Abstract
Cell fate decision processes are regulated by networks which contain different molecules and interactions. Different network topologies may exhibit synergistic or antagonistic effects on cellular functions. Here, we analyze six most common small networks with regulatory logic AND or OR, trying to clarify the relationship between network topologies and synergism (or antagonism) related to cell fate decisions. We systematically examine the contribution of both network topologies and regulatory logic to the cell fate synergism by bifurcation and combinatorial perturbation analysis. Initially, under a single set of parameters, the synergism of three types of networks with AND and OR logic is compared. Furthermore, to consider whether these results depend on the choices of parameter values, statistics on the synergism of five hundred parameter sets is performed. It is shown that the results are not sensitive to parameter variations, indicating that the synergy or antagonism mainly depends on the network topologies rather than the choices of parameter values. The results indicate that the topology with “Dual Inhibition” shows good synergism, while the topology with “Dual Promotion” or “Hybrid” shows antagonism. The results presented here may help us to design synergistic networks based on network structure and regulation combinations, which has promising implications for cell fate decisions and drug combinations.











Similar content being viewed by others
Data availability
No public data are used. The data we used are generated by randomly chosen values of model parameters in certain ranges.
References
Ralf, J.: Cell fate reprogramming through engineering of native transcription factors. Curr. Opin. Genet. Dev. 52, 109–116 (2018)
Jukam, D., Desplan, C.: Binary fate decisions in differentiating neurons. Curr. Opin. Neurobiol. 29(1), 6–13 (2010)
Chen, X., Hartman, A., Guo, S.: Choosing cell fate through a dynamic cell cycle. Curr. Stem Cell Rep. 1(3), 129–138 (2015)
Yu, X.X., Qiu, W.L., Yang, L., Zhang, Y., He, M.Y., Li, L.C., Xu, C.R.: Defining multistep cell fate decision pathways during pancreatic development at single-cell resolution. EMBO J. 38(8), (2019)
Evan, D., Danielle, T., Christopher, J.: Cell fate decision making through oriented cell division. J. Dev. Biol. 3(4), 129–157 (2015)
Ding, X.M.: Micrornas: regulators of cancer metastasis and epithelial-mesenchymal transition (emt). Chin. J. Cancer 33(3), 140 (2014)
Nieto, M.A.: The ins and outs of the epithelial to mesenchymal transition in health and disease. Annu. Rev. Cell Dev. Biol. 27, 347–376 (2011)
Fischer, N.W., Prodeus, A., Malkin, D., Gariépy, J.: p53 oligomerization status modulates cell fate decisions between growth, arrest and apoptosis. Cell Cycle 15(23), 3210–3219 (2016)
Bieging, K.T., Mello, S.S., Attardi, L.D.: Unravelling mechanisms of p53-mediated tumour suppression. Nat. Rev. Cancer 14(5), 359–370 (2014)
Hat, B., Kochańczyk, M., Bogdał, M.N., Lipniacki, T.: Feedbacks, bifurcations, and cell fate decision-making in the p53 system. Plos Comput. Biol. 12(2), e1004787 (2016)
Tatapudy, S., Aloisio, F., Barber, D., Nystul, T.: Cell fate decisions: emerging roles for metabolic signals and cell morphology. EMBO Rep. 18(12), 2105–2118 (2017)
Frey, U., Kottenberg, E., Kamler, M., Leineweber, K., Manthey, I., Heusch, G., Siffert, W., Peters, J.: Genetic interactions in the β-adrenoceptor g-protein signal transduction pathway and survival after coronary artery bypass grafting: A pilot study. Br. J. Anaesth. 107(6), 869–878 (2011)
Casey, M.J., Stumpf, P.S., MacArthur, B.D.: Theory of cell fate. Wiley Interdiscip. Rev. Syst. Biol. 12(2), (2020)
Maeda, M., Lu, S., Shaulsky, G., Miyazaki, Y., Kuwayama, H., Tanaka, Y., Kuspa, A., Loomis, W.: Periodic signaling controlled by an oscillatory circuit that includes protein kinases erk2 and pka. Science 304(5672), 875–878 (2004)
Zhang, Y., Liu, H., Yan, F., Zhou, J.: Oscillatory dynamics of p38 activity with transcriptional and translational time delays. Sci. Rep. 7(1), 11495 (2017)
Zeng, Y.T., Liu, X.F., Yang, W.T., Zheng, P.S.: Rex1 promotes emt-induced cell metastasis by activating the jak2/stat3-signaling pathway by targeting socs1 in cervical cancer. Oncogene 38(43), 1–18 (2019)
Pfeuty, B., Kaneko, K.: The combination of positive and negative feedback loops confers exquisite flexibility to biochemical switches. Phys. Biol. 6(4), (2009)
Yu, P., Nie, Q., Tang, C., Zhang, L.: Nanog induced intermediate state in regulating stem cell differentiation and reprogramming. Bmc Syst. Biol. 12(1), 22 (2018)
Zheng, X., Jin, S., Nie, Q., Zou, X.: scrcmf: Identification of cell subpopulations and transition states from single-cell transcriptomes. IEEE Trans. Biomed. Eng. 67(5), 1418–1428 (2020)
Collens, J., Pusuluri, K., Kelley, A., Knapper, D.E., Shilnikov, A.: Dynamics and bifurcations in multistable 3-cell neural networks. Chaos 30(7), 072101(2020)
Abdelaziz, A., Sarma, A.K.: Effective control and switching of optical multistability in a three-level v-type atomic system. Phys. Rev. A 102(4),(2020)
Liu, Y., Li, S., Liu, Z., Wang, R.: Bifurcation-based approach reveals synergism and optimal combinatorial perturbation. J. Biol. Phys. 42(3), 399–414 (2016)
Luo, M., Huang, D., Jiao, J., Wang, R.: Detection of synergistic combinatorial perturbations by a bifurcation-based approach. Int. J. Bifurcat. Chaos 31(12), 2150175 (2021)
Chou, T.C.: Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 58(3), 621–681 (2006)
Chou, T.C.: Preclinical versus clinical drug combination studies. Leuk. Lymphoma 49(11), 2059–2080 (2008)
Chou, T.C.: Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70(2), 440–6 (2010)
Fitzgerald, J.B., Schoeberl, B., Nielsen, U.B., Sorger, P.K.: Systems biology and combination therapy in the quest for clinical efficacy. Nature Chemical Biol. 2(9), 458 (2006)
Zhang, J., Yuan, Z., Zhou, T.: Synchronization and clustering of synthetic genetic networks: A role for cis-regulatory modules. Phys. Rev. E 79, (2009)
Goh, K.I., Kahng, B., Cho, K.H.: Sustained oscillations in extended genetic oscillatory systems. Biophys. J. 94(11), 4270–4276 (2008)
Deritei, D., Rozum, J., Regan, E.R., Albert, R.: A feedback loop of conditionally stable circuits drives the cell cycle from checkpoint to checkpoint. Sci. Rep. 9(1), 1–19 (2019)
Mitrophanov, A.Y., Groisman, E.A.: Positive feedback in cellular control systems. BioEssays 30(6), 542–555 (2008)
Díaz-López, A., Moreno-Bueno, G., Cano, A.: Role of microrna in epithelial to mesenchymal transition and metastasis and clinical perspectives. Cancer Manag. Res. 6, 205–216 (2014)
Huang, B., Lu, M., Galbraith, M., Levine, H., Onuchic, J.N., Jia, D.: Decoding the mechanisms underlying cell-fate decision-making during stem cell differentiation by random circuit perturbation. J. R. Soc. Interface 17(169), 20200500 (2020)
Gelens, L., Anderson, G.A., Ferrell, J.E.: Spatial trigger waves: positive feedback gets you a long way. Mol. Biol. Cell 25(22), 3486–3493 (2014)
Katebi, A., Kohar, V., Lu, M.: Random parametric perturbations of gene regulatory circuit uncover state transitions in cell cycle. iScience 23(6), 101150 (2020)
Inoue, D., Sagata, N.: The polo-like kinase plx1 interacts with and inhibits myt1 after fertilization of Xenopus eggs. EMBO J. 24(5), 1057–1067 (2005)
Yan, F., Liu, Z., Liu, H.: Dynamic analysis of the combinatorial regulation involving transcription factors and microRNAs in cell fate decisions. BBA-Proteins Proteomics 1844(1), 248–257 (2014)
Tokuda, I.T., Okamoto, A., Matsumura, R., Takumi, T., Akashi, M.: Potential contribution of tandem circadian enhancers to nonlinear oscillations in clock gene expression. Mol. Biol. Cell 28(17), 2333–2342 (2017)
Jia, D., Lu, M., Jung, K.H., Park, J.H., Yu, L., Onuchic, J.N., Kaipparettu, B.A., Levine, H.: Elucidating cancer metabolic plasticity by coupling gene regulation with metabolic pathways. Proc. Natl. Acad. Sci. USA 116(9), 3909–3918 (2019)
García-Gómez, M.L., Ornelas-Ayala, D., Garay-Arroyo, A., Garcia-Ponce, B., Lvarez-Buylla, E.R.: A system-level mechanistic explanation for asymmetric stem cell fates: Arabidopsis thaliana root niche as a study system. Sci. Rep. 10(1), 3525 (2020)
Hoa, T.T., Tortosa, P., Albano, M., Dubnau, D.: Rok (YkuW) regulates genetic competence in Bacillus subtilis by directly repressing comK. Mol. Microbiol. 43(1), 15–26 (2002)
Guan, Y., Li, Z., Wang, S., Barnes, P.M., Liu, X., Xu, H., Jin, M., Liu, A.P., Yang, Q.: A robust and tunable mitotic oscillator in artificial cells. eLife 7, e33549 (2018)
Lehár, J., Zimmermann, G.R., Krueger, A.S., Molnar, R.A., Ledell, J.T., Heilbut, A.M., Short, G.F., Giusti, L.C., Nolan, G.P., Magid, O.A.: Chemical combination effects predict connectivity in biological systems. Mol. Syst. Biol. 3(1), 80 (2007)
Duan, J., Li, B., Bhakta, M., Xie, S., Zhou, P., Munshi, N.V., Hon, G.C.: Rational reprogramming of cellular states by combinatorial perturbation. Cell Rep. 27(12), 3486–3499 (2019)
Yeo, G.T., Lin, L., Qi, C.Y., Cha, M., Gifford, D.K., Sherwood, R.I.: A multiplexed barcodelet single-cell RNA-seq approach elucidates combinatorial signaling pathways that drive ESC differentiation. Cell Stem Cell 26(6), 938–950 (2020)
Kim, J.R., Yoon, Y., Cho, K.H.: Coupled feedback loops form dynamic motifs of cellular networks. Biophys. J. 94(2), 359–365 (2008)
Miguel, R.P.J., Agata, O., Gerhard, M., Simona, S., Dominic, V.E., Jim, K.: Zbtb7a is a transducer for the control of promoter accessibility by NF-kappa B and multiple other transcription factors. Plos Biol. 16(5), e2004526 (2018)
Takaoka, K., Hamada, H.: Cell fate decisions and axis determination in the early mouse embryo. Development 139(1), 3–14 (2012)
Loewe, S.: The problem of synergy and antagonism of combined drugs. Arzneimittel-Forschung 3(7), 285–290 (1953)
Huang, B., Lu, M., Jia, D., Ben-Jacob, E., Levine, H., Onuchic, J.N.: Interrogating the topological robustness of gene regulatory circuits by randomization. Plos Comput. Biol. 13(3), e1005456 (2017)
Wang, N., Lefaudeux, D., Mazumder, A., Li, J.J., Hoffmann, A.: Identifying the combinatorial control of signal-dependent transcription factors. Plos Comput. Biol. 17(6), e1009095 (2021)
Dhooge, A., Govaerts, W., Kuznetsov, Y.A.: MATCONT: a MATLAB package for numerical bifurcation analysis of ODEs. ACM Trans. Math. Softw. 29(2), 141–164 (2003)
Funding
This research is supported by the National Natural Science Foundation of China (Grant No. 11971297).
Author information
Authors and Affiliations
Contributions
Menghan Chen: apply statistical and mathematical methods to analyze dynamical models and interpret simulation results; write and edit. Ruiqi Wang: provide ideas; formulate overall research goals and aims; supervise and revise this paper.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by the authors.
Informed consent
No individual participants are included in this study and no informed consent needs to be obtained.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendices
Appendix A
1.1 Formulation of logic gates
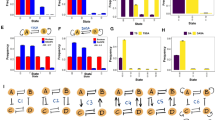
To identify the dependence of synergy on different logic, we enumerate six small networks, as shown in Fig. 1. Based on the different logic modeling methods (Section 4.2), ordinary differential equations of AND logic and OR logic for these six networks are given.

OR logic
AND logic

OR logic
AND logic

OR logic
AND logic

OR logic
AND logic

OR logic
AND logic

OR logic
AND logic
Here, variables A, B,and C represent the concentration of the three proteins at time t. The parameters used for the following calculations are listed in Table 5. All the bifurcation diagrams in the above text are drawn by software Matcont (a subpackage available under Matlab) [52], and all random and statistical processes are completed by Matlab.
Appendix B
1.1 Steady-state comparison of OR logic and AND logic
In this section, the steady state of AND and OR logic in Fig. 1a(1) are compared by theoretical calculation. Models (15) and (16) are used to calculate, \(K_b\) is regarded as the bifurcation parameter. Here, \((A^*_{OR}, B^*_{OR}, C^*_{OR})\) and \((A^*_{AND}, B^*_{AND}, C^*_{AND})\) denotes the low-state equilibrium point of the OR logic and AND logic. \((\tilde{A}_{OR}, \tilde{B}_{OR}, \tilde{C}_{OR})\) and \((\tilde{A}_{AND}, \tilde{B}_{AND}, \tilde{C}_{AND})\) denotes the high-state equilibrium point of the OR logic and AND logic.
First, the low steady state of OR logic is given. From Eq. (15), we know that when \(K_b\) \(\rightarrow\)0, the variable B\(\rightarrow\)0, i.e., B is in the low state. The A and C can be approximated
Substituting Eq. (27) into the formula B in model (15), we get
Substituting Eq. (28) into model (15), equilibrium points \(A^*_{OR}\) and \(C^*_{OR}\) can be obtained
For AND logic, substituting Eq. (27) into the formula B in model (16), we get
Substituting Eq. (31) into model (16), equilibrium points \(A^*_{AND}\) and \(C^*_{AND}\) can be obtained
When \(K_b\) is in the low state, the values of formulas (28) and (31) can be used to compare the B low steady state of OR logic and AND logic. The parameter values are replaced in Table 5, \(B_{OR}>B_{AND}\) can be obtained, which shows good consistency with Fig. 2.
Next, the high steady state of OR logic and AND logic in Fig. 1a(1) are compared. For OR logic, when \(K_b\rightarrow \infty\), \(B\rightarrow \frac{V_y}{b\left( 1+\left( \frac{C}{K_c}\right) ^2\right) }\). B is substituted into Eq. (15), we can obtain
The root of A and C can be determined by Eq. (34). Substituting the roots into model (15), we can get
Substituting Eq. (35) into model (15), \(\tilde{A}_{OR}\) and \(\tilde{C}_{OR}\) can be obtained
For AND logic, when \(K_b\rightarrow \infty\), \(B\rightarrow \frac{V_y}{b}\). A and C is obtained
Substituting Eq. (38) into the formula B of model (16), we can get
Substituting Eq. (39) into model (16), equilibrium points \(\tilde{A}_{AND}\) and \(\tilde{C}_{AND}\) are obtained
When \(K_b\) is in high steady state, Eqs. (35) and (39) can be used to compare the high steady state of OR logic and AND logic. \(K_b\) takes a larger value, and other values are replaced by Table 5, the calculation result is \(B_{AND}>B_{OR}\), which is consistent with Fig. 2.
1.2 Saddle-node bifurcation analysis of OR logic and AND logic
In this section, the bifurcation points of OR logic and AND logic are analyzed. Here, \(K_b\) is regarded as a bifurcation parameter. First, the bifurcation point of OR logic (i.e., model (15)), is calculated. It assumed that \((A^*, B^*, C^*)\) is the equilibrium point of the bifurcation point of \(K_b\).
Let \(\bar{A}=A-A^*\), \(\bar{B}=B-B^*\) and \(\bar{C}=C-C^*\). \(\bar{A}\), \(\bar{B}\) and \(\bar{C}\) are still represented by A, B and C respectively. Therefore, the linear form of the system (15) is expressed as follows
where
where
Therefore, the Jacobian matrix of system (15) is
The characteristic equation of the linearized system (15) is
Then, the exponential polynomial equation is obtained
As we all know, when the system produces saddle-node bifurcation, at least one of all eigenvalues is zero. Therefore, substituting \(\lambda =0\) into the polynomial Eq. (45), we can get
The next goal is to solve the bifurcation point \(K_b\) and equilibrium point \(B^*\). From Eqs. (46) and (15), we get
Then, substituting all the parameters in Table 5 except the control parameter \(K_b\) into Eq. (47). The bifurcation point \(K_b\) and the equilibrium point \(B^*\) can be obtained, i.e., (\(K_b=1.30864\), \(B^*=1.91313\)) and (\(K_b=1.56867\), \(B^*=0.917858\)), which are consistent with the blue curve in Fig. 2a and b.
Next, the bifurcation point of AND logic (i.e., model (16)), is analyzed. Here, it is supposed that \((\widetilde{A}, \widetilde{B}, \widetilde{C})\) is the equilibrium point of the bifurcation point \(K_b\).
Let \(\bar{A}=A-\widetilde{A}\), \(\bar{B}=B-\widetilde{B}\) and \(\bar{C}=C-\widetilde{C}\). \(\bar{A}\), \(\bar{B}\) and \(\bar{C}\) are still represented by A, B, C. Therefore, the linear form of the system (16) is described as follows
where
where
Therefore, the Jacobian matrix of system (16) is
The characteristic equation of the linearized system (15) is
Then, the exponential polynomial equation is obtained
Substituting this zero eigenvalue into Eq. (51), the following equation is obtained
From Eqs. (16) and (52), we get
Substituting all the parameters in Table 5 except the control parameter \(K_b\) into Eq. (53). The bifurcation point \(K_b\) and the equilibrium point \(\widetilde{B}\) of the AND logic are obtained, i.e., (\(K_b\)=0.366907, \(\widetilde{B}\)= 2.53285) and (\(K_b\)= 1.44897, \(\widetilde{B}\)= 0.5), which are consistent with red curve in Fig. 2a and b.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, M., Wang, R. Computational analysis of synergism in small networks with different logic. J Biol Phys 49, 1–27 (2023). https://doi.org/10.1007/s10867-022-09620-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10867-022-09620-0