Abstract
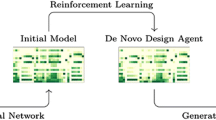
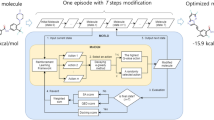
Generative approaches to molecular design are an area of intense study in recent years as a method to generate new pharmaceuticals with desired properties. Often though, these types of efforts are constrained by limited experimental activity data, resulting in either models that generate molecules with poor performance or models that are overfit and produce close analogs of known molecules. In this paper, we reduce this data dependency for the generation of new chemotypes by incorporating docking scores of known and de novo molecules to expand the applicability domain of the reward function and diversify the compounds generated during reinforcement learning. Our approach employs a deep generative model initially trained using a combination of limited known drug activity and an approximate docking score provided by a second machine learned Bayes regression model, with final evaluation of high scoring compounds by a full docking simulation. This strategy results in molecules with docking scores improved by 10–20% compared to molecules of similar size, while being 130 × faster than a docking only approach on a typical GPU workstation. We also show that the increased docking scores correlate with (1) docking poses with interactions similar to known inhibitors and (2) result in higher MM-GBSA binding energies comparable to the energies of known DDR1 inhibitors, demonstrating that the Bayesian model contains sufficient information for the network to learn to efficiently interact with the binding pocket during reinforcement learning. This outcome shows that the combination of the learned latent molecular representation along with the feature-based docking regression is sufficient for reinforcement learning to infer the relationship between the molecules and the receptor binding site, which suggest that our method can be a powerful tool for the discovery of new chemotypes with potential therapeutic applications.




Similar content being viewed by others
Data availability
All necessary data and code to evaluate or reproduce this work are publicly available through the cited sources, or included as supplemental files with this submission. Data: Training data as curated by our team is available as a supplemental file. The sources of the molecules are indicated in Supplementary Table 1. Software: Smina can be obtained from https://sourceforge.net/projects/smina/. The GENTRL source code can be obtained from https://github.com/insilicomedicine/GENTRL. Information for how to obtain scikit-learn is available at https://scikit-learn.org/.
References
Lyu J, Irwin JJ, Shoichet BK (2023) Modeling the expansion of virtual screening libraries. Nat Chem Biol 19:712–718. https://doi.org/10.1038/s41589-022-01234-w
Lyu J, Wang S, Balius TE et al (2019) Ultra-large library docking for discovering new chemotypes. Nature 566:224–229. https://doi.org/10.1038/s41586-019-0917-9
Irwin JJ, Tang KG, Young J et al (2020) ZINC20—a free ultralarge-scale chemical database for ligand discovery. J Chem Inf Model 60:6065–6073
Shivanyuk AN, Ryabukhin SV, Tolmachev A et al (2007) Enamine real database: making chemical diversity real. Chemistry today 25:58–59
Varela-Rial A, Majewski M, De Fabritiis G (2022) Structure based virtual screening: Fast and slow. WIREs Comput Mol Sci 12:e1544. https://doi.org/10.1002/wcms.1544
Bragina ME, Daina A, Perez MA et al (2022) The SwissSimilarity 2021 web tool: novel chemical libraries and additional methods for an enhanced ligand-based virtual screening experience. Int J Mol Sci 23:811
Martinelli DD (2022) Generative machine learning for de novo drug discovery: a systematic review. Comput Biol Med 145:105403. https://doi.org/10.1016/j.compbiomed.2022.105403
Coleman RG, Carchia M, Sterling T et al (2013) Ligand pose and orientational sampling in molecular docking. PLoS ONE 8:e75992. https://doi.org/10.1371/journal.pone.0075992
Xu W, Lucke AJ, Fairlie DP (2015) Comparing sixteen scoring functions for predicting biological activities of ligands for protein targets. J Mol Graph Model 57:76–88. https://doi.org/10.1016/j.jmgm.2015.01.009
Zhavoronkov A, Ivanenkov YA, Aliper A et al (2019) Deep learning enables rapid identification of potent DDR1 kinase inhibitors. Nat Biotechnol 37:1038–1040. https://doi.org/10.1038/s41587-019-0224-x
Gainor JF, Chabner BA (2015) Ponatinib: accelerated disapproval. Oncologist 20:847–848. https://doi.org/10.1634/theoncologist.2015-0253
Zeng X, Wang F, Luo Y et al (2022) Deep generative molecular design reshapes drug discovery. Cell Rep Med. https://doi.org/10.1016/j.xcrm.2022.100794
Li Y, Zhang L, Wang Y et al (2022) Generative deep learning enables the discovery of a potent and selective RIPK1 inhibitor. Nat Commun 13:6891. https://doi.org/10.1038/s41467-022-34692-w
Grant LL, Sit CS (2021) De novo molecular drug design benchmarking. RSC Med Chem 12:1273–1280. https://doi.org/10.1039/D1MD00074H
Vella D, Ebejer J-P (2022) Few-shot learning for low-data drug discovery. J Chem Inf Model. https://doi.org/10.1021/acs.jcim.2c00779
Rogers D, Hahn M (2010) Extended-connectivity fingerprints. J Chem Inf Model 50:742–754
Jeon W, Kim D (2020) Autonomous molecule generation using reinforcement learning and docking to develop potential novel inhibitors. Sci Rep 10:22104. https://doi.org/10.1038/s41598-020-78537-2
Thomas M, Smith RT, O’Boyle NM et al (2021) Comparison of structure- and ligand-based scoring functions for deep generative models: a GPCR case study. J Cheminform 13:39. https://doi.org/10.1186/s13321-021-00516-0
Sadybekov AA, Sadybekov AV, Liu Y et al (2022) Synthon-based ligand discovery in virtual libraries of over 11 billion compounds. Nature 601:452–459. https://doi.org/10.1038/s41586-021-04220-9
Gentile F, Yaacoub JC, Gleave J et al (2022) Artificial intelligence–enabled virtual screening of ultra-large chemical libraries with deep docking. Nat Protoc 17:672–697
Berenger F, Kumar A, Zhang KYJ, Yamanishi Y (2021) Lean-docking: exploiting ligands’ predicted docking scores to accelerate molecular docking. J Chem Inf Model 61:2341–2352. https://doi.org/10.1021/acs.jcim.0c01452
Bucinsky L, Bortňák D, Gall M et al (2022) Machine learning prediction of 3CL SARS-CoV-2 docking scores. Comput Biol Chem 98:107656. https://doi.org/10.1016/j.compbiolchem.2022.107656
MolFinder: an evolutionary algorithm for the global optimization of molecular properties and the extensive exploration of chemical space using SMILES | Journal of Cheminformatics | Full Text. https://jcheminf.biomedcentral.com/articles/https://doi.org/10.1186/s13321-021-00501-7. Accessed 21 Jun 2023
Ciepliński T, Danel T, Podlewska S, Jastrzȩbski S (2023) Generative models should at least be able to design molecules that dock well: a new benchmark. J Chem Inf Model 63:3238–3247. https://doi.org/10.1021/acs.jcim.2c01355
Gómez-Bombarelli R, Wei JN, Duvenaud D et al (2018) Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent Sci 4:268–276
Kusner MJ, Paige B, Hernández-Lobato JM (2017) Grammar variational autoencoder. In: International conference on machine learning. PMLR, pp 1945–1954
Olivecrona M, Blaschke T, Engkvist O, Chen H (2017) Molecular de-novo design through deep reinforcement learning. J Cheminform 9:48. https://doi.org/10.1186/s13321-017-0235-x
Gao Y, Zhou J, Li J (2021) Discoidin domain receptors orchestrate cancer progression: a focus on cancer therapies. Cancer Sci 112:962–969. https://doi.org/10.1111/cas.14789
Moll S, Desmoulière A, Moeller MJ et al (2019) DDR1 role in fibrosis and its pharmacological targeting. Biochimica et Biophysica Acta (BBA) - Mol Cell Res 1866:118474. https://doi.org/10.1016/j.bbamcr.2019.04.004
Tian Y, Bai F, Zhang D (2022) New target DDR1: A “double-edged sword” in solid tumors. Biochimica et Biophysica Acta (BBA) -Rev Cancer 1878:188829
Hinton GE, Roweis S (2002) Stochastic neighbor embedding. Advances in neural information processing systems 15. https://proceedings.neurips.cc/paper_files/paper/2002/hash/6150ccc6069bea6b5716254057a194ef-Abstract.html
Koes DR, Baumgartner MP, Camacho CJ (2013) Lessons learned in empirical scoring with smina from the CSAR 2011 benchmarking exercise. J Chem Inf Model 53:1893–1904
Pedregosa F, Varoquaux G, Gramfort A et al (2011) Scikit-learn: machine learning in python. J Machine Learn Res 12:2825–2830
Kohonen T (1990) The self-organizing map. Proc IEEE 78:1464–1480
Kaiser TM, Burger PB, Butch CJ et al (2018) A machine learning approach for predicting HIV reverse transcriptase mutation susceptibility of biologically active compounds. J Chem Inf Model 58:1544–1552
Kaiser TM, Dentmon ZW, Dalloul CE et al (2020) Accelerated discovery of novel ponatinib analogs with improved properties for the treatment of parkinson’s disease. ACS Med Chem Lett 11:491–496
Pribut N, Kaiser TM, Wilson RJ et al (2020) Accelerated discovery of potent fusion inhibitors for respiratory syncytial virus. ACS Infect Dis 6:922–929
Cox BD, Prosser AR, Sun Y et al (2015) Pyrazolo-piperidines exhibit dual inhibition of CCR5/CXCR4 HIV entry and reverse transcriptase. ACS Med Chem Lett 6:753–757
Shi Q, Kaiser TM, Dentmon ZW et al (2015) Design and validation of FRESH, a drug discovery paradigm resting on robust chemical synthesis. ACS Med Chem Lett 6:518–522
Lipinski CA (2004) Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Pan Y, Huang N, Cho S, MacKerell AD (2003) Consideration of molecular weight during compound selection in virtual target-based database screening. J Chem Inf Comput Sci 43:267–272
Bajusz D, Rácz A, Héberger K (2015) Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations? J Chem 7:1–13
Bouysset C, Fiorucci S (2021) ProLIF: a library to encode molecular interactions as fingerprints. J Cheminform 13:72. https://doi.org/10.1186/s13321-021-00548-6
Eastman P, Swails J, Chodera JD et al (2017) OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLoS Comput Biol 13:e1005659
Tuccinardi T (2021) What is the current value of MM/PBSA and MM/GBSA methods in drug discovery? Expert Opin Drug Discov 16:1233–1237. https://doi.org/10.1080/17460441.2021.1942836
Altae-Tran H, Ramsundar B, Pappu AS, Pande V (2017) Low data drug discovery with one-shot learning. ACS Cent Sci 3:283–293. https://doi.org/10.1021/acscentsci.6b00367
Mendez D, Gaulton A, Bento AP et al (2019) ChEMBL: towards direct deposition of bioassay data. Nucleic Acids Res 47:D930–D940
Gabrielson SW (2018) SciFinder. J Med Libr Assoc: JMLA 106:588
Polykovskiy D, Zhebrak A, Sanchez-Lengeling B et al (2020) Molecular sets (MOSES): a benchmarking platform for molecular generation models. Front Pharmacol 11:565644
Sterling T, Irwin JJ (2015) ZINC 15–ligand discovery for everyone. J Chem Inf Model 55:2324–2337
Trott O, Olson AJ (2009) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem NA-NA. https://doi.org/10.1002/jcc.21334
Richter H, Satz AL, Bedoucha M et al (2018) DNA-encoded library-derived DDR1 inhibitor prevents fibrosis and renal function loss in a genetic mouse model of Alport syndrome. ACS Chem Biol 14:37–49
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Bento AP, Hersey A, Félix E et al (2020) An open source chemical structure curation pipeline using RDKit. J Cheminform 12:1–16
Halgren TA (1996) Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J Comput Chem 17:490–519
O’Boyle NM, Banck M, James CA et al (2011) Open Babel: an open chemical toolbox. J Cheminform 3:1–14
Vettigli G (2022) MiniSom
Acknowledgements
Support for this work was provided by start up funds issued by Nanjing University and by Ice-Kredit.
Funding
Support for this work was provided by startup funds issued by Nanjing University and by Ice-Kredit.
Author information
Authors and Affiliations
Contributions
YWs numbered as in the author order. YX YW1 YW2 PY CB and JW collected and analyzed data and created figures. YW3 LG and CB developed and guided the research. CB wrote the main manuscript text. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
LG is the CEO and shareholder of IceKredit.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiong, Y., Wang, Y., Wang, Y. et al. Improving drug discovery with a hybrid deep generative model using reinforcement learning trained on a Bayesian docking approximation. J Comput Aided Mol Des 37, 507–517 (2023). https://doi.org/10.1007/s10822-023-00523-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-023-00523-3