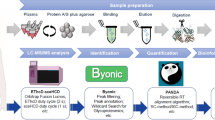
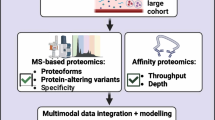
Abstract
Alpha-1,6 fucosylation of N-glycans (core fucosylation, CF) represents a unique form of N-glycans and is widely involved in disease progression. In order to accurately identify CF glycoproteins, several approaches have been developed based on sequential cleavage with different glycosidases to truncate the N-glycans. Since multi-step sample treatments may introduce quantitation bias and affect the practicality of these approaches in large-scale applications. Here, we systematically evaluated the performance of the single-step treatment of intact glycopeptides by endoglycosidase F3 for CF glycoproteome. The single-step truncation (SST) strategy demonstrated higher quantitative stability and higher efficiency compared with previous approaches. The strategy was further practiced on both cell lines and serum samples. The dysregulation of CF glycopeptides between preoperative and postoperative serum from patients with pancreatic ductal adenocarcinoma was revealed, and the CF modifications of BCHE_N369, CDH5_N112 and SERPIND1_N49 were found to be potential prognostic markers. This study thus provides an efficient solution for large-scale quantitative analysis of the CF glycoproteome.




Similar content being viewed by others
Data Availability
The data in this article will be available on reasonable request to the corresponding author.
References
Ohtsubo, K., Marth, J.D.: Glycosylation in cellular mechanisms of health and disease. Cell. 126(5), 855–867 (2006). https://doi.org/10.1016/j.cell.2006.08.019
Moremen, K.W., Tiemeyer, M., Nairn, A.V.: Vertebrate protein glycosylation: Diversity, synthesis and function. Nat. Rev. Mol. Cell. Biol. 13(7), 448–462 (2012). https://doi.org/10.1038/nrm3383
Bastian, K., Scott, E., Elliott, D.J., Munkley, J.: FUT8 Alpha-(1,6)-Fucosyltransferase in Cancer. Int. J. Mol. Sci. 22(1) (2021). https://doi.org/10.3390/ijms22010455
Takahashi, M., Kuroki, Y., Ohtsubo, K., Taniguchi, N.: Core fucose and bisecting GlcNAc, the direct modifiers of the N-glycan core: Their functions and target proteins. Carbohydr. Res. 344(12), 1387–1390 (2009). https://doi.org/10.1016/j.carres.2009.04.031
Zhao, Y.P., Xu, X.Y., Fang, M., Wang, H., You, Q., Yi, C.H., Ji, J., Gu, X., Zhou, P.T., Cheng, C., Gao, C.F.: Decreased core-fucosylation contributes to malignancy in gastric cancer. PLoS One. 9(4) (2014). https://doi.org/10.1371/journal.pone.0094536 e94536
Saldova, R., Fan, Y., Fitzpatrick, J.M., Watson, R.W., Rudd, P.M.: Core fucosylation and alpha2-3 sialylation in serum N-glycome is significantly increased in prostate cancer comparing to benign prostate hyperplasia. Glycobiology. 21(2), 195–205 (2011). https://doi.org/10.1093/glycob/cwq147
Agrawal, P., Fontanals-Cirera, B., Sokolova, E., Jacob, S., Vaiana, C.A., Argibay, D., Davalos, V., McDermott, M., Nayak, S., Darvishian, F., Castillo, M., Ueberheide, B., Osman, I., Fenyö, D., Mahal, L.K., Hernando, E.: A Systems Biology Approach identifies FUT8 as a driver of Melanoma Metastasis. Cancer Cell. 31(6), 804–819e807 (2017). https://doi.org/10.1016/j.ccell.2017.05.007
Zhou, J., Yang, W., Hu, Y., Hoti, N., Liu, Y., Shah, P., Sun, S., Clark, D., Thomas, S., Zhang, H.: Site-specific fucosylation analysis identifying Glycoproteins Associated with aggressive prostate Cancer cell lines using Tandem Affinity enrichments of Intact Glycopeptides followed by Mass Spectrometry. Anal. Chem. 89(14), 7623–7630 (2017). https://doi.org/10.1021/acs.analchem.7b01493
Totten, S.M., Adusumilli, R., Kullolli, M., Tanimoto, C., Brooks, J.D., Mallick, P., Pitteri, S.J.: Multi-lectin Affinity Chromatography and quantitative proteomic analysis Reveal Differential Glycoform levels between prostate Cancer and Benign Prostatic Hyperplasia Sera. Sci. Rep. 8(1), 6509 (2018). https://doi.org/10.1038/s41598-018-24270-w
Yang, S.J., Zhang, H.: Glycan analysis by reversible reaction to hydrazide beads and mass spectrometry. Anal. Chem. 84(5), 2232–2238 (2012). https://doi.org/10.1021/ac202769k
Zhu, J., Wang, F., Chen, R., Cheng, K., Xu, B., Guo, Z., Liang, X., Ye, M., Zou, H.: Centrifugation assisted microreactor enables facile integration of trypsin digestion, hydrophilic interaction chromatography enrichment, and on-column deglycosylation for rapid and sensitive N-glycoproteome analysis. Anal. Chem. 84(11), 5146–5153 (2012). https://doi.org/10.1021/ac3000732
Sun, Z., Fu, B., Wang, G., Zhang, L., Xu, R., Zhang, Y., Lu, H.: High-throughput site-specific N-glycoproteomics reveals glyco-signatures for liver disease diagnosis. Natl. Sci. Rev. 10(1), nwac059 (2023). https://doi.org/10.1093/nsr/nwac059
Zhang, Y., Zheng, S., Mao, Y., Cao, W., Zhao, L., Wu, C., Cheng, J., Liu, F., Li, G., Yang, H.: Systems analysis of plasma IgG intact N-glycopeptides from patients with chronic kidney diseases via EThcD-sceHCD-MS/MS. Analyst. 146(23), 7274–7283 (2021). https://doi.org/10.1039/d1an01657a
Cerrato, A., Cavaliere, C., Montone, C.M., Piovesana, S.: New hydrophilic material based on hydrogel polymer for the selective enrichment of intact glycopeptides from serum protein digests. Anal. Chim. Acta. 1245, 340862 (2023). https://doi.org/10.1016/j.aca.2023.340862
Ruhaak, L.R., Xu, G., Li, Q., Goonatilleke, E., Lebrilla, C.B.: Mass Spectrometry approaches to glycomic and glycoproteomic analyses. Chem. Rev. 118(17), 7886–7930 (2018). https://doi.org/10.1021/acs.chemrev.7b00732
Segu, Z.M., Hussein, A., Novotny, M.V., Mechref, Y.: Assigning N-glycosylation sites of glycoproteins using LC/MSMS in conjunction with endo-M/exoglycosidase mixture. J. Proteome Res. 9(7), 3598–3607 (2010). https://doi.org/10.1021/pr100129n
Zhang, W., Cao, W., Huang, J., Wang, H., Wang, J., Xie, C., Yang, P.: PNGase F-mediated incorporation of (18)O into glycans for relative glycan quantitation. Analyst. 140(4), 1082–1089 (2015). https://doi.org/10.1039/c4an02073a
Ma, J., Sanda, M., Wei, R., Zhang, L., Goldman, R.: Quantitative analysis of core fucosylation of serum proteins in liver diseases by LC-MS-MRM. J Proteom. 189, 67–74 (2018). https://doi.org/10.1016/j.jprot.2018.02.003
Lang, R., Leinenbach, A., Karl, J., Swiatek-de Lange, M., Kobold, U., Vogeser, M.: An endoglycosidase-assisted LC-MS/MS-based strategy for the analysis of site-specific core-fucosylation of low-concentrated glycoproteins in human serum using prostate-specific antigen (PSA) as example. Clin. Chim. Acta. 480, 1–8 (2018). https://doi.org/10.1016/j.cca.2018.01.040
Donald, L.J., Spearman, M., Mishra, N., Komatsu, E., Butler, M., Perreault, H.: Mass spectrometric analysis of core fucosylation and sequence variation in a human-camelid monoclonal antibody. Mol. Omics. 16(3), 221–230 (2020). https://doi.org/10.1039/c9mo00168a
Ma, C., Qu, J., Li, X., Zhao, X., Li, L., Xiao, C., Edmunds, G., Gashash, E., Song, J., Wang, P.G.: Improvement of core-fucosylated glycoproteome coverage via alternating HCD and ETD fragmentation. J Proteom. 146, 90–98 (2016). https://doi.org/10.1016/j.jprot.2016.06.003
Cao, Q., Zhao, X., Zhao, Q., Lv, X., Ma, C., Li, X., Zhao, Y., Peng, B., Ying, W., Qian, X.: Strategy integrating stepped fragmentation and glycan diagnostic ion-based spectrum refinement for the identification of core fucosylated glycoproteome using mass spectrometry. Anal. Chem. 86(14), 6804–6811 (2014). https://doi.org/10.1021/ac501154a
Liu, M.Q., Zeng, W.F., Fang, P., Cao, W.Q., Liu, C., Yan, G.Q., Zhang, Y., Peng, C., Wu, J.Q., Zhang, X.J., Tu, H.J., Chi, H., Sun, R.X., Cao, Y., Dong, M.Q., Jiang, B.Y., Huang, J.M., Shen, H.L., Wong, C.C.L., He, S.M., Yang, P.Y.: pGlyco 2.0 enables precision N-glycoproteomics with comprehensive quality control and one-step mass spectrometry for intact glycopeptide identification. Nat. Commun. 8(1), 438 (2017). https://doi.org/10.1038/s41467-017-00535-2
Chen, Z., Shen, J., Dong, W., Li, P., Xin, M., Liu, D., Jia, L., Zhu, B., Li, W., Sun, S.: Recognition of Core-Fucosylated Glycopeptides based on the Y1 + Fuc/Y1 ratio in low-energy HCD Spectra. Anal. Chem. (2022). https://doi.org/10.1021/acs.analchem.2c03182
Jia, W., Lu, Z., Fu, Y., Wang, H.P., Wang, L.H., Chi, H., Yuan, Z.F., Zheng, Z.B., Song, L.N., Han, H.H., Liang, Y.M., Wang, J.L., Cai, Y., Zhang, Y.K., Deng, Y.L., Ying, W.T., He, S.M., Qian, X.H.: A strategy for precise and large scale identification of core fucosylated glycoproteins. Mol. Cell. Proteomics. 8(5), 913–923 (2009). https://doi.org/10.1074/mcp.M800504-MCP200
Zhao, X., Yu, Z., Huang, Y., Liu, C., Wang, M., Li, X., Qian, X., Ying, W.: Integrated Strategy for large-scale investigation on protein core Fucosylation Stoichiometry based on glycan-simplification and paired-peaks-extraction. Anal. Chem. 92(4), 2896–2901 (2020). https://doi.org/10.1021/acs.analchem.9b05276
Yang, X., Leslie, G., Doroszuk, A., Schneider, S., Allen, J., Decker, B., Dunning, A.M., Redman, J., Scarth, J., Plaskocinska, I., Luccarini, C., Shah, M., Pooley, K., Dorling, L., Lee, A., Adank, M.A., Adlard, J., Aittomäki, K., Andrulis, I.L., Ang, P., Barwell, J., Bernstein, J.L., Bobolis, K., Borg, Ã., Blomqvist, C., Claes, K.B.M., Concannon, P., Cuggia, A., Culver, J.O., Damiola, F., de Pauw, A., Diez, O., Dolinsky, J.S., Domchek, S.M., Engel, C., Evans, D.G., Fostira, F., Garber, J., Golmard, L., Goode, E.L., Gruber, S.B., Hahnen, E., Hake, C., Heikkinen, T., Hurley, J.E., Janavicius, R., Kleibl, Z., Kleiblova, P., Konstantopoulou, I., Kvist, A., Laduca, H., Lee, A.S.G., Lesueur, F., Maher, E.R., Mannermaa, A., Manoukian, S., McFarland, R., McKinnon, W., Meindl, A., Metcalfe, K., Mohd Taib, N.A., Moilanen, J., Nathanson, K.L., Neuhausen, S., Ng, P.S., Nguyen-Dumont, T., Nielsen, S.M., Obermair, F., Offit, K., Olopade, O.I., Ottini, L., Penkert, J., Pylkäs, K., Radice, P., Ramus, S.J., Rudaitis, V., Side, L., Silva-Smith, R., Silvestri, V., Skytte, A.B., Slavin, T., Soukupova, J., Tondini, C., Trainer, A.H., Unzeitig, G., Usha, L., van Overeem Hansen, T., Whitworth, J., Wood, M., Yip, C.H., Yoon, S.Y., Yussuf, A., Zogopoulos, G., Goldgar, D., Hopper, J.L., Chenevix-Trench, G., Pharoah, P., George, S.H.L., Balmaña, J., Houdayer, C., James, P., El-Haffaf, Z., Ehrencrona, H., Janatova, M., Peterlongo, P., Nevanlinna, H., Schmutzler, R., Teo, S.H., Robson, M., Pal, T., Couch, F., Weitzel, J.N., Elliott, A., Southey, M., Winqvist, R., Easton, D.F., Foulkes: W.D., Antoniou, A.C., Tischkowitz, M.: Cancer Risks Associated With Germline PALB2 Pathogenic Variants: An International Study of 524 Families. J Clin Oncol 38(7), 674–685 doi: (2020). https://doi.org/10.1200/jco.19.01907
Siegel, R.L., Miller, K.D., Jemal, A.: Cancer statistics, 2020. CA Cancer J Clin. 70(1), 7–30 (2020). https://doi.org/10.3322/caac.21590
Tan, Z., Yin, H., Nie, S., Lin, Z., Zhu, J., Ruffin, M.T., Anderson, M.A., Simeone, D.M., Lubman, D.M.: Large-scale identification of core-fucosylated glycopeptide sites in pancreatic cancer serum using mass spectrometry. J. Proteome Res. 14(4), 1968–1978 (2015). https://doi.org/10.1021/acs.jproteome.5b00068
Tada, K., Ohta, M., Hidano, S., Watanabe, K., Hirashita, T., Oshima, Y., Fujnaga, A., Nakanuma, H., Masuda, T., Endo, Y., Takeuchi, Y., Iwashita, Y., Kobayashi, T., Inomata, M.: Fucosyltransferase 8 plays a crucial role in the invasion and metastasis of pancreatic ductal adenocarcinoma. Surg. Today. 50(7), 767–777 (2020). https://doi.org/10.1007/s00595-019-01953-z
Liang, C., Fukuda, T., Isaji, T., Duan, C., Song, W., Wang, Y., Gu, J.: α1,6-Fucosyltransferase contributes to cell migration and proliferation as well as to cancer stemness features in pancreatic carcinoma. Biochim. Biophys. Acta Gen. Subj. 1865(6), 129870 (2021). https://doi.org/10.1016/j.bbagen.2021.129870
Turiák, L., Sugár, S., Ács, A., Tóth, G., Gömöry, Ã., Telekes, A., Vékey, K., Drahos, L.: Site-specific N-glycosylation of HeLa cell glycoproteins. Sci. Rep. 9(1), 14822 (2019). https://doi.org/10.1038/s41598-019-51428-x
Ma, C., Zhang, Q., Qu, J., Zhao, X., Li, X., Liu, Y., Wang, P.G.: A precise approach in large scale core-fucosylated glycoprotein identification with low- and high-normalized collision energy. J Proteom. 114, 61–70 (2015). https://doi.org/10.1016/j.jprot.2014.09.001
Cao, L., Lih, T.M., Hu, Y., Schnaubelt, M., Chen, S.Y., Zhou, Y., Guo, C., Dong, M., Yang, W., Eguez, R.V., Chen, L., Clark, D.J., Sodhi, A., Li, Q.K., Zhang, H.: Characterization of core fucosylation via sequential enzymatic treatments of intact glycopeptides and mass spectrometry analysis. Nat. Commun. 13(1), 3910 (2022). https://doi.org/10.1038/s41467-022-31472-4
Li, H., Li, L., Cheng, K., Ning, Z., Mayne, J., Zhang, X., Walker, K., Chen, R., Twine, S., Li, J., Figeys, D.: Chemoenzymatic method for glycoproteomic N-Glycan type quantitation. Anal. Chem. 92(1), 1618–1627 (2020). https://doi.org/10.1021/acs.analchem.9b04937
Yu, Z., Zhao, X., Tian, F., Zhao, Y., Zhang, Y., Huang, Y., Qian, X., Ying, W.: Sequential fragment ion filtering and endoglycosidase-assisted identification of intact glycopeptides. Anal. Bioanal Chem. 409(12), 3077–3087 (2017). https://doi.org/10.1007/s00216-017-0195-z
Product Information Sheet - E2264.pdf>.:
Kong, R., Qian, X., Ying, W.: Pancreatic cancer cells spectral library by DIA-MS and the phenotype analysis of gemcitabine sensitivity. Sci. Data. 9(1), 283 (2022). https://doi.org/10.1038/s41597-022-01407-1
Mehra, S., Deshpande, N., Nagathihalli, N.: Targeting PI3K pathway in pancreatic ductal adenocarcinoma: Rationale and Progress. Cancers (Basel). 13(17) (2021). https://doi.org/10.3390/cancers13174434
Fu, Y., Yao, N., Ding, D., Zhang, X., Liu, H., Ma, L., Shi, W., Zhu, C., Tang, L.: TMEM158 promotes pancreatic cancer aggressiveness by activation of TGFβ1 and PI3K/AKT signaling pathway. J. Cell. Physiol. 235(3), 2761–2775 (2020). https://doi.org/10.1002/jcp.29181
Biankin, A.V., Waddell, N., Kassahn, K.S., Gingras, M.C., Muthuswamy, L.B., Johns, A.L., Miller, D.K., Wilson, P.J., Patch, A.M., Wu, J., Chang, D.K., Cowley, M.J., Gardiner, B.B., Song, S., Harliwong, I., Idrisoglu, S., Nourse, C., Nourbakhsh, E., Manning, S., Wani, S., Gongora, M., Pajic, M., Scarlett, C.J., Gill, A.J., Pinho, A.V., Rooman, I., Anderson, M., Holmes, O., Leonard, C., Taylor, D., Wood, S., Xu, Q., Nones, K., Fink, J.L., Christ, A., Bruxner, T., Cloonan, N., Kolle, G., Newell, F., Pinese, M., Mead, R.S., Humphris, J.L., Kaplan, W., Jones, M.D., Colvin, E.K., Nagrial, A.M., Humphrey, E.S., Chou, A., Chin, V.T., Chantrill, L.A., Mawson, A., Samra, J.S., Kench, J.G., Lovell, J.A., Daly, R.J., Merrett, N.D., Toon, C., Epari, K., Nguyen, N.Q., Barbour, A., Zeps, N., Kakkar, N., Zhao, F., Wu, Y.Q., Wang, M., Muzny, D.M., Fisher, W.E., Brunicardi, F.C., Hodges, S.E., Reid, J.G., Drummond, J., Chang, K., Han, Y., Lewis, L.R., Dinh, H., Buhay, C.J., Beck, T., Timms, L., Sam, M., Begley, K., Brown, A., Pai, D., Panchal, A., Buchner, N., De Borja, R., Denroche, R.E., Yung, C.K., Serra, S., Onetto, N., Mukhopadhyay, D., Tsao, M.S., Shaw, P.A., Petersen, G.M., Gallinger, S., Hruban, R.H., Maitra, A., Iacobuzio-Donahue, C.A., Schulick, R.D., Wolfgang, C.L., Morgan, R.A., Lawlor, R.T., Capelli, P., Corbo, V., Scardoni, M., Tortora, G., Tempero, M.A., Mann, K.M., Jenkins, N.A., Perez-Mancera, P.A., Adams, D.J., Largaespada, D.A., Wessels, L.F., Rust, A.G., Stein, L.D., Tuveson, D.A., Copeland, N.G., Musgrove, E.A., Scarpa, A., Eshleman, J.R., Hudson, T.J., Sutherland, R.L., Wheeler: Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature. 491(7424), 399–405 (2012). D.A., Pearson, J.V., McPherson, J.D., Gibbs, R.A., Grimmond, S.M https://doi.org/10.1038/nature11547
Jurcak, N.R., Rucki, A.A., Muth, S., Thompson, E., Sharma, R., Ding, D., Zhu, Q., Eshleman, J.R., Anders, R.A., Jaffee, E.M., Fujiwara, K., Zheng, L.: Axon Guidance Molecules promote Perineural Invasion and Metastasis of Orthotopic pancreatic tumors in mice. Gastroenterology. 157(3), 838–850e836 (2019). https://doi.org/10.1053/j.gastro.2019.05.065
Müller, M.W., Giese, N.A., Swiercz, J.M., Ceyhan, G.O., Esposito, I., Hinz, U., Büchler, P., Giese, T., Büchler, M.W., Offermanns, S., Friess, H.: Association of axon guidance factor semaphorin 3A with poor outcome in pancreatic cancer. Int. J. Cancer. 121(11), 2421–2433 (2007). https://doi.org/10.1002/ijc.22949
Gao, Y., Liu, X., Li, T., Wei, L., Yang, A., Lu, Y., Zhang, J., Li, L., Wang, S., Yin, F.: Cross-validation of genes potentially associated with overall survival and drug resistance in ovarian cancer. Oncol. Rep. 37(5), 3084–3092 (2017). https://doi.org/10.3892/or.2017.5534
Willis, S., Villalobos, V.M., Gevaert, O., Abramovitz, M., Williams, C., Sikic, B.I., Leyland-Jones, B.: Single gene prognostic biomarkers in ovarian Cancer: A Meta-analysis. PLoS One. 11(2), e0149183 (2016). https://doi.org/10.1371/journal.pone.0149183
Rochefort, P., Chabaud, S., Pierga, J.Y., Tredan, O., Brain, E., Bidard, F.C., Schiffler, C., Polena, H., Khalil-Mgharbel, A., Vilgrain, I., Bachelot, T.: Soluble VE-cadherin in metastatic breast cancer: An independent prognostic factor for both progression-free survival and overall survival. Br. J. Cancer. 116(3), 356–361 (2017). https://doi.org/10.1038/bjc.2016.427
Higuchi, K., Inokuchi, M., Takagi, Y., Ishikawa, T., Otsuki, S., Uetake, H., Kojima, K., Kawano, T.: Cadherin 5 expression correlates with poor survival in human gastric cancer. J. Clin. Pathol. 70(3), 217–221 (2017). https://doi.org/10.1136/jclinpath-2016-203640
Huang, H., Zhang, Q., Zhang, Y., Sun, X., Liu, C., Wang, Q., Huang, Y., Li, Q., Wu, Z., Pu, C., Sun, A.: Identification of the level of Exosomal protein by parallel reaction Monitoring Technology in HCC Patients. Int. J. Gen. Med. 15, 7831–7842 (2022). https://doi.org/10.2147/ijgm.S384140
Cavalcante Mde, S., Torres-Romero, J.C., Lobo, M.D., Moreno, F.B., Bezerra, L.P., Lima, D.S., Matos, J.C., Rde, M., Monteiro-Moreira, A.: A panel of glycoproteins as candidate biomarkers for early diagnosis and treatment evaluation of B-cell acute lymphoblastic leukemia. Biomark. Res. 4, 1 (2016). https://doi.org/10.1186/s40364-016-0055-6
Funding
This work was supported by the National Natural Science Foundation of China (32001048).
Author information
Authors and Affiliations
Contributions
All authors have given approval to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
The serum samples used in this study were obtained from the Peking University Cancer Hospital in China, with the approval of the Research Ethics Committee.
Competing interests
The authors declare no competing conflicts of interest.
Supporting information
The Supporting Information is available free of charge on the Publications website.
Experimental details and supporting figures and tables(PDF).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10719_2023_10130_MOESM3_ESM.xlsx
Supplementary Material 3: Table S5. The proteins and site-specific CF glycopeptides differentially expressed between pre- and postoperative serum of patients with PDAC (XLSX)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Min, Y., Wu, J., Hou, W. et al. Differential analysis of core-fucosylated glycoproteomics enabled by single-step truncation of N-glycans. Glycoconj J 40, 541–549 (2023). https://doi.org/10.1007/s10719-023-10130-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10719-023-10130-x