Abstract
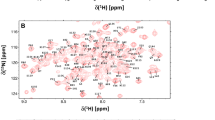
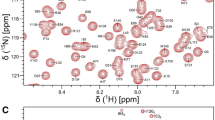
N-methyl-D-aspartate receptors (NMDARs) consist of glycine-binding GluN1 and glutamate-binding GluN2 subunits that form tetrameric ion channels. NMDARs in the neuronal post-synaptic membrane are important for controlling neuroplasticity and synaptic transmission in the brain. Calmodulin (CaM) binds to the cytosolic C0 domains of both GluN1 (residues 841–865) and GluN2 (residues 1004–1024) that may play a role in the Ca2+-dependent desensitization of NMDAR channels. Mutations that disrupt Ca2+-dependent desensitization of NMDARs are linked to Alzheimer’s disease, depression, stroke, epilepsy, and schizophrenia. NMR chemical shift assignments are reported here for Ca2+-saturated CaM bound to the GluN2A C0 domain of NMDAR (BMRB no. 51821).



Similar content being viewed by others
Data availability
The assignments have been deposited to the BMRB under the accession code: 51821.
References
Babu YS, Bugg CE, Cook WJ (1988) Structure of calmodulin refined at 2.2 A resolution. J Mol Biol 204:191–204
Bajaj G, Hau AM, Hsu P, Gafken PR, Schimerlik MI, Ishmael JE (2014) Identification of an atypical calcium-dependent calmodulin binding site on the C-terminal domain of GluN2A. Biochem Biophys Res Commun 444:588–594
Bej A, Ames JB (2022a) Chemical shift assignments of calmodulin bound to the beta-subunit of a retinal cyclic nucleotide-gated channel (CNGB1). Biomol NMR Assign 16:147–151
Bej A, Ames JB (2022b) Chemical shift assignments of calmodulin under standard conditions at neutral pH. Biomol NMR Assign 16:213–218
Bej A, Ames JB (2022c) NMR structures of calmodulin bound to two separate regulatory sites in the retinal cyclic nucleotide-gated channel. Biochemistry 61:1955–1965
Bej A, Ames JB (2023) Chemical shift assignments of calmodulin bound to the GluN1 C0 domain (residues 841–865) of the NMDA receptor. Biomolecular NMR Assignments in Press: https://doi.org/10.1007/s12104-023-10121-x
Benveniste M, Mayer ML (1991) Kinetic analysis of antagonist action at N-methyl-D-aspartic acid receptors. Two binding sites each for glutamate and glycine. Biophys J 59:560–573
Chou TH, Tajima N, Romero-Hernandez A, Furukawa H (2020) Structural basis of functional transitions in mammalian NMDA receptors. Cell 182(357–371):e13
Clements JD, Westbrook GL (1991) Activation kinetics reveal the number of glutamate and glycine binding sites on the N-methyl-D-aspartate receptor. Neuron 7:605–613
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeiffer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Halling DB, Georgiou DK, Black DJ, Yang G, Fallon JL, Quiocho FA, Pedersen SE, Hamilton SL (2009) Determinants in CaV1 channels that regulate the Ca2+ sensitivity of bound calmodulin. J Biol Chem 284:20041–20051
Iacobucci GJ, Popescu GK (2017) Resident calmodulin primes NMDA receptors for Ca(2+)-dependent inactivation. Biophys J 113:2236–2248
Iacobucci GJ, Popescu GK (2019) Spatial coupling tunes NMDA receptor responses via Ca(2+) diffusion. J Neurosci off J Soc Neurosci 39:8831–8844
Iacobucci GJ, Popescu GK (2020) Ca(2+)-dependent inactivation of GluN2A and GluN2B NMDA receptors occurs by a common kinetic mechanism. Biophys J 118:798–812
Jalali-Yazdi F, Chowdhury S, Yoshioka C, Gouaux E (2018) Mechanisms for zinc and proton inhibition of the GluN1/GluN2A NMDA receptor. Cell 175(1520–1532):e15
Karakas E, Furukawa H (2014) Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344:992–997
Krupp JJ, Vissel B, Thomas CG, Heinemann SF, Westbrook GL (1999) Interactions of calmodulin and alpha-actinin with the NR1 subunit modulate Ca2+-dependent inactivation of NMDA receptors. J Neurosci 19:1165–1178
Kunz PA, Roberts AC, Philpot BD (2013) Presynaptic NMDA receptor mechanisms for enhancing spontaneous neurotransmitter release. J Neurosci 33:7762–7769
Lee CH, Lu W, Michel JC, Goehring A, Du J, Song X, Gouaux E (2014) NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511:191–197
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Neri D, Szyperski T, Otting G, Senn H, Wuthrich K (1989) Stereospecific nuclear magnetic resonance assignments of the methyl groups of valine and leucine in the DNA-binding domain of the 434 repressor by biosynthetically directed fractional 13C labeling. Biochemistry 28:7510–7516
Peng TI, Jou MJ, Sheu SS, Greenamyre JT (1998) Visualization of NMDA receptor-induced mitochondrial calcium accumulation in striatal neurons. Exp Neurol 149:1–12
Puri BK (2020) Calcium signaling and gene expression. Adv Exp Med Biol 1131:537–545
Regan MC, Grant T, McDaniel MJ, Karakas E, Zhang J, Traynelis SF, Grigorieff N, Furukawa H (2018) Structural mechanism of functional modulation by gene splicing in NMDA receptors. Neuron 98(521–529):e3
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Sprenger J, Trifan A, Patel N, Vanderbeck A, Bredfelt J, Tajkhorshid E, Rowlett R, Lo Leggio L, Akerfeldt KS, Linse S (2021) Calmodulin complexes with brain and muscle creatine kinase peptides. Curr Res Struct Biol 3:121–132
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, Hansen KB, Yuan H, Myers SJ, Dingledine R (2010) Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev 62:405–496
Wadel K, Neher E, Sakaba T (2007) The coupling between synaptic vesicles and Ca2+ channels determines fast neurotransmitter release. Neuron 53:563–575
Wang C, Wang HG, Xie H, Pitt GS (2008) Ca2+/CaM controls Ca2+-dependent inactivation of NMDA receptors by dimerizing the NR1 C termini. J Neurosci off J Soc Neurosci 28:1865–1870
Williamson MP (2013) Using chemical shift perturbation to characterise ligand binding. Prog Nucl Magn Reson Spectrosc 73:1–16
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651
Zhang X, Majerus PW (1998) Phosphatidylinositol signalling reactions. Semin Cell Dev Biol 9:153–160
Acknowledgements
We thank Derrick Kaseman and Ping Yu for help with NMR experiments performed at the UC Davis NMR Facility.
Funding
Work supported by NIH grants to J.B.A (R01 EY012347) and to the UC Davis NMR Facility (RR11973).
Author information
Authors and Affiliations
Contributions
A.B. performed all experiments, analyzed data and helped write the manuscript. J.B.A directed the overall project and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing conflict of interest.
Ethical approval
The experiments comply with the current laws of the United States.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bej, A., Ames, J.B. Chemical shift assignments of calmodulin bound to a cytosolic domain of GluN2A (residues 1004–1024) from the NMDA receptor. Biomol NMR Assign 17, 89–93 (2023). https://doi.org/10.1007/s12104-023-10125-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10125-7