Abstract
Background
Alpha-fetoprotein (AFP) has been used as a biomarker for early diagnosis of hepatocellular carcinoma (HCC) for a long time. Aptamer is a type of material that is receiving attention as a new alternative in the diagnosis and treatment of diseases.
Objective
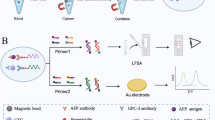
We applied aptamer to AFP detection and utilized in silico analysis techniques. Here, we describe the selection of aptamers that can bind to AFP as well as in vitro and in silico validation of the binding site. We also developed an aptamer-based detection platform called hybrid-aptablotting sandwich assay (Hybrid-ABSA) and a point-of-care AFP detection kit.
Results
To construct the AFP detection platform, we screened a total of 18 candidate AFP-binding aptamers and identified a pair of aptamers without non-overlapping-binding sites using dot blotting. We validated results of dot blotting using in silico 3D binding structure docking simulation analysis. The aptamer-based Hybris-ABSA for AFP detection using this pair of aptamers demonstrated superior detection performance to antibody-based ELISA detection methods. Furthermore, we confirmed that AFP could be detected using the developed point-of-care lateral flow In Vitro Diagnostic (IVD) kit.
Conclusion
Results of this study demonstrate that aptamer-based disease diagnostic technology has potential of replace conventional diagnostic methods. This study also confirms that in silico techniques have sufficient utility in biological assays.






Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ahn JY et al (2018) Surface plasmon resonance aptamer biosensor for discriminating pathogenic bacteria Vibrio parahaemolyticus. J Nanosci Nanotechnol 18:1599–1605. https://doi.org/10.1166/jnn.2018.14212
AlphaFold and Beyond (2023) Nat Methods 20:163. https://doi.org/10.1038/s41592-023-01790-6
Bairoch A, Boeckmann B, Ferro S, Gasteiger E (2004) Swiss-Prot: juggling between evolution and stability. Brief Bioinform 5:39–55. https://doi.org/10.1093/bib/5.1.39
Balogh J et al (2016) Hepatocellular carcinoma: a review. J Hepatocell Carcinoma 3:41–53
Bei R, Mizejewski GJ (2011) Alpha fetoprotein is more than a hepatocellular cancer biomarker: from spontaneous immune response in cancer patients to the development of an AFP-based cancer vaccine. Curr Mol Med 11:564–581. https://doi.org/10.2174/156652411800615162
Cassinotto C, Aubé C, Dohan A (2017) Diagnosis of hepatocellular carcinoma: an update on international guidelines. Diagn Interv Imaging 98:379–391
Debruyne EN, Delanghe JR (2008) Diagnosing and monitoring hepatocellular carcinoma with alpha-fetoprotein: new aspects and applications. Clin Chim Acta 395:19–26
El-Serag HB et al (2023) Prevention of hepatocellular carcinoma (HCC). White paper of the Texas collaborative center for hepatocellular cancer (TeCH) Multi-stakeholder conference. Clin Gastroenterol Hepatol. https://doi.org/10.1016/j.cgh.2023.03.029
Guilbert C, James TL (2008) Docking to RNA via root-mean-square-deviation-driven energy minimization with flexible ligands and flexible targets. J Chem Inf Model 48:1257–1268. https://doi.org/10.1021/ci8000327
Jakab S S (2010) Clinical hepatology—principles and practice of hepatobiliary diseases. Report No. 0192-0790, (LWW)
Johnson PJ (2001) The role of serum alpha-fetoprotein estimation in the diagnosis and management of hepatocellular carcinoma. Clin Liver Dis 5:145–159
Kim M et al (2009) Arsenic removal from Vietnamese groundwater using the arsenic-binding DNA aptamer. Environ Sci Technol 43:9335–9340. https://doi.org/10.1021/es902407g
Kim YH et al (2011) An RNA aptamer that specifically binds pancreatic adenocarcinoma up-regulated factor inhibits migration and growth of pancreatic cancer cells. Cancer Lett 313:76–83. https://doi.org/10.1016/j.canlet.2011.08.027
Kokudo N et al (2019) Clinical practice guidelines for hepatocellular carcinoma: the Japan Society of Hepatology 2017 (4th JSH-HCC guidelines) 2019 update. Hepatol Res 49:1109–1113
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786. https://doi.org/10.1021/ci200227u
Lee KA et al (2015) Aptamer-based sandwich assay and its clinical outlooks for detecting lipocalin-2 in hepatocellular carcinoma (HCC). Sci Rep 5:10897. https://doi.org/10.1038/srep10897
Lee J-P et al (2021) Structure-based molecular docking approach for identifying S-formylglutathione hydrolase from Sphingobium chungbukense. Toxicol Environ Heal Sci 13:407–416
Marrero JA, Feng Z (2010) Alpha-fetoprotein in early hepatocellular carcinoma. Gastroenterology 138:400–401. https://doi.org/10.1053/j.gastro.2009.08.076
National Cancer Institute (2022) https://www.cancer.gov/
National Institutes of Health (2012) PubMed, C &. Thomson, G. S. Karger AG, Basel
Oh IH et al (2020) Docking simulation and sandwich assay for aptamer-based botulinum neurotoxin type C detection. Biosensors (basel). https://doi.org/10.3390/bios10080098
Park H, Park JY (2013) Clinical significance of AFP and PIVKA-II responses for monitoring treatment outcomes and predicting prognosis in patients with hepatocellular carcinoma. BioMed Res Int. https://doi.org/10.1155/2013/310427
Perlinska AP et al (2023) AlphaFold predicts novel human proteins with knots. Protein Sci 32:e4631. https://doi.org/10.1002/pro.4631
Rabal O et al (2016) In silico aptamer docking studies: from a retrospective validation to a prospective case study-TIM3 aptamers binding. Mol Ther Nucleic Acids 5:e376. https://doi.org/10.1038/mtna.2016.84
Sekhon SS et al (2017a) Aptabody-aptatope interactions in aptablotting assays. Nanoscale 9:7464–7475. https://doi.org/10.1039/c7nr01827d
Sekhon SS et al (2017b) Defining the copper binding aptamotif and aptamer integrated recovery platform (AIRP). Nanoscale 9:2883–2894. https://doi.org/10.1039/c6nr09408b
Sekhon SS et al (2018) Aptasensors for pesticide detection. Toxicol Environ Heal Sci 10:229–236
Sekhon SS et al (2022) Cyclophilin A-mediated mitigation of coronavirus SARS-CoV-2. Bioeng Transl Med 8:e10436. https://doi.org/10.1002/btm2.10436
Shin WR et al (2018a) Aptamer-based paper strip sensor for detecting Vibrio fischeri. ACS Comb Sci 20:261–268. https://doi.org/10.1021/acscombsci.7b00190
Shin WR et al (2018b) Aptamer-based pathogen monitoring for Salmonella enterica ser. Typhimurium. J Biomed Nanotechnol 14:1992–2002. https://doi.org/10.1166/jbn.2018.2634
Shin W-R et al (2020) Aptamer-linked immobilized sorbent assay for detecting GMO marker, phosphinothricin acetyltransferase (PAT). Mol Cell Toxicol 16:253–261
Shin WR et al (2022) Structure based innovative approach to analyze aptaprobe-GPC3 complexes in hepatocellular carcinoma. J Nanobiotechnology 20:204. https://doi.org/10.1186/s12951-022-01391-z
Siegel AB, Zhu AX (2009) Metabolic syndrome and hepatocellular carcinoma: two growing epidemics with a potential link. Cancer 115:5651–5661
Song MS et al (2017) Detecting and discriminating Shigella sonnei using an aptamer-based fluorescent biosensor platform. Molecules. https://doi.org/10.3390/molecules22050825
Song MS et al (2018) Aptamer-immobilized surface plasmon resonance biosensor for rapid and sensitive determination of virulence determinant. J Nanosci Nanotechnol 18:3095–3101. https://doi.org/10.1166/jnn.2018.14697
Stoltenburg R, Reinemann C, Strehlitz B (2007) SELEX—A (r) evolutionary method to generate high-affinity nucleic acid ligands. Biomol Eng 24:381–403
Sung H et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J Clin 71:209–249
Toyoda H et al (2014) Changes in highly sensitive alpha-fetoprotein for the prediction of the outcome in patients with hepatocellular carcinoma after hepatectomy. Cancer Med 3:643–651. https://doi.org/10.1002/cam4.218
Trevisani F et al (2001) Serum α-fetoprotein for diagnosis of hepatocellular carcinoma in patients with chronic liver disease: influence of HBsAg and anti-HCV status. J Hepatol 34:570–575
Um HJ, Kim M, Lee SH, Kim YH (2012) Preventing the formation of positive transcription elongation factor b by human cyclin T1-binding RNA aptamer for anti-HIV transcription. AIDS 26:1599–1605. https://doi.org/10.1097/QAD.0b013e3283554f7d
UniProt C (2023) UniProt: the universal protein knowledgebase in 2023. Nucleic Acids Res 51:D523–D531. https://doi.org/10.1093/nar/gkac1052
World Health Organization (2020) WHO report on cancer: setting priorities, investing wisely and providing care for all. Report No. 9240001298
Zhang X-F et al (2009) Prognostic factors after liver resection for hepatocellular carcinoma with hepatitis B virus-related cirrhosis: the surgeon’s role in survival. Eur J Surg Oncol (EJSO) 35:622–628
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415. https://doi.org/10.1093/nar/gkg595
Acknowledgements
This research was supported by Chungbuk National University Korea National University Development Project (2021).
Author information
Authors and Affiliations
Contributions
W-RS and D-YP: conceived and designed the study; H-JU, GA, and W-RS: performed data analysis; D-YP: performed the biological methodology and acquisition of data; SYK, J-YA, and Y-HK: drafted the manuscript (assign co-first authors order according to workload). All the authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
Woo-Ri Shin declares that she has no conflict of interest. Dae-Young Park declares that he has no conflict of interest. Hyun-Ju Um declares that she has no conflict of interest. Gna Ahn declares that she has no conflict of interest. Sang Yong Kim declares that he has no conflict of interest. Ji-Young Ahn declares that she has no conflict of interest. Yang-Hoon Kim declares that he has no conflict of interest.
Ethical approval
Clinical samples were collected from the Institute of Clinical Medicine, Department of Internal Medicine, Chonbuk National University Hospital, South Korea. All subjects provided their written informed consent prior to participating in this study. The Ethics Committee and IRB (Institutional Review Board) of Chonbuk National University Hospital approved all experimental procedures.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shin, WR., Park, DY., Um, HJ. et al. 3D structural analysis of aptamer and diagnostic platforms for detecting hepatocellular carcinoma. Mol. Cell. Toxicol. 19, 621–634 (2023). https://doi.org/10.1007/s13273-023-00369-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13273-023-00369-8