Abstract
Backgrounds
Among tumor microenvironment, the immune components in it have an important influence on gene expression and clinical efficacy. We aim to find out the role of those in skin cutaneous melanoma (SKCM).
Objectives
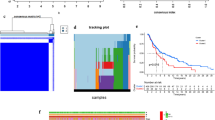
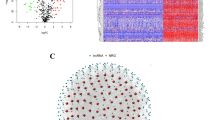
Gene expression profile and homologous clinical information of SKCM patients were obtained by TCGA (The Cancer Genome Atlas) and UCSC Toil. SsGSEA method was used to evaluate the immune cell infiltration of 468 TCGA-SKCM samples divided into high immune cell infiltration group (HICI) and low immune cell infiltration group (LICI). We used the Edger packet to conduct difference analysis on normal samples (GTEx) and cancer samples (TCGA), and combined it with the difference of the HICI group and LICI group, to find out the common differential expression of lncRNA in both groups. The prognostic value of immune-related lncRNAs was studied by univariate Cox, Lasso-Cox and multivariate Cox regression analysis, and a prognostic model was established. C index and calibration diagram were used to judge the accuracy of the model, and DCA was used to judge the net benefit.
Results
Six prognostic markers of immune-related lncRNA genes were established, which could be used as independent prognostic factors. The net benefit and prediction accuracy are significantly higher than TNM Stage.
Conclusion
The prognostic model identified in this study is a reliable biomarker for SKCM. The Nomogram survival prediction model based on it is a reliable way to predict the median survival time of patients, which may lay the foundation for future treatment of this disease.






Similar content being viewed by others
Data availability
The datasets analyzed in this study were obtained from The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/).
References
Arun G, Diermeier SD, Spector DL (2018) Therapeutic targeting of long non-coding RNAs in cancer. Trends Mol Med 24(3):257–277
Cameron-Mowat D et al (1973) Cellular immunity in human malignant melanoma and melanoma histology. Br J Cancer 28(1):77
Coe EA et al (2019) The MITF-SOX10 regulated long non-coding RNA DIRC3 is a melanoma tumour suppressor. PLoS Genet 15(12):e1008501
Dashti N et al (2018) Evaluation of ITGB2 (CD18) and SELL (CD62L) genes expression and methylation of ITGB2 promoter region in patients with systemic sclerosis. Rheumatol Int 38(3):489–498
Galatro TF et al (2017) Transcriptomic analysis of purified human cortical microglia reveals age-associated changes. Nat Neurosci 20(8):1162
Goldstein AM, Tucker MA (2001) Genetic epidemiology of cutaneous melanoma: a global perspective. Arch Dermatol 137(11):1493–1496
Gong J et al (2017) Long noncoding RNA linc00462 promotes hepatocellular carcinoma progression. Biomed Pharmacother 93:40–47
He Y et al (2018) Classification of triple-negative breast cancers based on Immunogenomic profiling. J Exp Clin Cancer Res 37(1):1–13
He S et al (2020) TRG-AS1 is a potent driver of oncogenicity of tongue squamous cell carcinoma through microRNA-543/Yes-associated protein 1 axis regulation. Cell Cycle 19(15):1969–1982
He H et al (2021) Tape strips detect distinct immune and barrier profiles in atopic dermatitis and psoriasis. J Allergy Clin Immunol 147(1):199–212
Hu T, Wang F, Han G (2020) LncRNA PSMB8-AS1 acts as ceRNA of miR-22-3p to regulate DDIT4 expression in glioblastoma. Neurosci Lett 728:134896
Hyland PL et al (2014) Constitutional promoter methylation and risk of familial melanoma. Epigenetics 9(5):685–692
Iasonos A et al (2008) How to build and interpret a nomogram for cancer prognosis. J Clin Oncol 26(8):1364–1370
Jiao X et al (2019) Recent advances targeting CCR5 for cancer and its role in immuno-oncology. Can Res 79(19):4801–4807
Jin L et al (2020) A potential prognostic prediction model of colon adenocarcinoma with recurrence based on prognostic lncRNA signatures. Hum Genomics 14(1):1–11
Jin D, et al. Identification of a Seven-lncRNA Immune Risk Signature and Construction of a Predictive Nomogram for Lung Adenocarcinoma. BioMed Research International, 2020. 2020.
Lagou V. et al. (2018) Genetic architecture of adaptive immune system identifies key immune regulators. Cell reports. 25(3): 798–810. e6.
Lee K, Hwang H, Nam KT (2014) Immune response and the tumor microenvironment: how they communicate to regulate gastric cancer. Gut and Liver 8(2):131
Lipson EJ et al. (2015) Antagonists of PD-1 and PD-L1 in cancer treatment. in Seminars in oncology. Elsevier.
Mariathasan S et al (2018) TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554(7693):544–548
Mascaro J et al (1984) Why do melanomas ulcerate? J Cutan Pathol 11(4):269–273
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10(3):155–159
Miller KD et al. Cancer treatment and survivorship statistics, 2016. CA: a cancer journal for clinicians, 2016. 66(4): 271–289.
Newman AM et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12(5):453–457
Paul S et al (2016) Intratumoral natural killer cells show reduced effector and cytolytic properties and control the differentiation of effector Th1 cells. Oncoimmunology 5(12):e1235106
Rowling EJ et al (2020) Cooperative behaviour and phenotype plasticity evolve during melanoma progression. Pigment Cell Melanoma Res 33(5):695
Schank TE, Hassel JC (2019) Immunotherapies for the treatment of uveal Melanoma—History and future. Cancers 11(8):1048
Shen G et al (2020) PSMB8-AS1 activated by ELK1 promotes cell proliferation in glioma via regulating miR-574-5p/RAB10. Biomed Pharmacother 122:109658
Šitum M et al (2014) Melanoma–clinical, dermatoscopical, and histopathological morphological characteristics. Acta Dermatovenerol Croat 22(1):2–2
Souri Z et al (2019) HLA expression in uveal melanoma: an indicator of malignancy and a modifiable immunological target. Cancers 11(8):1132
Therneau TM, Grambsch PM (2000) The Cox model. Modeling survival data: extending the Cox model. Springer, pp 39–77
Vickers AJ, Elkin EB (2006) Decision curve analysis: a novel method for evaluating prediction models. Med Decis Making 26(6):565–574
Wang T et al (2015) BRAF inhibition stimulates melanoma-associated macrophages to drive tumor growth. Clin Cancer Res 21(7):1652–1664
Whiteman DC, Green AC, Olsen CM (2016) The growing burden of invasive melanoma: projections of incidence rates and numbers of new cases in six susceptible populations through 2031. J Investig Dermatol 136(6):1161–1171
Xiao W, Yin A (2019) LINC0638 lncRNA is involved in the local recurrence of melanoma following surgical resection. Oncol Lett 18(1):101–108
Xiong Tf et al. (2020) Prognostic value of the expression of chemokines and their receptors in regional lymph nodes of melanoma patients. J Cell Mol Med 24(6): 3407–3418.
Yoshida K, Bohn J, Yoshida MK (2020) Package ‘tableone’. R Foundation for Statistical Computing, Vienna, Austria (30 November 2016).
Yoshihara K et al (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4(1):1–11
Young HL et al (2017) An adaptive signaling network in melanoma inflammatory niches confers tolerance to MAPK signaling inhibition. J Exp Med 214(6):1691–1710
Zhao Y et al. (2019) PD-L1: CD80 cis-heterodimer triggers the co-stimulatory receptor CD28 while repressing the inhibitory PD-1 and CTLA-4 pathways. Immunity. 51(6): 1059–1073. e9.
Zhou B et al (2018) Linc00462 promotes pancreatic cancer invasiveness through the miR-665/TGFBR1-TGFBR2/SMAD2/3 pathway. Cell Death Dis 9(6):1–15
Acknowledgements
We gratefully acknowledge Chengcheng He for his technical support on this manuscript.
Funding
This study was supported by the Luzhou Science and Technology Department Project (2020-JYJ-44), Sichuan Science and Technology of Traditional Chinese Medicine Administration Project (2020JC0134), and the Science Foundation of Southwest Medical University (2020ZRZD001).
Author information
Authors and Affiliations
Contributions
YC and GZ conceived and designed the study; NH, CH and YH performed data analysis; SL and JY assisted in data curation and analyzed the references; NH and YC wrote the paper; all authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Compliance with ethical standards
The article contains no research in which animals were used.
Statement of Ethics
This article does not contain any studies with human participants or animals performed by any of the authors; thus, ethical approval and informed consent are not required.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, N., Huang, C., He, Y. et al. A novel immune-related LncRNA prognostic signature for cutaneous melanoma. Mol. Cell. Toxicol. 20, 377–387 (2024). https://doi.org/10.1007/s13273-023-00351-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13273-023-00351-4