Abstract
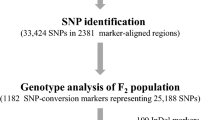
High-quality molecular markers are essential for marker-assisted selection to accelerate breeding progress. Compared with diploid species, recently diverged polyploid crop species tend to have highly similar homeologous subgenomes, which is expected to limit the development of broadly applicable locus-specific single-nucleotide polymorphism (SNP) assays. Furthermore, it is particularly challenging to make genome-wide marker sets for species that lack a reference genome. Here, we report the development of a genome-wide set of kompetitive allele specific PCR (KASP) markers for marker-assisted recurrent selection (MARS) in the tetraploid minor crop perilla. To find locus-specific SNP markers across the perilla genome, we used genotyping-by-sequencing (GBS) to construct linkage maps of two F2 populations. The two resulting high-resolution linkage maps comprised 2326 and 2454 SNP markers that spanned a total genetic distance of 2133 cM across 16 linkage groups and 2169 cM across 21 linkage groups, respectively. We then obtained a final genetic map consisting of 22 linkage groups with 1123 common markers from the two genetic maps. We selected 96 genome-wide markers for MARS and confirmed the accuracy of markers in the two F2 populations using a high-throughput Fluidigm system. We confirmed that 91.8% of the SNP genotyping results from the Fluidigm assay were the same as the results obtained through GBS. These results provide a foundation for marker-assisted backcrossing and the development of new varieties of perilla.


Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available in the NABIC (National Agricultural Biotechnology Information Center, https://nabic.rda.go.kr/) repository, [NN-5662 ~ 5712].
References
Ahmadikhah A, Mirarab M, Pahlevani MH, Nayyeripasand L (2015) Marker-assisted backcrossing to develop an elite cytoplasmic male sterility line in rice. Plant Genome. https://doi.org/10.3835/plantgenome2014.07.0031
Ahmed HM (2018) Ethnomedicinal, phytochemical and pharmacological investigations of perilla frutescens(l.) britt. Molecules. https://doi.org/10.3390/molecules24010102
Allen GC, Flores-Vergara MA, Krasynanski S, Kumar S, Thompson WF (2006) A modified protocol for rapid dna isolation from plant tissues using cetyltrimethylammonium bromide. Nat Protoc 1(5):2320–2325. https://doi.org/10.1038/nprot.2006.384
Asif M (2011) Health effects of omega-3,6,9 fatty acids: Perilla frutescens is a good example of plant oils. Orient Pharm Exp Med 11(1):51–59. https://doi.org/10.1007/s13596-011-0002-x
Bielenberg DG, Rauh B, Fan S, Gasic K, Abbott AG, Reighard GL, Okie WR, Wells CE (2015) Genotyping by sequencing for SNP-based linkage map construction and QTL analysis of chilling requirement and bloom date in peach [Prunus persica (L.) batsch]. PLoS One 10(10):e0139406
Cox MP, Peterson DA, Biggs PJ (2010) SolexaQA: At-a-glance quality assessment of illumina second-generation sequencing data. BMC Bioinformatics 11:485. https://doi.org/10.1186/1471-2105-11-485
Dhyani A, Chopra R, Garg M (2019) A review on nutritional value, functional properties and pharmacological application of perilla (Perilla frutescens L.). Biomed Pharmacol J 12(2):649–660. https://doi.org/10.13005/bpj/1685
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6(5):e19379. https://doi.org/10.1371/journal.pone.0019379
Eun MH, Han JH, Yoon JB, Lee J (2016) QTL mapping of resistance to the cucumber mosaic virus P1 strain in pepper using a genotyping-by-sequencing analysis. Hortic Environ Biotechnol 57(6):589–597. https://doi.org/10.1007/s13580-016-0128-3
Hussain W, Baenziger PS, Belamkar V, Guttieri MJ, Venegas JP, Easterly A, Sallam A, Poland J (2017) Genotyping-by-sequencing derived high-density linkage map and its application to qtl mapping of flag leaf traits in bread wheat. Sci Rep 7(1):16394. https://doi.org/10.1038/s41598-017-16006-z
Kang YJ, Lee BM, Nam M, Oh KW, Lee MH, Kim TH, Jo SH, Lee JH (2019) Identification of quantitative trait loci associated with flowering time in perilla using genotyping-by-sequencing. Mol Biol Rep 46(4):4397–4407. https://doi.org/10.1007/s11033-019-04894-5
Kim JE, Oh SK, Lee JH, Lee BM, Jo SH (2014) Genome-wide SNP calling using next generation sequencing data in tomato. Mol Cells 37(1):36–42. https://doi.org/10.14348/molcells.2014.2241
Kim H, Yoon JB, Lee J (2017) Development of fluidigm SNP type genotyping assays for marker-assisted breeding of chili pepper (Capsicum annuum L.). Korean J Hortic Sci Technol 35(4):465–479
Kim WJ, Ryu J, Im J, Kim SH, Kang SY, Lee JH, Jo SH, Ha BK (2018) Molecular characterization of proton beam-induced mutations in soybean using genotyping-by-sequencing. Mol Genet Genomics 293(5):1169–1180. https://doi.org/10.1007/s00438-018-1448-z
Kosambi DD (2016) The estimation of map distances from recombination values. Dd kosambi. Springer. p. 125–130. https://doi.org/10.1111/j.1469-1809.1943.tb02321.x
Lee MH, Oh KW, Kim MS, Kim SU, Kim JI, Oh EY, Pae SB, Yeo US, Kim T-H, Lee JH (2018) Detection of QTLs in an interspecific cross between Perilla citriodora× P. hirtella mapping population. Korean J Breed Sci 50(1):13–20
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li L, Zhao S, Su J, Fan S, Pang C, Wei H, Wang H, Gu L, Zhang C, Liu G et al (2017) High-density genetic linkage map construction by F2 populations and QTL analysis of early-maturity traits in upland cotton (Gossypium hirsutum L.). PLoS One 12(8):e0182918. https://doi.org/10.1371/journal.pone.0182918
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data Processing S (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Makhoul M, Rambla C, Voss-Fels K, Hickey L, Snowdon R, Obermeier C (2020) Overcoming polyploidy pitfalls: a user guide for effective SNP conversion into KASP markers in wheat. Theor Appl Genet 133:2413–2430. https://doi.org/10.1007/s00122-020-03608-x
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1):3. https://doi.org/10.14806/ej.17.1.200
Nitta M, Lee JK, Kang CW, Katsuta M, Yasumoto S, Liu D, Nagamine T, Ohnishi O (2005) The distribution of perilla species. Genet Resour Crop Evol 52(7):797–804. https://doi.org/10.1007/s10722-003-6017-5
Oh E, Lee MH, Kim JI, Kim S, Pae SB, Ha TJ (2018) Estimation of oil yield of perilla by seed characteristics and crude fat content. Korean J Crop Sci 63(2):158–163. https://doi.org/10.7740/kjcs.2018.63.2.158
Ooijen JW et al. (2006) Joinmap® 4: software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen 33(10.1371)
Park G, Jang HA, Jo SH, Park Y, Oh SK, Nam M (2018) Development of SNP marker set for marker-assisted backcrossing (MABC) in cultivating tomato varieties. Korean J Agri Sci 45(3):385–400. https://doi.org/10.7744/kjoas.20180061
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS One 7(2):e32253. https://doi.org/10.1371/journal.pone.0032253
Rasheed A, Hao Y, Xia X, Khan A, Xu Y, Varshney RK, He Z (2017) Crop breeding chips and genotyping platforms: progress, challenges, and perspectives. Mol Plant 10(8):1047–1064. https://doi.org/10.1016/j.molp.2017.06.008
Rossetto M, Henry RJ (2014) Escape from the laboratory: new horizons for plant genetics. Trends Plant Sci 19(9):554–555. https://doi.org/10.1016/j.tplants.2014.06.011
Sa KJ, Choi IY, Park KC, Lee JK (2018) Genetic diversity and population structure among accessions of Perilla frutescens (L.) britton in east asia using new developed microsatellite markers. Genes Genom 40(12):1319–1329. https://doi.org/10.1007/s13258-018-0727-8
Tamura K, Sakamoto M, Tanizawa Y, Mochizuki T, Matsushita S, Kato Y, Ishikawa T, Okuhara K, Nakamura Y, Bono H (2023) A highly contiguous genome assembly of red perilla (Perilla frutescens) domesticated in Japan. DNA Res 30(1):1–8. https://doi.org/10.1093/dnares/dsac044
Verma S, Gupta S, Bandhiwal N, Kumar T, Bharadwaj C, Bhatia S (2015) High-density linkage map construction and mapping of seed trait QTLs in chickpea (Cicer arietinum L.) using genotyping-by-sequencing (GBS). Sci Rep 5:17512. https://doi.org/10.1038/srep17512
Voorrips R (2002) Mapchart: software for the graphical presentation of linkage maps and qtls. J Hered 93(1):77–78. https://doi.org/10.1093/jhered/93.1.77
Ye G, Smith KF (2008) Marker-assisted gene pyramiding for inbred line development: basic principles and practical guidelines. Int J Plant Breed 2(1):1–10
Funding
This work was supported by a grant from the National Agricultural Genome Project (No. PJ013355032021), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Contributions
JO, JE, SH, and JH conceived and designed the experiments; JO and JE conducted the SNP analysis, linkage map construction, and data analysis; JK constructed the GBS library; MH developed the mapping population; TH provided the draft genome; KL participated in discussions about the experiments; SH and JH wrote and reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there are no relevant financial or non-financial conflicts of interest related to the content of this article.
Ethical approval and consent to participate
Not applicable.
Consent for publication
All authors consent to the publication of this paper.
Additional information
Communicated by Bing Yang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Oh, JE., Kim, JE., Kim, J. et al. Development of an SNP marker set for marker-assisted backcrossing using genotyping-by-sequencing in tetraploid perilla. Mol Genet Genomics 298, 1435–1447 (2023). https://doi.org/10.1007/s00438-023-02066-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-023-02066-6