Abstract
Retinal membrane guanylyl cyclases (RetGCs) in vertebrate rod and cone photoreceptors are activated by a family of neuronal Ca2+ sensor proteins called guanylyl cyclase activating proteins (GCAP1-7). GCAP5 from zebrafish photoreceptors binds to RetGC and confers Ca2+/Fe2+-dependent regulation of RetGC enzymatic activity that promotes the recovery phase of visual phototransduction. We report NMR chemical shift assignments of GCAP5 with a R22A mutation (called GCAP5R22A) that abolishes protein dimerization and activates RetGC with 3-fold higher activity than that of wild type GCAP5 (BMRB No. 51,783).
Similar content being viewed by others
Biological context
Guanylyl cyclase activating proteins (GCAP1-7) are Ca2+-binding proteins in the retina that belong to a sub-branch of the calmodulin superfamily (Burgoyne 2007, Lim, Dizhoor et al., 2014). GCAP proteins contain an N-terminal myristoyl group and four EF-hand motifs that bind to Ca2+ at EF2, EF3 and EF4 (Ames 2021). The first EF-hand contains residues that disable Ca2+ binding and the Ca2+-free EF1 interacts with the N-terminal myristoyl group (Cudia, Roseman et al., 2021, Stephen, Bereta et al., 2007). The Ca2+-bound GCAPs bind to RetGC and inhibit its cyclase activity, whereas Ca2+-free GCAPs activate RetGC enzymatic activity during the recovery phase of visual phototransduction (Koch and Stryer 1988, Palczewski, Subbaraya et al., 1994, Peshenko and Dizhoor 2007). Light activation of retinal photoreceptor cells causes a decrease in the cytosolic Ca2+ concentration that serves as a coordinating signal for visual recovery (Arshavsky and Burns 2014). The light-induced drop in cytosolic Ca2+ concentration is sensed by GCAPs that promote Ca2+-sensitive activation of RetGC to replenish cGMP levels in order to restore the dark state (Koch and Helten 2008; Koch and Stryer 1988). Mutations in GCAP1 that weaken Ca2+ binding or otherwise alter Ca2+-sensitive activation of RetGC are genetically linked to retinal diseases (Jiang and Baehr 2010, Payne, Downes et al., 1998).
GCAP5 in zebrafish photoreceptors binds to both Ca2+ and Fe2+ (Lim et al. 2017). The Ca2+-free forms of GCAP1 (Peshenko and Dizhoor 2006) and GCAP5 (Lim et al. 2017) both activate RetGC activity in light-adapted photoreceptors, whereas the Ca2+-bound GCAP1 (Peshenko and Dizhoor 2007) and Fe2+-bound GCAP5 (Lim et al. 2017) both inhibit RetGC in dark-adapted photoreceptors. The NMR structure of GCAP5 (Cudia et al. 2021) revealed that GCAP5 forms a dimer in solution with key amino acid residues at the dimer interface (H18, Y21, R22, M25, F72, V76 and W93) that are important for cyclase activation. The GCAP5 mutations H18E, M25E and V76E each abolish GCAP5 dimerization and prevent activation of RetGC (Cudia et al. 2021). These results suggested that GCAP5 dimerization might be essential for RetGC activation (Ames 2021, 2022). However, this hypothesis was refuted by the discovery that the R22A mutation of GCAP5 not only abolishes GCAP5 dimerization but also causes a 300% increase in RetGC activation compared to that of wild type (Cudia et al. 2021). We hypothesize that the R22A mutation might somehow alter the structure of GCAP5 to abolish its dimerization and increase its potency for activating RetGC. We report here NMR resonance assignments for the Ca2+-free activator and monomeric form of Ca2+-free GCAP5 with the R22A mutation (called GCAP5R22A) to understand how this mutation abolishes protein dimerization and causes a 300% increase of RetGC activity compared to that of wild type GCAP5.
Methods and experiments
Preparation of GCAP5
Samples of recombinant myristoylated GCAP5R22A (residues 2-198) uniformly labeled with 15 N and 13 C were prepared as described previously for wild type GCAP5 (Cudia and Ames 2019; Cudia et al. 2021).
NMR spectroscopy
NMR samples of Ca2+-free and myristoylated GCAP5R22A were prepared as described previously for wild type GCAP5 (Cudia et al. 2021). The NMR samples consisted of 0.3 mM protein dissolved in 5 mM TRIS-d11 (pH 7.4), 2 mM DTT-d10, 1 mM EDTA, 1 mM EGTA, 0.04% w/v NaN3, and 92% H2O/7% D2O. All NMR experiments were performed at 32 °C on a Bruker Avance 600 MHz spectrometer equipped with a triple resonance cryogenic (TCI probe) as described previously (Cudia and Ames 2019). The following 3D NMR experiments (HNCA, HNCACB, HNCOCACB, HNCO, HBHACONH, and HBHANH) were analyzed to obtain backbone assignments (Ikura, Kay et al., 1990). Side chain resonances were assigned by analyzing HBCBCGCDHD, HBCBCGCDHDCEHE, and HCCH-TOCSY (Ikura, Spera et al., 1991). The software NMRPipe (Delaglio, Grzesiek et al., 1995) was used to process all NMR data, and Sparky NMRFAM (Lee, Tonelli et al., 2015) was used to obtain resonance assignments.
Extent of assignments and data deposition
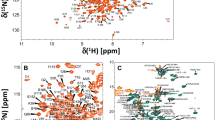
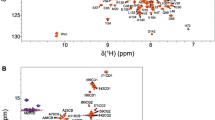
Representative NMR assignments are illustrated by two-dimensional NMR spectra of Ca2+-free GCAP5R22A (15 N-1 H HSQC, Fig. 1A-B and 13 C-1 H HSQC, Fig. 1C). The resonance assignments were determined by analyzing 3D triple resonance NMR spectra of 13 C/15 N-labeled GCAP5R22A. The highly resolved NMR peaks with uniform intensities indicate a stable and folded structure. Amide resonances assigned to Q19, L33 and I70 exhibited noteworthy downfield shifts, perhaps because these residues are flanked by nearby aromatic rings (W20, F35 and F72 respectively) (Fig. 1A). The amide resonances assigned to G68 and G147 have downfield chemical shifts that are caused by a strong hydrogen bond between the backbone NH of G68 (EF2)/G147 (EF4) with side chain carboxyl groups of D63 (EF2)/D142 (EF4), respectively. These strong hydrogen bonds are stabilized by an open conformation for both EF2 and EF4. It is unusual for Ca2+-free EF-hands to occupy an open conformation that is typically only formed by Ca2+-bound EF-hands (Ikura 1996, Yap, Ames et al., 1999). However, the NMR structure of wild type Ca2+-free GCAP5 revealed that the Ca2+-free structures of EF2, EF3 and EF4 each adopt a pre-formed open conformation (Cudia et al. 2021), which might explain why the GCAP proteins exhibit such high affinity Ca2+ binding in the nanomolar range (Lim, Peshenko et al., 2009). Spectral assignments were obtained for more than 94% of the main chain 13 C resonances (13Cα, 13Cβ, and 13CO), 97% of non-proline backbone amide resonances (1HN, 15 N), and 87% of side chain resonances (Fig. 1C). The unassigned residues (A22, N46, E74, Y75, and I136) had weak HSQC peaks caused by exchange broadening that prevented their assignment. Complete chemical shift assignments (1H, 15N, 13C) of Ca2+-free GCAP5R22A have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under accession number 51,783.
A Two-dimensional 15 N-1 H HSQC spectrum of 15 N-labeled Ca2+-free GCAP5R22A illustrates backbone amide assignments. B Expanded view of the spectrally crowded central region surrounded by a box in panel A. C Constant-time 13 C-1 H HSQC spectrum of 13 C-labeled Ca2+-free GCAP5R22A illustrates side-chain methyl assignments indicated by residue labels
Chemical shift index (Wishart, Sykes et al., 1992) and secondary structure prediction software using TALOS+ (Shen, Delaglio et al., 2009) were both used to calculate the secondary structure of Ca2+-free GCAP5R22A (Fig. 2A, B). GCAP5R22A has the same secondary structure that was reported previously for wild type GCAP5 (Cudia and Ames 2019): The protein has 10 α-helices: H1 (residues 8–14), H2 (residues 18–26), H3 (residues 35–41), H4 (residues 49–62), H5 (residues 74–82), H6 (residues 87–95), H7 (residues 110–117), H8 (residues 129–135), H9 (residues 150–160) and H10 (residues 162–172) shown as cylinders in Fig. 2B. Helices H2–H9 form four EF-hand motifs as seen in previous structures of GCAP1 (Lim, Peshenko et al., 2016, Stephen et al. 2007) and GCAP5 (Cudia et al. 2021). A 3-residue β-strand is observed in the Ca2+-free binding loops of EF1 and EF2 (shown as red arrows in Fig. 2A). This β-strand is only partially formed in the third and fourth EF-hands of Ca2+-free GCAP5. The final 14 residues from the C-terminus in GCAP5R22A (residues 184–198) are dynamically disordered and unstructured like was seen in previous structures of wild type GCAP5 (Cudia et al. 2021) and GCAP1 (Stephen et al. 2007).
Secondary structure and RCI order parameter (S2) of Ca2+-free GCAP5R22A predicted from the assigned backbone chemical shifts. A Probability of secondary structural elements (blue cylinders for helix and red arrow for strand) and B RCI S2 of Ca2+-free GCAP5R22A were calculated using TALOS + server (Shen et al. 2009)
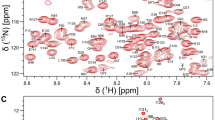
The assigned amide chemical shifts of Ca2+-free GCAP5R22A (BMRB 51,783) are compared to those of Ca2+-free GCAP5 wild type (BMRB 51,784) to help identify residues that are structurally affected by the R22A mutation (Fig. 3A). Not surprisingly, the GCAP5 residues in EF1 (Q19, W20, Y21 and K23) that are closest to R22A exhibit the largest chemical shift perturbation (Fig. 3A, B). In addition, C-terminal residues R176, I177 and V178 also exhibit detectably large chemical shift perturbations. In the wild type GCAP5 structure (Cudia et al. 2021), the side-chain methyl groups of I177 are in close proximity with the side chain indole group of W20, and both side chains make close contact with the N-terminal myristoyl group (Fig. 3C). Interestingly, the myristoyl group contacts with both W20 and I177 are both important for the proposed Ca2+-myristoyl tug mechanism that transmits Ca2+-induced conformational changes from the EF-hands to the myristoyl group (Peshenko, Olshevskaya et al., 2012). We suggest that the R22A mutation may stabilize the Ca2+-free GCAP5 activator conformation by disrupting the Ca2+-myristoyl tug (Peshenko et al. 2012). The NMR assignments of Ca2+-free GCAP5R22A presented here suggest the R22A mutation affects the structure in both EF1 (W20) and C-terminal region (I177) that may play a role in disrupting GCAP5 dimerization and enhancing cyclase activation.
Chemical shift perturbation (CSP) for Ca2+-free GCAP5R22A versus wild type GCAP5. A Backbone amide CSP was calculated as: \(CSP= \sqrt{{\left(\varDelta {H}^{N}\right)}^{2}+{\left(0.14\times \varDelta N\right)}^{2}}\). ΔHN and ΔN are the observed difference in the 1HN and 15 N chemical shifts, respectively between Ca2+-free GCAP5R22A (BMRB 51,783) and wild type GCAP5 (BMRB 51,783). B CSPs mapped on the structure of GCAP5 (Cudia et al. 2021). C Close-up view of the myristoyl group binding site environment in GCAP5. The side chains of W20 and I177 make close contacts with the myristoyl group
Data availability
The assignments have been deposited to the BMRB under the accession code: 51,783.
References
Ames JB (2021) Structural insights into retinal guanylate cyclase activator proteins (GCAPs). Int J Mol Sci 22:8731
Ames JB (2022) Structural basis of retinal membrane guanylate cyclase regulation by GCAP1 and RD3. Front Mol Neurosci 15:988142
Arshavsky VY, Burns ME (2014) Current understanding of signal amplification in phototransduction. Cell logistics 4:e29390
Burgoyne RD (2007) Neuronal calcium sensor proteins: generating diversity in neuronal Ca2 + signalling. Nat Rev Neurosci 8:182–193
Cudia D, Ames J (2019) Chemical shift assignments of retinal guanylyl cyclase activating protein 5 (GCAP5). Biomol NMR Assign 13:201–205
Cudia D, Roseman GP, Assafa TE, Shahu MK, Scholten A, Menke-Sell SK, Yamada H, Koch KW, Milhauser G, Ames JB (2021) NMR and EPR-DEER structure of a Dimeric Guanylate Cyclase activator Protein-5 from zebrafish photoreceptors. Biochemistry 60:3058–3070
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeiffer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Ikura M (1996) Calcium binding and conformational response in EF-hand proteins. Trends Biochem Sci 21:14–17
Ikura M, Kay LE, Bax A (1990) A novel approach for sequential assignment of 1H, 13 C, and 15 N spectra of proteins: heteronuclear triple-resonance three-dimensional NMR spectroscopy. Application to calmodulin. Biochemistry 29:4659–4667
Ikura M, Spera S, Barbato G, Kay LE, Krinks M, Bax A (1991) Secondary structure and side-chain 1H and 13 C resonance assignments of calmodulin in solution by heteronuclear multidimensional NMR spectroscopy. Biochemistry 30:9216–9228
Jiang L, Baehr W (2010) GCAP1 mutations associated with autosomal dominant cone dystrophy. Adv Exp Med Biol 664:273–282
Koch KW, Helten A (2008) Guanylate cyclase-based signaling in photoreceptors and retina. Signal Transduction in the retina. Taylor and Francis CRC Press, Boca Raton, pp 121–143
Koch KW, Stryer L (1988) Highly cooperative feedback control of retinal rod guanylate cyclase by calcium ions. Nature 334:64–66
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Lim S, Peshenko IV, Dizhoor AM, Ames JB (2009) Effects of Ca2+, Mg2+, and myristoylation on guanylyl cyclase activating protein 1 structure and stability. Biochemistry 48:850–862
Lim S, Dizhoor AM, Ames JB (2014) Structural diversity of neuronal calcium sensor proteins and insights for activation of retinal guanylyl cyclase by GCAP1. Front Mol Neurosci 7:19
Lim S, Peshenko IV, Olshevskaya EV, Dizhoor AM, Ames JB (2016) Structure of guanylyl cyclase activator protein 1 (GCAP1) mutant V77E in a Ca2+-free/Mg2+-bound activator state. J Biol Chem 291:4429–4441
Lim S, Scholten A, Manchala G, Cudia D, Zlomke-Sell SK, Koch KW, Ames JB (2017) Structural characterization of Ferrous Ion binding to retinal guanylate cyclase activator protein 5 from zebrafish photoreceptors. Biochemistry 56:6652–6661
Palczewski K, Subbaraya I, Gorczyca WA, Helekar BS, Ruiz CC, Ohguro H, Huang J, Zhao X, Crabb JW, Johnson RS (1994) Molecular cloning and characterization of retinal photoreceptor guanylyl cyclase-activating protein. Neuron 13:395–404
Payne AM, Downes SM, Bessant DA, Taylor R, Holder GE, Warren MJ, Bird AC, Bhattacharya SS (1998) A mutation in guanylate cyclase activator 1A (GUCA1A) in an autosomal dominant cone dystrophy pedigree mapping to a new locus on chromosome 6p21.1. Hum Mol Genetics 7:273–277
Peshenko IV, Dizhoor AM (2006) Ca2 + and Mg2 + binding properties of GCAP-1. Evidence that Mg2+-bound form is the physiological activator of photoreceptor guanylyl cyclase. J Biol Chem 281:23830–23841
Peshenko IV, Dizhoor AM (2007) Activation and inhibition of photoreceptor guanylyl cyclase by guanylyl cyclase activating protein 1 (GCAP-1): the functional role of Mg2+/Ca2 + exchange in EF-hand domains. J Biol Chem 282:21645–21652
Peshenko IV, Olshevskaya EV, Lim S, Ames JB, Dizhoor AM (2012) Calcium-myristoyl tug. J Biol Chem 287:13972–13984
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Stephen R, Bereta G, Golczak M, Palczewski K, Sousa MC (2007) Stabilizing function for myristoyl group revealed by the crystal structure of a neuronal calcium sensor, guanylate cyclase-activating protein 1. Structure 15:1392–1402
Wishart DS, Sykes BD, Richards FM (1992) The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31:1647–1651
Yap KL, Ames JB, Swindells MB, Ikura M (1999) Diversity of conformational states and changes within the EF-hand protein superfamily. Proteins 37:499–507
Acknowledgements
We thank Derrick Kaseman and Ping Yu for help with NMR experiments performed at the UC Davis NMR Facility.
Funding
Work supported by NIH grants to J.B.A (R01 EY012347) and to the UC Davis NMR Facility (RR11973).
Author information
Authors and Affiliations
Contributions
DC performed experiments, analyzed data and helped write the manuscript. EOA. performed experiments and prepared NMR samples. JBA directed the overall project and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing conflict of interest.
Ethical approval
The experiments comply with the current laws of the United States.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cudia, D., Ahoulou, E.O. & Ames, J.B. Chemical shift assignments of retinal guanylyl cyclase activating protein 5 (GCAP5) with a mutation (R22A) that abolishes dimerization and enhances cyclase activation. Biomol NMR Assign 17, 115–119 (2023). https://doi.org/10.1007/s12104-023-10129-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10129-3