Abstract
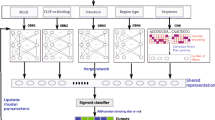
Circular RNAs (circRNAs) participate in the regulation of biological processes by binding to specific proteins and thus influence transcriptional processes. In recent years, circRNAs have become an emerging hotspot in RNA research. Due to powerful learning ability, the various deep learning frameworks have been used to predict the binding sites of RNA-binding protein (RPB) on circRNAs. These methods usually perform only single-level feature extraction of sequence information. However, the feature acquisition may be inadequate for single-level extraction. Generally, the features of deep and shallow layers of neural network can complement each other and are both important for binding site prediction tasks. Based on this concept, we propose a method that combines deep and shallow features, namely CRBP-HFEF. Specifically, features are first extracted and expanded for different levels of network. Then, the expanded deep and shallow features are fused and fed into the classification network, which finally determines whether they are binding sites. Compared to several existing methods, the experimental results on multiple datasets show that the proposed method achieves significant improvements in a number of metrics (with an average AUC of 0.9855). Moreover, much sufficient ablation experiments are also performed to verify the effectiveness of the hierarchical feature expansion strategy.
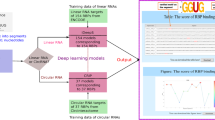
Graphical Abstract















Similar content being viewed by others
Data availability
We used freely available data as described in Methods. The data are available at [https://github.com/wzf171/CRPBsites] and [https://github.com/kavin525zhang/CRIP].
References
Adelman K, Egan E (2017) More uses for genomic junk. Nature 543(7644):183–185. https://doi.org/10.1038/543183a
Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss R et al (2015) Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell 58(5):870–885. https://doi.org/10.1016/j.molcel.2015.03.027
Meng X, Li X, Zhang P, Wang J, Zhou Y, Chen M (2017) Circular RNA: an emerging key player in RNA world. Brief Bioinform 18(4):547–557. https://doi.org/10.1093/bib/bbw045
Zang J, Lu D, Xu A (2020) The interaction of circRNAs and RNA binding proteins: an important part of circRNA maintenance and function. J Neurosci Res 98(1):87–97. https://doi.org/10.1002/jnr.24356
Wang J, Wang L (2019) Deep learning of the back-splicing code for circular RNA formation. Bioinformatics 35(24):5235–5242. https://doi.org/10.1093/bioinformatics/btz382
Jakobi T, Dieterich C (2019) Computational approaches for circular RNA analysis. Wiley Interdiscip Rev 10(3):1528. https://doi.org/10.1002/wrna.1528
Ivanov A, Memczak S, Wyler E, Torti F, Porath HT, Orejuela MR, Piechotta M, Levanon EY, Landthaler M, Dieterich C et al (2015) Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep 10(2):170–177. https://doi.org/10.1016/j.celrep.2014.12.019
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495(7441):333–338. https://doi.org/10.1038/nature11928
Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G et al (2016) Circular RNA profiling reveals an abundant circhipk3 that regulates cell growth by sponging multiple mirnas. Nat Commun 7(1):1–13. https://doi.org/10.1038/ncomms11215
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, Kjems J (2013) Natural RNA circles function as efficient microrna sponges. Nature 495(7441):384–388. https://doi.org/10.1038/nature11993
Pamudurti NR, Bartok O, Jens M, Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E, Perez-Hernandez D, Ramberger E et al (2017) Translation of circRNAs. Mol Cell 66(1):9–21. https://doi.org/10.1016/j.molcel.2017.02.021
Barbagallo D, Caponnetto A, Brex D, Mirabella F, Barbagallo C, Lauretta G, Morrone A, Certo F, Broggi G, Caltabiano R et al (2019) Circsmarca5 regulates vegfa mrna splicing and angiogenesis in glioblastoma multiforme through the binding of srsf1. Cancers 11(2):194. https://doi.org/10.3390/cancers11020194
Okholm TLH, Sathe S, Park SS, Kamstrup AB, Rasmussen AM, Shankar A, Chua ZM, Fristrup N, Nielsen MM, Vang S et al (2020) Transcriptome-wide profiles of circular RNA and RNA-binding protein interactions reveal effects on circular RNA biogenesis and cancer pathway expression. Genome Med 12(1):1–22. https://doi.org/10.1186/s13073-020-00812-8
Zhang H-D, Jiang L-H, Sun D-W, Hou J-C, Ji Z-L (2018) Circrna: a novel type of biomarker for cancer. Breast Cancer 25(1):1–7. https://doi.org/10.1007/s12282-017-0793-9
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, Wu Y-M, Dhanasekaran SM, Engelke CG, Cao X et al (2019) The landscape of circular RNA in cancer. Cell 176(4):869–881. https://doi.org/10.1016/j.cell.2018.12.021
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, Marzluff WF, Sharpless NE (2013) Circular RNAs are abundant, conserved, and associated with alu repeats. RNA 19(2):141–157. https://doi.org/10.1261/rna.035667.112
Thakkar A, Lohiya R (2022) A survey on intrusion detection system: feature selection, model, performance measures, application perspective, challenges, and future research directions. Artif Intell Rev 55(1):453–563. https://doi.org/10.1007/s10462-021-10037-9
Thakkar A, Lohiya R (2021) Attack classification using feature selection techniques: a comparative study. J Ambient Intell Humaniz Comput 12:1249–1266. https://doi.org/10.1007/s12652-020-02167-9
Chaudhari K, Thakkar A (2023) Neural network systems with an integrated coefficient of variation-based feature selection for stock price and trend prediction. Expert Syst Appl. https://doi.org/10.1016/j.eswa.2023.119527
Chaudhari K, Thakkar A (2023) Data fusion with factored quantization for stock trend prediction using neural networks. Inf Process Manag 60(3):103293. https://doi.org/10.1016/j.ipm.2023.103293
Thakkar A, Lohiya R (2023) Fusion of statistical importance for feature selection in deep neural network-based intrusion detection system. Inf Fusion 90:353–363. https://doi.org/10.1016/j.inffus.2022.09.026
Ferre F, Colantoni A, Helmer-Citterich M (2016) Revealing protein-lncrna interaction. Brief Bioinform 17(1):106–116. https://doi.org/10.1093/bib/bbv031
Zhang K, Pan X, Yang Y, Shen H-B (2019) Crip: predicting circRNA-rbp-binding sites using a codon-based encoding and hybrid deep neural networks. RNA 25(12):1604–1615. https://doi.org/10.1261/rna.070565.119
Yang Y, Hou Z, Ma Z, Li X, Wong K-C (2021) icircrbp-dhn: identification of circRNA-rbp interaction sites using deep hierarchical network. Brief Bioinform 22(4):274. https://doi.org/10.1093/bib/bbaa274
Ju Y, Yuan L, Yang Y, Zhao H (2019) Circslnn: identifying rbp-binding sites on circRNAs via sequence labeling neural networks. Front Genet. https://doi.org/10.3389/fgene.2019.01184
Wang Z, Lei X (2021) Prediction of rbp binding sites on circRNAs using an lstm-based deep sequence learning architecture. Brief Bioinform 22(6):342. https://doi.org/10.1093/bib/bbab342
Zhang Q, Zhu L, Huang D-S (2018) High-order convolutional neural network architecture for predicting DNA-protein binding sites. IEEE/ACM Trans Comput Biol Bioinf 16(4):1184–1192. https://doi.org/10.1109/TCBB.2018.2819660
Rong X (2014) word2vec parameter learning explained. arXiv preprint. https://doi.org/10.48550/arXiv.1411.2738
Alam W, Ali SD, Tayara H, to Chong K (2020) A CNN-based RNA n6-methyladenosine site predictor for multiple species using heterogeneous features representation. IEEE Access 8:138203–138209. https://doi.org/10.1109/ACCESS.2020.3002995
Liu B, Gao X, Zhang H (2019) Bioseq-analysis2.0: an updated platform for analyzing DNA, RNA and protein sequences at sequence level and residue level based on machine learning approaches. Nucleic Acids Res 47(20):127. https://doi.org/10.1093/nar/gkz740
Nair AS, Sreenadhan SP (2006) A coding measure scheme employing electron-ion interaction pseudopotential (eiip). Bioinformation 1(6):197
Li Q, Xu L, Li Q, Zhang L et al (2020) Identification and classification of enhancers using dimension reduction technique and recurrent neural network. Comput Math Methods Med. https://doi.org/10.1155/2020/8852258
Liu Q, Chen J, Wang Y, Li S, Jia C, Song J, Li F (2021) Deeptorrent: a deep learning-based approach for predicting DNA n4-methylcytosine sites. Brief Bioinform 22(3):124. https://doi.org/10.1093/bib/bbaa124
Trabelsi A, Chaabane M, Ben-Hur A (2019) Comprehensive evaluation of deep learning architectures for prediction of DNA/RNA sequence binding specificities. Bioinformatics 35(14):269–277. https://doi.org/10.1093/bioinformatics/btz339
Patel R, Patel CI, Thakkar A (2012) Aggregate features approach for texture analysis. Nirma Univ Int Conf Eng (NUiCONE). https://doi.org/10.1109/NUICONE.2012.6493209
Thakkar A, Chaudhari K (2022) Information fusion-based genetic algorithm with long short-term memory for stock price and trend prediction. Appl Soft Comput 128:109428. https://doi.org/10.1016/j.asoc.2022.109428
Yu Y, Si X, Hu C, Zhang J (2019) A review of recurrent neural networks: Lstm cells and network architectures. Neural Comput 31(7):1235–1270. https://doi.org/10.1162/neco_a_01199
Thakkar A, Chaudhari K (2020) Predicting stock trend using an integrated term frequency-inverse document frequency-based feature weight matrix with neural networks. Appl Soft Comput 96:106684. https://doi.org/10.1016/j.asoc.2020.106684
Chen T, Xu R, He Y, Wang X (2017) Improving sentiment analysis via sentence type classification using bilstm-crf and cnn. Expert Syst Appl 72:221–230. https://doi.org/10.1016/j.eswa.2016.10.065
Huang Z, Xu W, Yu K (2015) Bidirectional lstm-crf models for sequence tagging. arXiv preprint. https://doi.org/10.48550/arXiv.1508.01991
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, Kaiser Ł, Polosukhin I (2017) Attention is all you need. Adv Neural Inf Process Syst. https://doi.org/10.48550/arXiv.1706.03762
Funding
This work was supported by a National Natural Science Foundation of China (No. 61972002).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ma, Z., Sun, ZL. & Liu, M. CRBP-HFEF: Prediction of RBP-Binding Sites on circRNAs Based on Hierarchical Feature Expansion and Fusion. Interdiscip Sci Comput Life Sci 15, 465–479 (2023). https://doi.org/10.1007/s12539-023-00572-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-023-00572-0