97 aa protein expressed from an alternative ORF in the N locus. It was suggested that it suppresses type I IFN induction by targeting the MAVS signalosome [50].
Overall, the accessory proteins clearly play a major role in antagonizing activation of the IFN system. However, it seems that antagonism of innate immunity is only a part of their function, and many of them have additional roles, e.g., in virion assembly and egress of the virus.
Concluding remarks
IFNs are a key component of the innate immune system that set cells in an anti-viral state by inducing hundreds of (anti-viral) ISGs. Evidently, the unfortunate success of SARS-CoV-2 is critically dependent on its ability to counteract both IFN induction and signaling to evade the host’s innate defenses. Since the discovery of SARS-CoV-2 in late 2019, our knowledge about this pathogen has literally exploded and is still rapidly expanding. However, despite much progress in an amazingly short time, many questions on IFN antagonists encoded by SARS-CoV-2 still remain.
While the extend of SARS-CoV-2 proteins that inhibit IFN induction and signaling is established (Fig. 1, Table 1), the respective underlying molecular mechanism(s) often remain elusive [17,18,19]. Of note, most viral proteins were studied in the context of type I interferon responses. A recent study suggests that most type I IFN signaling antagonists of SARS-CoV-2 may also affect type II and III signaling, albeit to varying extend [17]. For many viral proteins, however, it remains to be determined whether they also target type II or III IFN induction and/or responses. In addition, the relevance and contribution of the individual proteins to the immune escape of SARS-CoV-2, as well as functional interactions and synergisms, are currently largely unclear. In a replicon setting, a mutant lacking Nsp1 was more sensitive toward type I IFN-mediated inhibition [26]. Furthermore, recombinant SARS-CoV-2 lacking ORF6 induces higher levels of ISGs in vitro [23]. However, more studies using recombinant viruses and in vivo models are required to better understand the individual contribution of IFN counteraction mechanisms to viral spread, replication, and pathogenesis.
SARS-CoV-2 targets IFN induction and signaling cascades at multiple levels using more than half of its proteins [17,18,19] (Fig. 1, Table 1). This highlights how crucial it is for successful viruses to tightly control these signaling pathways. None of the individual SARS-CoV-2 proteins inhibits the IFN system entirely. Thus, multiple factors need to synergize to allow efficient viral immune evasion and spread. Especially, IFN production and release need to be kept at a minimum by the virus to avoid setting uninfected cells in an anti-viral state and recruiting/activating the adaptive immune responses.
It was reported that SARS-CoV-1 is more resistant to inhibition by type I IFNs [51]. In vitro evidence suggested that Nsp15 of SARS-CoV-1 is a stronger type I IFN signaling antagonist than Nsp15 of SARS-CoV-2, possibly providing one explanation for the difference between the two coronaviruses [17]. How SARS-CoV-2 compares to middle east respiratory syndrome coronavirus (MERS) or seasonal coronaviruses in terms of IFN resistance is currently unknown.
SARS-CoV-2 continues to adapt for an efficient spread in the human population resulting in the emergence of variants [52, 53]. It was reported that variants of ORF3b that are either longer and shorter forms are either more or less active than the original ORF3b variant, respectively [44]. Variants of Nsp1 were detected, which show increased efficiencies in IFN antagonism [54]. It was noted, that variants of concern (VOC) differ in their resistance toward exogenous IFN, with the alpha VOC being consistently the most resistant [55, 56]. Mechanistic analysis revealed that the alpha VOC has increased relative expression levels of the IFN antagonists N, ORF9b, and ORF6 compared to an early 2020 SARS-CoV-2 strain [57].
As SARS-CoV-2 was shown to be sensitive to innate immune activation, treatment with IFNs may be beneficial in COVID-19 [58, 59]. Accordingly, the early presence of IFNs was shown to protect COVID-19 patients from severe disease [51, 60]. However, IFNs are detrimental in the long run by promoting inflammation. The excessive presence of IFNs (and other pro-inflammatory cytokines) often defines the severity of the diseases and possibly even long-term consequences of the infection [61, 62]. Consequently, anti-viral IFN therapy is usually efficient but also associated with severe side effects. Therefore, exact timing and dosing are paramount, e.g., using the most potent IFNs or synergies in combinatorial approaches with multiple different IFNs [17]. In addition, a better understanding of the mechanistic details of the IFN antagonism by SARS-CoV-2 proteins may allow us to more efficiently interfere with and perhaps even prevent viral immune evasion.
In summary, while the past more than two years have brought astounding progress in the characterization of the interplay between the IFN system and SARS-CoV-2, we are still only beginning to understand the intricate details. Many of the proteins described here also antagonize other pathways of anti-viral immunity, besides the IFN system, such as autophagy [17, 63]. Especially for therapeutic intervention, e.g., with IFNs, we need to better understand the interplay between the IFN system and SARS-CoV-2 to define the best dose, timing, and synergistic combinations of different IFNs to approach safe and effective COVID-19 therapy based on innate immune modulation [64].
References
Janeway CA, Medzhitov R (2002) Innate immune recognition. Annu Rev Immunol 20:197–216. https://doi.org/10.1146/annurev.immunol.20.083001.084359
Zheng M, Karki R, Williams EP et al (2021) TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nat Immunol 22:829–838. https://doi.org/10.1038/s41590-021-00937-x
Tripathi U, Nchioua R, Prata LGPL et al (2021) SARS-CoV-2 causes senescence in human cells and exacerbates the senescence-associated secretory phenotype through TLR-3. Aging 13:21838–21854. https://doi.org/10.18632/aging.203560
Kell AM, Gale M (2015) RIG-I in RNA virus recognition. Virology 479–480:110–121. https://doi.org/10.1016/j.virol.2015.02.017
Aboudounya MM, Holt MR, Heads RJ (2021) SARS-CoV-2 Spike S1 glycoprotein is a TLR4 agonist, upregulates ACE2 expression and induces pro-inflammatory M1 macrophage polarisation. 2021.08.11.455921
Zhao Y, Kuang M, Li J et al (2021) SARS-CoV-2 spike protein interacts with and activates TLR41. Cell Res 31:818–820. https://doi.org/10.1038/s41422-021-00495-9
Polyfunctional responses by human T cells result from sequential release of cytokines | PNAS. https://www.pnas.org/content/109/5/1607.short. Accessed 1 Feb 2022
Platanias LC (2005) Mechanisms of type-I- and type-II-interferon-mediated signalling. Nat Rev Immunol 5:375–386. https://doi.org/10.1038/nri1604
Stanifer ML, Guo C, Doldan P, Boulant S (2020) Importance of type I and III interferons at respiratory and intestinal barrier surfaces. Front Immunol 11:608645. https://doi.org/10.3389/fimmu.2020.608645
Broggi A, Tan Y, Granucci F, Zanoni I (2017) IFN-λ suppresses intestinal inflammation by non-translational regulation of neutrophil function. Nat Immunol 18:1084–1093. https://doi.org/10.1038/ni.3821
Hemann EA, Green R, Turnbull JB et al (2019) Interferon-λ modulates dendritic cells to facilitate T cell immunity during infection with influenza A virus. Nat Immunol 20:1035–1045. https://doi.org/10.1038/s41590-019-0408-z
Koepke L, Gack MU, Sparrer KM (2021) The antiviral activities of TRIM proteins. Curr Opin Microbiol 59:50–57. https://doi.org/10.1016/j.mib.2020.07.005
Sparrer KM, Gack MU (2015) Intracellular detection of viral nucleic acids. Curr Opin Microbiol 26:1–9. https://doi.org/10.1016/j.mib.2015.03.001
Schneider WM, Chevillotte MD, Rice CM (2014) Interferon-stimulated genes: a complex web of host defenses. Annu Rev Immunol 32:513–545. https://doi.org/10.1146/annurev-immunol-032713-120231
Schoggins JW (2019) Interferon-stimulated genes: what do they all do? Annu Rev Virol 6:567–584. https://doi.org/10.1146/annurev-virology-092818-015756
Lee J-H, Chiang C, Gack MU (2019) Endogenous nucleic acid recognition by RIG-I-like receptors and cGAS. J Interferon Cytokine Res 39:450–458. https://doi.org/10.1089/jir.2019.0015
Hayn M, Hirschenberger M, Koepke L et al (2021) Systematic functional analysis of SARS-CoV-2 proteins uncovers viral innate immune antagonists and remaining vulnerabilities. Cell Rep 35:109126. https://doi.org/10.1016/j.celrep.2021.109126
Xia H, Cao Z, Xie X et al (2020) Evasion of type I interferon by SARS-CoV-2. Cell Rep 33:108234. https://doi.org/10.1016/j.celrep.2020.108234
Lei X, Dong X, Ma R et al (2020) Activation and evasion of type I interferon responses by SARS-CoV-2. Nat Commun 11:3810. https://doi.org/10.1038/s41467-020-17665-9
V’kovski P, Kratzel A, Steiner S, et al (2021) Coronavirus biology and replication: implications for SARS-CoV-2. Nat Rev Microbiol 19:155–170. https://doi.org/10.1038/s41579-020-00468-6
Jungreis I, Nelson CW, Ardern Z et al (2021) Conflicting and ambiguous names of overlapping ORFs in the SARS-CoV-2 genome: a homology-based resolution. Virology 558:145–151. https://doi.org/10.1016/j.virol.2021.02.013
Gordon DE, Jang GM, Bouhaddou M et al (2020) A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583:459–468. https://doi.org/10.1038/s41586-020-2286-9
Schroeder S, Pott F, Niemeyer D et al (2021) Interferon antagonism by SARS-CoV-2: a functional study using reverse genetics. Lancet Microbe 2:e210–e218. https://doi.org/10.1016/S2666-5247(21)00027-6
Thoms M, Buschauer R, Ameismeier M et al (2020) Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Science 369:1249–1255. https://doi.org/10.1126/science.abc8665
Schubert K, Karousis ED, Jomaa A et al (2020) SARS-CoV-2 Nsp1 binds the ribosomal mRNA channel to inhibit translation. Nat Struct Mol Biol 27:959–966. https://doi.org/10.1038/s41594-020-0511-8
Ricardo-Lax I, Luna JM, Thao TTN et al (2021) Replication and single-cycle delivery of SARS-CoV-2 replicons. Science 374:1099–1106. https://doi.org/10.1126/science.abj8430
Clementz MA, Chen Z, Banach BS et al (2010) Deubiquitinating and interferon antagonism activities of coronavirus papain-like proteases. J Virol 84:4619–4629. https://doi.org/10.1128/JVI.02406-09
Harcourt BH, Jukneliene D, Kanjanahaluethai A et al (2004) Identification of severe acute respiratory syndrome coronavirus replicase products and characterization of papain-like protease activity. J Virol 78:13600–13612. https://doi.org/10.1128/JVI.78.24.13600-13612.2004
Liu G, Lee J-H, Parker ZM et al (2021) ISG15-dependent activation of the sensor MDA5 is antagonized by the SARS-CoV-2 papain-like protease to evade host innate immunity. Nat Microbiol 6:467–478. https://doi.org/10.1038/s41564-021-00884-1
Shin D, Mukherjee R, Grewe D et al (2020) Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587:657–662. https://doi.org/10.1038/s41586-020-2601-5
Wu Y, Ma L, Zhuang Z et al (2020) Main protease of SARS-CoV-2 serves as a bifunctional molecule in restricting type I interferon antiviral signaling. Signal Transduct Target Ther 5:221. https://doi.org/10.1038/s41392-020-00332-2
Liu Y, Qin C, Rao Y et al (2021) SARS-CoV-2 Nsp5 Demonstrates Two Distinct Mechanisms Targeting RIG-I and MAVS To Evade the Innate Immune Response. mBio 12:e0233521. https://doi.org/10.1128/mBio.02335-21
Wang W, Zhou Z, Xiao X et al (2021) SARS-CoV-2 nsp12 attenuates type I interferon production by inhibiting IRF3 nuclear translocation. Cell Mol Immunol 18:945–953. https://doi.org/10.1038/s41423-020-00619-y
Li A, Zhao K, Zhang B et al (2021) SARS-CoV-2 NSP12 protein is not an interferon-β antagonist. J Virol 95:e0074721. https://doi.org/10.1128/JVI.00747-21
Yuen C-K, Lam J-Y, Wong W-M et al (2020) SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 function as potent interferon antagonists. Emerg Microbes Infect 9:1418–1428. https://doi.org/10.1080/22221751.2020.1780953
Hoffmann H-H, Sánchez-Rivera FJ, Schneider WM et al (2021) Functional interrogation of a SARS-CoV-2 host protein interactome identifies unique and shared coronavirus host factors. Cell Host Microbe 29:267-280.e5. https://doi.org/10.1016/j.chom.2020.12.009
N7-Methylation of the Coronavirus RNA Cap Is Required for Maximal Virulence by Preventing Innate Immune Recognition | mBio. https://journals.asm.org/doi/https://doi.org/10.1128/mbio.03662-21. Accessed 1 Feb 2022
Pillon MC, Frazier MN, Dillard LB et al (2021) Cryo-EM structures of the SARS-CoV-2 endoribonuclease Nsp15 reveal insight into nuclease specificity and dynamics. Nat Commun 12:636. https://doi.org/10.1038/s41467-020-20608-z
Qiao J, Li Y-S, Zeng R et al (2021) SARS-CoV-2 Mpro inhibitors with antiviral activity in a transgenic mouse model. Science. https://doi.org/10.1126/science.abf1611
Owen DR, Allerton CMN, Anderson AS et al (2021) An oral SARS-CoV-2 Mpro inhibitor clinical candidate for the treatment of COVID-19. Science 374:1586–1593. https://doi.org/10.1126/science.abl4784
Oh SJ, Shin OS (2021) SARS-CoV-2 nucleocapsid protein targets RIG-I-like receptor pathways to inhibit the induction of interferon response. Cells 10:530. https://doi.org/10.3390/cells10030530
Mu J, Fang Y, Yang Q et al (2020) SARS-CoV-2 N protein antagonizes type I interferon signaling by suppressing phosphorylation and nuclear translocation of STAT1 and STAT2. Cell Discov 6:1–4. https://doi.org/10.1038/s41421-020-00208-3
Wang R, Yang X, Chang M et al (2021) ORF3a protein of severe acute respiratory syndrome coronavirus 2 inhibits interferon-activated janus kinase/signal transducer and activator of transcription signaling via elevating suppressor of cytokine signaling 1. Front Microbiol 12:752597. https://doi.org/10.3389/fmicb.2021.752597
Konno Y, Kimura I, Uriu K et al (2020) SARS-CoV-2 ORF3b is a potent interferon antagonist whose activity is increased by a naturally occurring elongation variant. Cell Rep 32:108185. https://doi.org/10.1016/j.celrep.2020.108185
Lam J-Y, Yuen C-K, Ip JD et al (2020) Loss of orf3b in the circulating SARS-CoV-2 strains. Emerg Microbes Infect 9:2685–2696. https://doi.org/10.1080/22221751.2020.1852892
Kato K, Ikliptikawati DK, Kobayashi A et al (2021) Overexpression of SARS-CoV-2 protein ORF6 dislocates RAE1 and NUP98 from the nuclear pore complex. Biochem Biophys Res Commun 536:59–66. https://doi.org/10.1016/j.bbrc.2020.11.115
Miorin L, Kehrer T, Sanchez-Aparicio MT et al (2020) SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc Natl Acad Sci U S A 117:28344–28354. https://doi.org/10.1073/pnas.2016650117
Kimura I, Konno Y, Uriu K et al (2021) Sarbecovirus ORF6 proteins hamper induction of interferon signaling. Cell Rep 34:108916. https://doi.org/10.1016/j.celrep.2021.108916
Cao Z, Xia H, Rajsbaum R et al (2021) Ubiquitination of SARS-CoV-2 ORF7a promotes antagonism of interferon response. Cell Mol Immunol 18:746–748. https://doi.org/10.1038/s41423-020-00603-6
Jiang H-W, Zhang H-N, Meng Q-F et al (2020) SARS-CoV-2 Orf9b suppresses type I interferon responses by targeting TOM70. Cell Mol Immunol 17:998–1000. https://doi.org/10.1038/s41423-020-0514-8
Lokugamage KG, Hage A, de Vries M et al (2020) Type I interferon susceptibility distinguishes SARS-CoV-2 from SARS-CoV. J Virol. https://doi.org/10.1128/JVI.01410-20
Jung C, Kmiec D, Koepke L et al (2022) Omicron: what makes the latest SARS-CoV-2 variant of concern so concerning? J Virol jvi0207721. https://doi.org/10.1128/jvi.02077-21
Harvey WT, Carabelli AM, Jackson B et al (2021) SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol 19:409–424. https://doi.org/10.1038/s41579-021-00573-0
Lin J-W, Tang C, Wei H-C et al (2021) Genomic monitoring of SARS-CoV-2 uncovers an Nsp1 deletion variant that modulates type I interferon response. Cell Host Microbe 29:489-502.e8. https://doi.org/10.1016/j.chom.2021.01.015
Guo K, Barrett BS, Mickens KL et al (2021) Interferon resistance of emerging SARS-CoV-2 variants. BioRxiv Prepr Serv Biol 2021.03.20.436257. https://doi.org/10.1101/2021.03.20.436257
Nchioua R, Schundner A, Klute S et al (2021) The Delta variant of SARS-CoV-2 maintains high sensitivity to interferons in human lung cells. Microbiology
Thorne LG, Bouhaddou M, Reuschl A-K et al (2021) Evolution of enhanced innate immune evasion by SARS-CoV-2. Nature. https://doi.org/10.1038/s41586-021-04352-y
Blanco-Melo D, Nilsson-Payant BE, Liu W-C et al (2020) Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181:1036-1045.e9. https://doi.org/10.1016/j.cell.2020.04.026
Hadjadj J, Yatim N, Barnabei L et al (2020) Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 369:718–724. https://doi.org/10.1126/science.abc6027
Sa Ribero M, Jouvenet N, Dreux M, Nisole S (2020) Interplay between SARS-CoV-2 and the type I interferon response. PLOS Pathog 16:e1008737. https://doi.org/10.1371/journal.ppat.1008737
Lee AJ, Ashkar AA (2018) The dual nature of Type I and Type II Interferons. Front Immunol 9:2061. https://doi.org/10.3389/fimmu.2018.02061
Hirschenberger M, Hunszinger V, Sparrer KMJ (2021) Implications of innate immunity in post-acute sequelae of non-persistent viral infections. Cells 10:2134. https://doi.org/10.3390/cells10082134
Koepke L, Hirschenberger M, Hayn M et al (2021) Manipulation of autophagy by SARS-CoV-2 proteins. Autophagy. https://doi.org/10.1080/15548627.2021.1953847
Zanoni I (2021) Interfering with SARS-CoV-2: are interferons friends or foes in COVID-19? Curr Opin Virol 50:119–127. https://doi.org/10.1016/j.coviro.2021.08.004
Acknowledgements
We apologize to the authors whose work on IFN and SARS-CoV-2 could not be cited or mentioned in this short review.
Funding
Open Access funding enabled and organized by Projekt DEAL. Work in the Sparrer lab is funded by the German Ministry for Research and Education (BMBF; Project IMMUNOMOD) and the German Research Foundation (DFG; SPP 1923, CRC 1279 and SP 1600/6–1). F.K. is supported by the DFG (SPP 1923, CRC 1279) and BMBF (Restrict SARS-CoV-2). L.K. is part of the International Graduate School of Molecular Medicine, Ulm (IGradU).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interests.
Additional information
Edited by Hanna-Mari Baldauf.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Special Issue on Immunobiology of Viral Infections.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, JH., Koepke, L., Kirchhoff, F. et al. Interferon antagonists encoded by SARS-CoV-2 at a glance. Med Microbiol Immunol 212, 125–131 (2023). https://doi.org/10.1007/s00430-022-00734-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-022-00734-9