Abstract
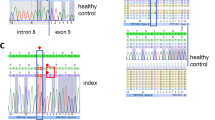
Type I inositol polyphosphate-4-phosphatase (INPP4A) belongs to the group of phosphoinositide phosphatases controlling proliferation, apoptosis, and endosome function by hydrolyzing phosphatidylinositol 3,4-bisphosphate. INPP4A produces multiple transcripts encoding shorter and longer INPP4A isoforms with hydrophilic or hydrophobic C-terminus. Biallelic INPP4A truncating variants cause a spectrum of neurodevelopmental disorders ranging from moderate intellectual disability to postnatal microcephaly with developmental and epileptic encephalopathy and (ponto)cerebellar hypoplasia. We report a girl with the novel homozygous INPP4A variant NM_001134224.2:c.2840del/p.(Gly947Glufs*12) (isoform d). She presented with postnatal microcephaly, global developmental delay, visual impairment, myoclonic seizures, and pontocerebellar hypoplasia and died at the age of 27 months. The level of mutant INPP4A mRNAs in proband-derived leukocytes was comparable to controls suggesting production of C-terminally altered INPP4A isoforms. We transiently expressed eGFP-tagged INPP4A isoform a (NM_004027.3) wildtype and p.(Gly908Glufs*12) mutant [p.(Gly947Glufs*12) according to NM_001134224.2] as well as INPP4A isoform b (NM_001566.2) wildtype and p.(Asp915Alafs*2) mutant, previously reported in family members with moderate intellectual disability, in HeLa cells and determined their subcellular distributions. While INPP4A isoform a was preferentially found in perinuclear clusters co-localizing with the GTPase Rab5, isoform b showed a net-like distribution, possibly localizing near and/or on microtubules. Quantification of intracellular localization patterns of the two INPP4A mutants revealed significant differences compared with the respective wildtype and similarity with each other. Our data suggests an important non-redundant function of INPP4A isoforms with hydrophobic or hydrophilic C-terminus in the brain.





Similar content being viewed by others
Data availability
Previously reported INPP4A variants and the INPP4A variant reported in this study have been deposited in the Leiden Open Variation Database (http://www.lovd.nl/INPP4A) with the accession IDs 0000872158, 0000872195, 0000872196, and 0000872197. The data that supports the findings of this study are available in the supporting information of this article. Exome sequencing data have not been made publicly available because of privacy protection.
References
Phan TK, Williams SA, Bindra GK, Lay FT, Poon IKH, Hulett MD (2019) Phosphoinositides: multipurpose cellular lipids with emerging roles in cell death. Cell Death Differ 26(5):781–793. https://doi.org/10.1038/s41418-018-0269-2
Di Paolo G, De Camilli P (2006) Phosphoinositides in cell regulation and membrane dynamics. Nature 443(7112):651–657. https://doi.org/10.1038/nature05185
Posor Y, Jang W, Haucke V (2022) Phosphoinositides as membrane organizers. Nat Rev Mol Cell Biol 23(12):797–816. https://doi.org/10.1038/s41580-022-00490-x
Li H, Marshall AJ (2015) Phosphatidylinositol (3,4) bisphosphate-specific phosphatases and effector proteins: a distinct branch of PI3K signaling. Cell Signal 27(9):1789–1798. https://doi.org/10.1016/j.cellsig.2015.05.013
Norris FA, Auethavekiat V, Majerus PW (1995) The isolation and characterization of cDNA encoding human and rat brain inositol polyphosphate 4-phosphatase. J Biol Chem 270(27):16128–16133. https://doi.org/10.1074/jbc.270.27.16128
Norris FA, Majerus PW (1994) Hydrolysis of phosphatidylinositol 3,4-bisphosphate by inositol polyphosphate 4-phosphatase isolated by affinity elution chromatography. J Biol Chem 269(12):8716–8720
Dyson JM, Fedele CG, Davies EM, Becanovic J, Mitchell CA (2012) Phosphoinositide phosphatases: just as important as the kinases. Subcell Biochem 58:215–279. https://doi.org/10.1007/978-94-007-3012-0_7
Ivetac I, Munday AD, Kisseleva MV, Zhang XM, Luff S, Tiganis T, Whisstock JC, Rowe T, Majerus PW, Mitchell CA (2005) The type Ialpha inositol polyphosphate 4-phosphatase generates and terminates phosphoinositide 3-kinase signals on endosomes and the plasma membrane. Mol Biol Cell 16(5):2218–2233. https://doi.org/10.1091/mbc.e04-09-0799
Norris FA, Atkins RC, Majerus PW (1997) Inositol polyphosphate 4-phosphatase is inactivated by calpain-mediated proteolysis in stimulated human platelets. J Biol Chem 272(17):10987–10989. https://doi.org/10.1074/jbc.272.17.10987
Shearn CT, Walker J, Norris FA (2001) Identification of a novel spliceoform of inositol polyphosphate 4-phosphatase type Ialpha expressed in human platelets: structure of human inositol polyphosphate 4-phosphatase type I gene. Biochem Biophys Res Commun 286(1):119–125. https://doi.org/10.1006/bbrc.2001.5331
Nystuen A, Legare ME, Shultz LD, Frankel WN (2001) A null mutation in inositol polyphosphate 4-phosphatase type I causes selective neuronal loss in weeble mutant mice. Neuron 32(2):203–212. https://doi.org/10.1016/s0896-6273(01)00468-8
Sasaki J, Kofuji S, Itoh R, Momiyama T, Takayama K, Murakami H, Chida S, Tsuya Y, Takasuga S, Eguchi S, Asanuma K, Horie Y, Miura K, Davies EM, Mitchell C, Yamazaki M, Hirai H, Takenawa T, Suzuki A, Sasaki T (2010) The PtdIns(3,4)P(2) phosphatase INPP4A is a suppressor of excitotoxic neuronal death. Nature 465(7297):497–501. https://doi.org/10.1038/nature09023
Sachs AJ, David SA, Haider NB, Nystuen AM (2009) Patterned neuroprotection in the Inpp4a(wbl) mutant mouse cerebellum correlates with the expression of Eaat4. PLoS One 4(12):e8270. https://doi.org/10.1371/journal.pone.0008270
Najmabadi H, Hu H, Garshasbi M, Zemojtel T, Abedini SS, Chen W, Hosseini M, Behjati F, Haas S, Jamali P, Zecha A, Mohseni M, Puttmann L, Vahid LN, Jensen C, Moheb LA, Bienek M, Larti F, Mueller I, Weissmann R, Darvish H, Wrogemann K, Hadavi V, Lipkowitz B, Esmaeeli-Nieh S, Wieczorek D, Kariminejad R, Firouzabadi SG, Cohen M, Fattahi Z, Rost I, Mojahedi F, Hertzberg C, Dehghan A, Rajab A, Banavandi MJ, Hoffer J, Falah M, Musante L, Kalscheuer V, Ullmann R, Kuss AW, Tzschach A, Kahrizi K, Ropers HH (2011) Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature 478(7367):57–63. https://doi.org/10.1038/nature10423
Sheffer R, Bennett-Back O, Yaacov B, Edvardson S, Gomori M, Werner M, Fahham D, Anteby I, Frumkin A, Meiner V, Elpeleg O (2015) Hindbrain malformation and myoclonic seizures associated with a deleterious mutation in the INPP4A gene. Neurogenetics 16(1):23–26. https://doi.org/10.1007/s10048-014-0428-7
Banihashemi S, Tahmasebi-Birgani M, Mohammadiasl J, Hajjari M (2020) Whole exome sequencing identified a novel nonsense INPP4A mutation in a family with intellectual disability. Eur J Med Genet 63(4):103846. https://doi.org/10.1016/j.ejmg.2020.103846
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, Chen X, Kim Y, Beyter D, Krusche P, Saunders CT (2018) Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods 15(8):591–594. https://doi.org/10.1038/s41592-018-0051-x
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR, Thormann A, Flicek P, Cunningham F (2016) The ensembl variant effect predictor. Genome Biol 17(1):122. https://doi.org/10.1186/s13059-016-0974-4
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q, Collins RL, Laricchia KM, Ganna A, Birnbaum DP, Gauthier LD, Brand H, Solomonson M, Watts NA, Rhodes D, Singer-Berk M, England EM, Seaby EG, Kosmicki JA, Walters RK, Tashman K, Farjoun Y, Banks E, Poterba T, Wang A, Seed C, Whiffin N, Chong JX, Samocha KE, Pierce-Hoffman E, Zappala Z, O’Donnell-Luria AH, Minikel EV, Weisburd B, Lek M, Ware JS, Vittal C, Armean IM, Bergelson L, Cibulskis K, Connolly KM, Covarrubias M, Donnelly S, Ferriera S, Gabriel S, Gentry J, Gupta N, Jeandet T, Kaplan D, Llanwarne C, Munshi R, Novod S, Petrillo N, Roazen D, Ruano-Rubio V, Saltzman A, Schleicher M, Soto J, Tibbetts K, Tolonen C, Wade G, Talkowski ME, Genome Aggregation Database C, Neale BM, Daly MJ, MacArthur DG (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581(7809):434–443. https://doi.org/10.1038/s41586-020-2308-7
Coban-Akdemir Z, White JJ, Song X, Jhangiani SN, Fatih JM, Gambin T, Bayram Y, Chinn IK, Karaca E, Punetha J, Poli C, Baylor-Hopkins Center for Mendelian G, Boerwinkle E, Shaw CA, Orange JS, Gibbs RA, Lappalainen T, Lupski JR, Carvalho CMB (2018) Identifying genes whose mutant transcripts cause dominant disease traits by potential gain-of-function alleles. Am J Hum Genet 103(2):171–87.https://doi.org/10.1016/j.ajhg.2018.06.009
Shin HW, Hayashi M, Christoforidis S, Lacas-Gervais S, Hoepfner S, Wenk MR, Modregger J, Uttenweiler-Joseph S, Wilm M, Nystuen A, Frankel WN, Solimena M, De Camilli P, Zerial M (2005) An enzymatic cascade of Rab5 effectors regulates phosphoinositide turnover in the endocytic pathway. J Cell Biol 170(4):607–618. https://doi.org/10.1083/jcb.200505128
Wang H, Loerke D, Bruns C, Muller R, Koch PA, Puchkov D, Schultz C, Haucke V (2020) Phosphatidylinositol 3,4-bisphosphate synthesis and turnover are spatially segregated in the endocytic pathway. J Biol Chem 295(4):1091–1104. https://doi.org/10.1074/jbc.RA119.011774
Acknowledgements
We thank the patient’s family for participation in this study. We further thank Henrike Wilshusen for skillful technical assistance and the UKE Microscopy Imaging Facility (UMIF) at the University Medical Center Hamburg-Eppendorf for technical support. This study was supported by grants from the Deutsche Forschungsgemeinschaft (KU 1240/13-1 to K.K.) and the University Children’s Hospital at the University Medical Center Hamburg-Eppendorf (ped.tracks to L.H.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Laura Hecher and Frederike L. Harms should be considered joint first authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hecher, L., Harms, F.L., Lisfeld, J. et al. INPP4A-related genetic and phenotypic spectrum and functional relevance of subcellular targeting of INPP4A isoforms. Neurogenetics 24, 79–93 (2023). https://doi.org/10.1007/s10048-023-00709-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-023-00709-9