Abstract
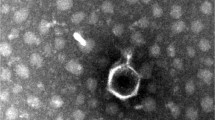
Enterobacter cloacae is a widespread opportunistic pathogen that causes urinary tract infection. The abuse of antibiotics enabled multidrug-resistant strains to spread. Bacteriophage therapy is a naturally, safe, and efficient alternative treatment technology against multi-resistant bacteria. In this study, a virulent phage vB_EclM_Q7622 (Q7622) was isolated from the sewage of Jiangcun poultry market in Guangzhou city. Transmission electron microscopy indicated that Q7622 had an icosahedral head (97.8 ± 5.6 nm in diameter) and a short, contractile tail (113.7 ± 4.5 nm). Its double-stranded DNA genome is composed of 173,871 bp with a GC content of 40.02%. It possesses 297 open reading frames and 9 tRNAs. No known virulence and resistance genes were detected, indicated that phage Q7622 could be used for pathogens prevention and control safely. Comparative genomic and phylogenetic analysis showed that Q7622 was highly similar to the phages vB_EclM_CIP9 and vB_EhoM-IME523. The highest nucleotide similarity between Q7622 and the similar phages in NCBI calculated by pyANI and VIRIDIC were 94.9% and 89.1% with vB_EhoM-IME523 respectively, below 95%. Thus, according to the result of nucleotide similarity calculation results, Q7622 was a novel virulent Enterobacter cloacae phage strain of the genus Kanagawavirus.







Similar content being viewed by others
Data Availability
The whole-genome sequence of vB_EclM_Q7622 was deposited in GenBank as an accession number OL989991.
References
Adriaenssens, E., & Brister, J. R. (2017). How to name and classify your phage: An informal guide. Viruses. https://doi.org/10.3390/v9040070
Ajuebor, J., Buttimer, C., Arroyo-Moreno, S., Chanishvili, N., Gabriel, E. M., O’Mahony, J., McAuliffe, O., Neve, H., Franz, C., & Coffey, A. (2018). Comparison of staphylococcus phage K with close phage relatives commonly employed in phage therapeutics. Antibiotics (Basel). https://doi.org/10.3390/antibiotics7020037
Bailly-Bechet, M., Vergassola, M., & Rocha, E. (2007). Causes for the intriguing presence of tRNAs in phages. Genome Research, 17, 1486–1495. https://doi.org/10.1101/gr.6649807
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A. A., Dvorkin, M., Kulikov, A. S., Lesin, V. M., Nikolenko, S. I., Pham, S., Prjibelski, A. D., Pyshkin, A. V., Sirotkin, A. V., Vyahhi, N., Tesler, G., Alekseyev, M. A., & Pevzner, P. A. (2012). SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology, 19, 455–477. https://doi.org/10.1089/cmb.2012.0021
Bin, J. H., Bolduc, B., Zablocki, O., Kuhn, J. H., Roux, S., Adriaenssens, E. M., Brister, J. R., Kropinski, A. M., Krupovic, M., Lavigne, R., Turner, D., & Sullivan, M. B. (2019). Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nature Biotechnology, 37, 632–639. https://doi.org/10.1038/s41587-019-0100-8
Chan, P. P., Lin, B. Y., Mak, A. J., & Lowe, T. M. (2021). tRNAscan-SE 2.0: Improved detection and functional classification of transfer RNA genes. Nucleic Acids Research, 49, 9077–9096.
D’Souza, A. W., Potter, R. F., Wallace, M., Shupe, A., Patel, S., Sun, X., Gul, D., Kwon, J. H., Andleeb, S., Burnham, C. D., & Dantas, G. (2019). Spatiotemporal dynamics of multidrug resistant bacteria on intensive care unit surfaces. Nature Communications, 10, 4569. https://doi.org/10.1038/s41467-019-12563-1
Jain, C., Rodriguez-R, L. M., Phillippy, A. M., Konstantinidis, K. T., & Aluru, S. (2018). High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nature Communications, 9, 5114. https://doi.org/10.1038/s41467-018-07641-9
Jing, J., Shuwei, W., Dehai, H., Gai, Z., Zhenjiang, L., Songjian, C., Yahui, L., & Zhongquan, W. (2015). Characteristics of Enterobacter cloacae phage Pyg1 and the antibacterial activity of the complement system. Journal of Pathogen Biology, 10(307–11), 324.
Kutter, E. (2009). Phage host range and efficiency of plating. Methods in Molecular Biology, 501, 141–149. https://doi.org/10.1007/978-1-60327-164-6_14
Li, G., Hu, S., Zhong, S., Tsui, W. L., & Chan, S. H. G. (2022). vContact: Private wifi-based IoT contact tracing with virus lifespan. IEEE Internet of Things Journal, 9, 3465–3480. https://doi.org/10.1109/JIOT.2021.3100276
Liu, B., & Pop, M. (2009). ARDB–antibiotic resistance genes database. Nucleic Acids Research, 37, D443–D447. https://doi.org/10.1093/nar/gkn656
Liu, B., Zheng, D., Jin, Q., Chen, L., & Yang, J. (2019). VFDB 2019: A comparative pathogenomic platform with an interactive web interface. Nucleic Acids Research, 47, D687–D692. https://doi.org/10.1093/nar/gky1080
Manohar, P., Tamhankar, A. J., Lundborg, C. S., & Nachimuthu, R. (2019). Therapeutic characterization and efficacy of bacteriophage cocktails Infecting Escherichia coli, Klebsiella pneumoniae, and Enterobacter Species. Frontiers in Microbiology, 10, 574. https://doi.org/10.3389/fmicb.2019.00574
Mezzatesta, M. L., Gona, F., & Stefani, S. (2012). Enterobacter cloacae complex: Clinical impact and emerging antibiotic resistance. Future Microbiology, 7, 887–902. https://doi.org/10.2217/fmb.12.61
Minh, B. Q., Schmidt, H. A., Chernomor, O., Schrempf, D., Woodhams, M. D., von Haeseler, A., & Lanfear, R. (2020). IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution, 37, 1530–1534.
Moraru, C., Varsani, A., & Kropinski, A. M. (2020). VIRIDIC-a novel tool to calculate the intergenomic similarities of prokaryote-infecting viruses. Viruses. https://doi.org/10.3390/v12111268
Nair, A., Vyawahare, R., & Khairnar, K. (2022). Characterization of a novel, biofilm dispersing, lytic bacteriophage against drug-resistant Enterobacter cloacae. Journal of Applied Microbiology, 132, 2721–2732. https://doi.org/10.1111/jam.15420
Otasek, D., Morris, J. H., Boucas, J., Pico, A. R., & Demchak, B. (2019). Cytoscape automation: Empowering workflow-based network analysis. Genome Biology, 20, 185. https://doi.org/10.1186/s13059-019-1758-4
Seemann, T. (2014). Prokka: Rapid prokaryotic genome annotation. Bioinformatics, 30, 2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Shi, Y., Chen, Y., Yang, Z., Zhang, Y., You, B., Liu, X., Chen, P., Liu, M., Zhang, C., Luo, X., Chen, Y., Yuan, Z., Chen, J., Gong, Y., & Peng, Y. (2020). Characterization and genome sequencing of a novel T7-like lytic phage, kpssk3, infecting carbapenem-resistant Klebsiella pneumoniae. Archives of Virology, 165, 97–104. https://doi.org/10.1007/s00705-019-04447-y
Sullivan, M. J., Petty, N. K., & Beatson, S. A. (2011). Easyfig: A genome comparison visualizer. Bioinformatics, 27, 1009–1010. https://doi.org/10.1093/bioinformatics/btr039
Tolstoy, I., Kropinski, A. M., & Brister, J. R. (2018). Bacteriophage taxonomy: An evolving discipline. Methods in Molecular Biology, 1693, 57–71. https://doi.org/10.1007/978-1-4939-7395-8_6
Wang, D., Shi, H. Y., Jing-Hua, L. I., Jing-Dan, Y. U., Huang, J. F., & Sun, Y. B. (2014). Experimental phage therapy against Enterobacter cloacae abscesses in rabbit model. Chinese Journal of Microecology. https://doi.org/10.13381/j.cnki.cjm.201401004
Wang, K., Tamayo, M. G., Penner, T. V., Cook, B., Court, D. A., & Theriault, S. S. (2020). Characterization of the enterobacter phage vB_EclM_CIP9. Microbiology Resource Announcements. https://doi.org/10.1128/MRA.01600-19
Wang, P., Wen, Z., Li, B., Zeng, Z., & Wang, X. (2016). Complete genome sequence of Vibrio alginolyticus ATCC 33787(T) isolated from seawater with three native megaplasmids. Marine Genomics, 28, 45–47. https://doi.org/10.1016/j.margen.2016.05.003
Wilson, B. M., El, C. N., Patel, S., Saade, E., Donskey, C. J., Bonomo, R. A., & Perez, F. (2017). Carbapenem-resistant Enterobacter cloacae in patients from the US Veterans health administration, 2006–2015. Emerging Infectious Diseases, 23, 878–880. https://doi.org/10.3201/eid2305.162034
Yang, M., Chen, H., Huang, Q., Xie, Z., Liu, Z., Zhang, J., Ding, Y., Chen, M., Xue, L., Wu, Q., & Wang, J. (2022). Characterization of the novel phage vB_VpaP_FE11 and Its Potential role in controlling vibrio parahaemolyticus biofilms. Viruses. https://doi.org/10.3390/v14020264
Yang, M., Hanfang, C., Sheng, G., Shilin, T., Zhuanbei, X., Zhang, J., Qingping, W., & Zhiyuan, T. (2022). Characterization and genome analysis of a novel Vibrio parahaemolyticus phage vB_VpP_DE17. Virus Research. https://doi.org/10.1016/j.virusres.2021.198580
Yang, M., Liang, Y., Huang, S., Zhang, J., Wang, J., Chen, H., Ye, Y., Gao, X., Wu, Q., & Tan, Z. (2020). Isolation and characterization of the novel phages vB_VpS_BA3 and vB_VpS_CA8 for lysing vibrio parahaemolyticus. Frontiers in Microbiology, 11, 259. https://doi.org/10.3389/fmicb.2020.00259
Yang, M., Liang, Y., Su, R., Chen, H., Wang, J., Zhang, J., Ding, Y., Kong, L., Zeng, H., Xue, L., Wu, H., & Wu, Q. (2019). Genome characterization of the novel lytic Vibrio parahaemolyticus phage vB_VpP_BA6. Archives of Virology, 164, 2627–2630. https://doi.org/10.1007/s00705-019-04351-5
Acknowledgements
The authors thank all the members of the laboratory for helping with the experiment.
Funding
The authors are grateful to the Guangzhou Key Research and Development Program (SL2022B03J01243), Guangdong Major Project of Basic and Applied Basic Research (2020B0301030005), National Natural Science Foundation of China (32202194, 31730070), Provincial Key Laboratory of Microbial Safety and Health (2020B121201009- PKLMSH004-2021) and National Key Research and Development Program (2021YFA0910200) for financial supports.
Author information
Authors and Affiliations
Contributions
Conceptualization, ST, SH and MY; methodology, ST, SH and MY; software, ST, and SH; writing—original draft preparation, ST; writing—review and editing, ST, SH HC and MY; visualization, ST, SH and ZL; project administration, MY; funding acquisition, MY. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tan, S., Huang, S., Liu, Z. et al. Genome Characterization of the Novel Lytic Enterobacter cloacae Phage vB_EclM_Q7622. Food Environ Virol 15, 236–245 (2023). https://doi.org/10.1007/s12560-023-09558-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-023-09558-z