Abstract
Three new halimane furanoditerpenoids (1–3) and three new clerodane furanoditerpenoids (4–6), along with seven known terpenoids including four pimarane diterpenoids (7–10) and three norisoprenoids (11–13) were isolated from the 95% EtOH extracts of the plants of Croton cnidophyllus. The 2D structures including absolute configuration of new furanoditerpenoids (1–6) were elucidated by analysis of their HRMS and NMR data as well as comparison of experimental and calculated ECD curves. Bioassay revealed that two compounds (8 and 9) possessed certain inhibitory effects against NO production stimulated by LPS, with IC50 values of 19.00 ± 1.76 and 21.61 ± 1.11 μM, respectively.
Graphical Abstract

Similar content being viewed by others
1 Introduction
Many plants of the genus Croton (Euphorbiaceae) such as C. tiglium, C. crassifolius, and C. kongensis are well-known traditional Chinese medicines, which have been used to treat stomachache, sore throat, rheumatism, and headache [1, 2]. Previous chemical investigation on some species of Croton revealed various types of diterpenoids including tiglianes [3], clerodanes [4], halimanes [1], labdanes [5], kauranes [6], casbanes [7], abietanes [6], isopimaranes [8], cembranes [9], and phytanes [10]. Some of these diterpenoids from the genus Croton exhibited diverse biological functions, such as cytotoxity [11], antiviral [12, 13], anti-plasmodial [14, 15], anti-microbial [15, 16], anti-inflammatory [1], and hypoglycemic activities [17]. The interesting structures of diterpenoids as well as their important bioactivities have made natural product chemists increasingly interested in studying plants of this genus.
Croton cnidophyllus Radcliffe-Smith and Govaerts (synonym: Croton urticifolius) are shrubs, ranging from 1 to 2 m tall, mainly distributed in Guangxi, South Guizhou, and South Yunnan of China [18]. To our knowledge, there are currently no reports about the chemical constituents and bioactivities of C. cnidophyllus. In our continuous research aiming at the discovery of novel structures with biologically active diterpenoids from Euphorbiaceae [1, 19,20,21], the EtOAc fraction of the alcohol extracts of C. cnidophyllus was subjected to repeated column chromatography. A total of 13 compounds (1–13) were isolated and identified from C. cnidophyllus for the first time (Fig. 1). Of them, compounds 1–6 are new natural products. In addition, their inhibitory effects against the production of nitric oxide (NO) were evaluated in the LPS-induced RAW 264.7 cell model. Hence, the purification, structural elucidation of all terpenoids together with their bioactive assay are described.
2 Results and discussion
Crodophylloid A (1) (Fig. 1) was acquired as colorless oil, and its molecular formula of C20H26O5 was determined by an HR-ESI-MS ion at m/z 369.1676 [M + Na]+ (calcd for 369.1672), corresponding to eight DOUs (degrees of unsaturation). The 1H NMR data of 1 (Table 1) exhibited resonances for one β-substituted furan [δH 6.76 (1H, dd, J = 1.9 and 0.8 Hz), 7.46 (1H, t, J = 1.6 Hz), and 8.11 (1H, br s)], three methyls [δH 0.83 (3H, d, J = 6.7 Hz), 0.96 (3H, s), and 1.24 (3H, s)], one isolated methylene [δH 2.89 (1H, d, J = 19.0 Hz) and 3.41 (1H, d, J = 19.0 Hz)], one oxygenated methine [δH 3.47 (1H, br s)], and other aliphatic multiplets. According to the DEPT and HSQC spectra, 20 carbon signals observed in 13C NMR spectrum were assigned as one conjugated ketocarbonyl (δC 192.6), one ester carbonyl (δC 177.8), a substituted furan moiety [δC 108.5 (CH), 127.6 (C), 144.6 (CH), and 147.6 (CH)], three quaternary carbons including an oxygenated one, three methines including an oxygenated one, five methylenes, and three methyls. The collective information suggested that compound 1 was a furanoditerpenoid.
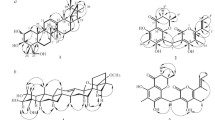
The 2D structure of 1 was determined on the basis of its 2D NMR spectra involving 1H–1H COSY and HMBC. The presence of fragments a and b as marked in Fig. 2 could be deduced by the 1H–1H COSY cross-peaks, which were linked to quaternary carbons C-4, C-5, and C-9 to constructed rings A and B, a 6/6 bicyclic skeleton with two methyls at C-4, by the HMBC correlations of H3-19/18 to C-5, C-4, and C-3, H2-6 to C-10, C-5, and C-4, H-8 to C-10 and C-9, and H3-17 to C-9. The 2-(furan-3-yl)-2-oxoethyl moiety could be confirmed by analyzing its 1D NMR data with the reported furanoditerpenoids crohalifuranes C–E [1], and its location at C-9 of ring B was proved by the HMBC correlations of H2-11 to C-10, C-9, and C-8. Besides, the ester carbonyl group (C-20) was also linked to C-9 as supported by the HMBC correlations from H2-11 and H-10 to C-20. The remaining one DOU (seven of the eight DOUs were represented by the ketocarbonyl, the ester carbonyl, the β-substituted furan moiety, and rings A and B) and the obvious downfield-shifted carbon chemical shift of C-5 (δC 89.5) manifested that there was a five-membered ring lactone formed between C-5 and C-20. Thus, the planar structure of 1 featured with a 5,20-γ-lactone moiety was elucidated.
The relative stereochemistry of compound 1 was assigned through its NOESY experiment. The NOE cross-peaks of H-10/H-8 and H-6 and H3-19/H-6 suggested that H-10, H-8, Me-19, and H-6 occupied the axial position of the chair conformer of the A- and B-rings, and these groups were arbitrarily assigned to be β-oriented (Fig. 3), while the lactone bridge was occupied the α-orientation. The NOE cross-peaks of Me-18 (19)/H-3 assigned H-3 as equatorial β-orientation. Furthermore, the experimental ECD data of 1 displayed a negative Cotton effect at approximately 217 nm, which was consistent with other halimane derivatives isolated from Croton [1]. Therefore, the absolute configuration 3R,5S,8R,9R,10R was assigned to compound 1, and it was the first example of halimane diterpenoid with a 5,20-γ-lactone moiety.
Crodophylloid B (2) (Fig. 1) had an HR-ESI-MS positive ion peak at m/z 383.1826 [M + Na]+ (calcd 383.1829), suggesting a molecular formula C21H28O5 for 2. The 1H and 13C NMR data (Table 1) showed signals for one isolated methylene, a β-substituted furan, and a conjugated ketocarbonyl, which were undoubtedly determined to be a 2-(furan-3-yl)-2-oxoethyl group with reference to 1D NMR data of 1. In addition, signals for an ester carbonyl, a tetrasubstituted double bond, a methyl doublet, a methoxy group, four methylenes, two methines including an oxygenated one, and two sp3 quaternary carbons were observed in its 1D NMR spectra. Comparison of these data with those of 1 suggested that 2 was similar to 1, and the main differences was that compound 2 had additional tetrasubstituted double bond and methoxy group. The locations of the double bond (Δ5,10) and methoxy group (20-OMe) were confirmed by the HMBC correlations of H2-6 to C-5, H2-1 to C-10, and 20-OMe to C-20. Its relative stereochemistry was identical to 1 by analysis of the NOE correlations of 2 (Fig. 3). The similar tendency shown in the ECD spectrum of 2 with that of crohalifurane D [1] indicated compound 2 had the (3R,8R,9R) absolute configuration. Therefore, compound 2 was elucidated as shown in Fig. 1.
Crodophylloid C (3) (Fig. 1) was deduced to possess a molecular formula of C21H28O5 based on the HR-ESI-MS data. The 1H and 13C NMR data of 3 (Table 1) had similar signals to 2, except for the resonances for a double bond. The appearance of a trisubstituted double bond signals in 3 instead of a tetrasubstituted one in 2 suggested the double bonds were in different positions. The double bond was switched to Δ5 in 3 confirmed through the observed 1H–1H COSY cross-peak of H-6/H2-7 together with the HMBC correlations of H-6 and H3-19 to C-5 (Fig. 2). The relative configuration of 3 was confirmed by analyzing its NOE correlations (Fig. 3). The ECD curve of 3 matched well with that of crodophylloid C (2), indicating that a 3R,8R,9R,10S absolute configuration was proposed for 3.
The molecular formula of crodophylloid D (4) (Fig. 1) was deduced to be C21H28O5 based on its HR-ESI-MS data together with its 13C NMR spectrum. By comparison of the 1D NMR data of 4 (Table 2) with those of the known clerodane diterpenoid 3,12-dioxo-15,16-epoxy-cleroda-13(16),14-dien-9-al [4], it was found that a methoxyl group [δC 51.3 (CH3) and 174.2 (C); δH 3.67 (3H, s)] had replaced the aldehyde group (C-20) in the known compound. This was further proven by the HMBC correlations of H2-11 and 20-OMe to C-20 (Fig. 2).
By comparing their 1H and 13C NMR data and NOESY correlations, the relative stereochemistry of C-5, C-8, C-9, and C-10 in 4 were determined to be the same as those of 3,12-dioxo-15,16-epoxy-cleroda-13(16),14-dien-9-al. While H-4 was determined as β-orientation by the NOESY signals of H-4/H-10 (Fig. 3). The absolute configuration of 4 was assigned via ECD calculation. The ECD curves of this pair of enantiomers, (4S,5R,8R,9R,10S)-4 and (4R,5S,8S,9S,10R)-4 were simulated by using the TDDFT method. By analysis of the experimental and calculated ECD curves (Fig. 4), the absolute configuration of 4 was determined as 4S,5R,8R,9R,10S.
Crodophylloid E (5) (Fig. 1) possessed the molecular formula C22H32O6 based on the analysis of the HR-ESI-MS data and 13C NMR spectrum of 5. By comparing its 1H and 13C NMR spectra with those of the known compound 3α,4β-dihydroxy-15,16-epoxy-12-oxo-cleroda-13(16),14-diene [22], 5 showed an ester carbonyl (δC 175.2) instead of the aldehyde group (C-20) of the known one together with the presence of two extra methoxyls (δC 50.0 and 51.0). These were supported by the HMBC correlations from 4-OMe (δH 3.18) to C-4 (δC 79.5) and from 20-OMe (δH 3.67) to C-20 (δC 175.2) (Fig. 2). The relative stereochemistry of 5 was ascertained the same as the known one [22] on the basis of its NOE correlations (Fig. 3). The 3R,4R,5R,8R,9R,10S absolute configuration of 5 was confirmed by comparing its ECD curve with that of 4, which showed similar tendency.
Compound 6 (Fig. 1) displayed an HR-ESI-MS ion at m/z 383.1829 [M + Na]+ (calcd 383.1829), suggesting the molecular formula C21H28O5. Comparison of the 1H and 13C NMR data (Table 1) of 6 with those of crodophylloid E (5) suggested that 6 was similar with 5, with the differences being a Δ5,18 exocyclic double bond in 6 instead of the oxygenated quaternary carbon with a methoxy group and the methyl in 5. These changes were further confirmed via the apparent downfield-shifted carbon chemical shifts of C-4 and C-18 and the HMBC correlations from H2-18 to C-4, C-3, and C-5. The relative configurations of C-3, C-5, C-8, C-9, and C-10 in 6 were identical to 5 by the NOESY correlations as shown in Fig. 3. Therefore, compound 6 was determined as illustrated in Fig. 1 and named as crodophylloid F.
Seven known compounds were determined as ent-3α-hydroxypimara-8(14),15-dien-12-one (7) [23], 12-hydroxy-13-methylpodocarpa-8,11,13-trien-3-one (8) [24], epi-isojatrogrossidione (9) [25], 12-hydroxy-13-methyl-ent-podocarp-6,8,11,13-tetraen-3-one (10) [26], 6-hydroxy-megastigm-7-en-3,9-dione (11) [27], (6S,7E)-6-hydroxy-4,7-megastigmadien-3,9-dione (12) [28], and (3S,5R,6S,7E)-5,6-epoxy-3-hydroxy-7-megastigmen-9-one (13) [27] based on their identical NMR data with the reported.
In the LPS-induced RAW 264.7 inflammatory cell model, the inhibitory effects of all isolates on NO production were tested by the Griess assay. Initially, all the tested compounds at a concentration of 50 μM had no cytotoxicity to RAW 264.7 cells. In comparison to the positive control (Quercetin, IC50 = 14.55 ± 0.8 μM), compounds 8 and 9 had certain inhibitory activities (IC50 = 19.0 ± 1.8 and 21.6 ± 1.1 μM, respectively), and the remaining terpenoids were inactive (IC50 > 50 μM).
3 Experimental section
3.1 General experimental procedures
For details see Additional file 1: S1.1.
3.2 Plant material
The plant material was obtained in June 2020 from Xishuangbanna of Yunnan Province, P. R. China, and it was authenticated to be Croton cnidophyllus Radcliffe-Smith and Govaerts by Dr. G.H. Tang. A voucher specimen (Accession No.: KCC201907) has been deposited in the School of Pharmaceutical Sciences, SYSU (Sun Yat-sen University).
3.3 Extraction and isolation
The air-dried powder of plant material (15 kg) was soaked in 95% EtOH (50 L × 3) at room temperature for a month. After removing the solvents under vacuum, 800 g of black crude extract was obtained, which was then suspended in water (3 L) and followed by partitioned with ethyl acetate (EtOAc, 3 L × 5). The obtained EtOAc fraction (315 g) was firstly separated over a silica gel column eluted with a gradient of petroleum ether (PE)/EtOAc (50:1 → 1:1) to obtained Frs. Ι–V.
Compounds 1 (1.7 mg) and 2 (2.4 mg) were obtained from Fr. Ι by various column chromatography including silica gel column, Sephadex LH-20 column, and HPLC. 8 (25 mg) and 4 (5 mg, tR = 21.5 min) were purified from Fr. ΙΙ, 3 (3 mg), 5 (1 mg), 6 (5 mg), 7 (11 mg), and 10 (30 mg) from Fr. III, 9 (2 mg) and 11 (8 mg) from Fr. IV, and 12 (8 mg) and 13 (3 mg) from Fr. V by similar separation methods. The detailed separation process can be found in Additional file 1: S1.2.
3.4 Spectroscopic data of compounds
3.4.1 Crodophylloid A (1)
Colorless oil; [α] 25D + 12.9 (c 0.09, MeCN); UV (MeCN) λmax (log ε) 190 (3.70) nm; ECD (c 5.5 × 10–4 M, MeCN) λmax (Δε) 190 (1.27), 217 (1.42) nm; IR (KBr) νmax 3444, 2954, 2924, 2854, 1751, 1678, 1156 cm–1; 1D NMR data see Table 1; positive ion HR-ESI-MS m/z 369.1676 [M + Na]+ (calcd for C20H26O5 Na+, 369.1672).
3.4.2 Crodophylloid B (2)
Colorless oil; [α] 25D + 65.6 (c 0.09, MeCN); UV (MeCN) λmax (log ε) 190 (3.71) nm; ECD (c 5.6 × 10–4 M, MeCN) λmax (Δε) 190 (1.17), 200 (1.68), 225 (5.80); IR (KBr) νmax 3423, 2954, 2924, 2854, 1718 cm−1; 1D NMR data see Table 1; positive ion HR-ESI-MS m/z 383.1830 [M + Na]+ (calcd for C21H28O5Na+, 383.1829).
3.4.3 Crodophylloid C (3)
Colorless oil; [α] 25D + 11.8 (c 0.12, MeCN); UV (MeCN) λmax (log ε) 190 (4.05) nm; ECD (c 5.0 × 10−4 M, MeCN) λmax (Δε) 192 (3.16), 208 (1.94), 226 (0.25); IR (KBr) νmax 3443, 2925, 1714, 1671, 1156 cm−1; 1D NMR data see Table 1; positive ion HR-ESI-MS m/z 383.1829 [M + Na]+ (calcd for C21H28O5Na+, 383.1829).
3.4.4 Crodophylloid D (4)
Colorless oil; [α] 25D + 40.3 (c 0.07, MeCN); UV (MeCN) λmax (log ε) 190 (3.83), 250 (3.25) nm; ECD (c 4.4 × 10−4 M, MeCN) λmax (Δε) 191 (2.25), 220 (0.64), 291 (0.79); IR (KBr) νmax 3441, 2953, 2925, 2854, 1712, 1156 cm−1; 1D NMR data see Table 2; positive ion HR-ESI-MS m/z 383.1830 [M + Na]+ (calcd for C21H28O5 Na+, 383.1829).
3.4.5 Crodophylloid E (5)
Colorless oil; [α] 25D + 7.6 (c 0.12, MeCN); UV (MeCN) λmax (log ε) 190 (3.32) nm; ECD (c 8.9 × 10−4 M, MeCN) λmax (Δε) 194 (+ 0.54), 220 (+ 0.42); IR (KBr) νmax 3445, 2917, 2849, 1708, 1462, 1155 cm−1; 1D NMR data see Table 2; positive ion HR-ESI-MS m/z 415.2092 [M + Na]+ (calcd for C22H32O6Na+, 415.2091).
3.4.6 Crodophylloid F (6)
Colorless oil; [α] 25D + 16.8 (c 0.11, MeCN); UV (MeCN) λmax (log ε) 190 (3.95); ECD (c 5.5 × 10−4 M, MeCN) λmax (Δε) 191 (0.91), 200 (2.87), 225 (0.05); IR (KBr) νmax 3443, 2923, 1723, 1155 cm−1; 1D NMR data see Table 2; positive ion HR-ESI-MS m/z 383.1829 [M + Na]+ (calcd for C21H28O5Na+, 383.1829).
3.5 ECD calculations
For details of the quantum chemical ECD calculation of 4, see Additional file 1: S1.5.
3.6 Anti-inflammatory activity assay
RAW 264.7 cells were obtained from Southern Medical University Cell Bank (Guangzhou, China). The cell viability and the NO concentration were evaluated by the MTT assay and the Griess reaction, respectively (for details see Additional file 1: S1.3 and S1.4).
Availability of data and materials
All data generated and analyzed during this study are included in this published article and its Additional file 1.
References
Wang R, Fan RZ, Ni FQ, Sang J, Xie XL, Luo SY, Tang GH, Yin S. 19-nor-, 20-nor-, and tetranor-halimane-type furanoditerpenoids from Croton crassifolius. J Nat Prod. 2020;83:255–67.
Hu R, Huang JL, Yuan FY, Wei X, Zou MF, Tang GH, Li W, Yin S. Crotonianoids A-C, three unusual tigliane diterpenoids from the seeds of Croton tiglium and their anti-prostate cancer activity. J Org Chem. 2022;87:9301–6.
Cui JJ, Ji KL, Liu HC, Zhou B, Liu QF, Xu CH, Ding J, Zhao JX, Yue JM. Cytotoxic tigliane diterpenoids from Croton damayeshu. J Nat Prod. 2019;82:1550–7.
Krebs HC, Ramiarantsoa H. Clerodane diterpenes of Croton hovarum. Phytochemistry. 1997;45:379–81.
Qi JJ, Zhou JS, Zhang Y, Fan YY, Zhou B, Liu HC, Zhao JX, Yue JM. Sublyratins A-O, labdane-type diterpenoids from Croton sublyratus. J Nat Prod. 2021;84:2971–80.
Santos HS, Barros FWA, Albuquerque MRJR, Bandeira PN, Pessoa C, Braz-Filho R, Monte FJQ, Leal-Cardoso JH, Lemos TLG. Cytotoxic diterpenoids from Croton argyrophylloides. J Nat Prod. 2009;72:1884–7.
Moura VLA, Monte FJO, Filho RB. A new casbane-type diterpenoid from Croton nepetaefolius. J Nat Prod. 1990;53:1566–71.
Baccelli C, Navarro I, Block S, Abad A, Morel N, Quetin-Leclercq J. Vasorelaxant activity of diterpenes from Croton zambesicus and synthetic trachylobanes and their structure-activity relationships. J Nat Prod. 2007;70:910–7.
Roengsumran S, Achayindee S, Petsom A, Pudhom K, Singtothong P, Surachetapan C, Vilaivan T. Two new cembranoids from Croton oblongifolius. J Nat Prod. 1998;61:652–4.
Kitazawa E, Ogiso A, Takahashi S, Aiya S, Kurabayashi M, Kuwano H, Hata T, Tamura C. Plaunol A and B, new anti-ulcer diterpenelactones from Croton sublyratus. Tetrahedron Lett. 1979;20:1117–20.
Rakotonandrasana OL, Raharinjato FH, Rajaonarivelo M, Dumontet V, Martin MT, Bignon J, Rasoanaivo P. Cytotoxic 3,4-seco-Atisane diterpenoids from Croton barorum and Croton goudotii. J Nat Prod. 2010;73:1730–3.
Wang GC, Li JG, Li GQ, Xu JJ, Wu X, Ye WC, Li YL. Clerodane diterpenoids from Croton crassifolius. J Nat Prod. 2012;75:2188–92.
Terefe EM, Okalebo FA, Derese S, Muriuki J, Rotich W, Mas-Claret E, Sadgrove N, Padilla-González GF, Prescott TAK, Siddique H, Langat MK. Constituents of Croton megalocarpus with potential anti-HIV activity. J Nat Prod. 2022;85:1861–6.
Adelekan AM, Prozesky EA, Hussein AA, Ureña LD, van Rooyen PH, Liles DC, Meyer JJM, Rodríguez B. Bioactive diterpenes and other constituents of Croton steenkampianus. J Nat Prod. 2008;71:1919–22.
Thongtan J, Kittakoop P, Ruangrungsi N, Saenboonrueng J, Thebtaranonth Y. New antimycobacterial and antimalarial 8,9-secokaurane diterpenes from Croton kongensis. J Nat Prod. 2003;66:868–70.
Liu CP, Xu JB, Zhao JX, Xu CH, Dong L, Ding J, Yue JM. Diterpenoids from Croton laui and their cytotoxic and antimicrobial activities. J Nat Prod. 2014;77:1013–20.
Jiang ZY, Liu CJ, Niu Q, Yan XY, Xiao D, Zhang HL, Huang CQ, Shi SL, Zuo AX, He HP. In vitro hypoglycemic diterpenoids from the roots of Croton yunnanensis. J Nat Prod. 2023;86:199–208.
Li B, Hans-Joachim E. Croton. Flora China. 2008;11:258–64.
Yuan FY, Tang ZY, Huang D, Li W, Wu SQ, Huang JL, Yan XL, Fan RZ, Tang GH, Yin S. Tigliane and rhamnofolane glycosides from Euphorbia wallichii prevent oxidative stress-induced neuronal death in PC-12 cells. Bioorg Chem. 2022;128: 106103.
Wu SQ, Fan RZ, Yuan FY, Li W, Huang D, Li S, Tang GH, Huang ZS, Yin S. Euphylonoids A and B, two highly modified jatrophane diterpenoids with potent lipid-lowering activity from Euphorbia hylonoma. Org Lett. 2022;24:8854–8.
Zou MF, Pan YH, Hu R, Yuan FY, Huang D, Tang GH, Li W, Yin S. Highly modified nor-clerodane diterpenoids from Croton yanhuii. Fitoterapia. 2021;153: 104979.
Krebs HC, Ramiarantsoa H. Clerodane diterpenes and other constituents of Croton hovarum. Phytochemistry. 1996;41:561–3.
Maslovskaya LA, Savchenko AI, Gordon VA, Reddell PW, Pierce CJ, Parsons PG, Williams CM. EBC-316, 325–327, and 345: New pimarane diterpenes from Croton insularis found in the Australian rainforest. Aust J Chem. 2015;68:652–9.
Itokawa H, Ichihara Y, Takeya K, Morita H, Motidome M. Diterpenes from Croton Salutaris. Phytochemistry. 1991;30(12):4071–3.
Ravindranath N, Ravinder Reddy M, Ramesh C, Ramu R, Prabhakar A, Jagadeesh B, Das B. New lathyrane and podocarpane diterpenoids from Jatropha curcas. Chem Pharm Bull. 2004;52:608–11.
Chao CH, Cheng JC, Shen DY, Wu TS. Anti-hepatitis C virus dinorditerpenes from the roots of Flueggea virosa. J Nat Prod. 2014;77:22–8.
Kim KH, Lee KH, Choi SU, Kim YH, Lee KR. Terpene and phenolic constituents of Lactuca indica L. Arch Pharm Res. 2008;31:983–8.
Kisiel W, Michalska K, Szneler E. Norisoprenoids from aerial parts of Cichorium pumilum. Biochem Syst Ecol. 2004;32:343–6.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 82273804), the Science and Technology Program of Guangzhou, China (No. 202201011613), the Southern Marine Science and Engineering Guangdong Laboratory (Zhuhai) (No. SML2021SP301), the Open Program of Shenzhen Bay Laboratory (No. SZBL2021080601007), the Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program (No. 2017BT01Y093), and the Guangdong Basic and Applied Basic Research Foundation, China (No. 2021B1515140062).
Author information
Authors and Affiliations
Contributions
XW carried out the experiments, analyzed the results, and wrote the manuscript; FYY did the ECD calculation; JLH and HHG screened the biological activities; GHT and SY designed and checked the whole manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that there are no competing interests associated with this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
The NMR, ECD, and HSMS spectra of 1–6, 13C NMR spectroscopic data for 7–13, general experimental procedures, extraction and isolation, and ECD calculation for 4.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wei, X., Huang, JL., Gao, HH. et al. New halimane and clerodane diterpenoids from Croton cnidophyllus. Nat. Prod. Bioprospect. 13, 21 (2023). https://doi.org/10.1007/s13659-023-00386-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13659-023-00386-z