Abstract
Background
This study aimed to screen out the potential diagnostic biomarkers for atherosclerosis (AS).
Methods
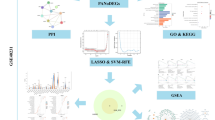
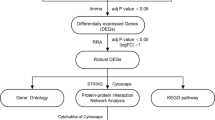
We downloaded the gene expression profiles GSE66360, GSE28829, GSE41571, GSE71226, and GSE100927 from the Gene Expression Omnibus (GEO) database. The differentially expressed genes (DEGs) were identified using the “limma” package in R. Weighted gene co-expression network analysis (WGCNA) was applied to reveal the correlation between genes in different samples. Subsequently, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed. The interaction pairs of proteins were retained by the STRING database, and the protein–protein interaction (PPI) network was visualized with the hub genes. Finally, the R packages “ggpubr” and “preprocessCore” were used to analyze immune cell infiltration.
Results
In total, 40 overlapping genes both in GSE66360 and GSE28829 were found to be related to the occurrence of AS. Further, the top 10 network hub genes including TYROBP, CSF1R, TLR2, CD14, CCL4, FCER1G, CD163, TREM1, PLEK, and C5AR1 were identified as significant key genes. Moreover, four genes (TYROBP, CSF1R, FCGR1B, and CD14) were verified that could efficiently diagnose AS. Finally, the gene TYROBP was found to have a strong correlation with immune-infiltrating cells.
Conclusion
Our study identified four genes (TYROBP, CSF1R, FCGR1B, and CD14) that may be effective biomarkers for AS, with the potential to guide the clinical diagnosis of AS.
Zusammenfassung
Hintergrund
Ziel der vorliegenden Studie war es, die potenziellen diagnostischen Biomarker für Arteriosklerose (AS) zu ermitteln.
Methoden
Dazu führten die Autoren einen Download der Genexpressionsprofile GSE66360, GSE28829, GSE41571, GSE71226 und GSE100927 von der Datenbank Gene Expression Omnibus (GEO) durch. Die unterschiedlich exprimierten Gene („differentially expressed genes“, DEG) wurden unter Verwendung der Software R „limma“ identifiziert. Unter Einsatz der gewichteten Gen-Koexpressions-Netzwerk-Analyse („weighted gene co-expression network analysis“, WGCNA) wurde die Korrelation zwischen Genen in verschiedenen Proben ermittelt. Anschließend wurden Signalweg-Anreicherungsanalysen mithilfe der Datenbanken Gene Ontology (GO) und Kyoto Encyclopedia of Genes and Genomes (KEGG) durchgeführt. Die in Wechselwirkung stehenden Proteinpaare wurden in der STRING-Datenbank erfasst, und das Protein-Protein-Interaktions(PPI)-Netzwerk wurde mit den Hub-Genen visualisiert. Schließlich wurden R „ggpubr“ und R „preprocessCore“ verwendet, um die Immunzellinfiltration zu analysieren.
Ergebnisse
Bei 40 sowohl in GSE66360 als auch in GSE28829 überlappenden Genen wurde festgestellt, dass sie mit dem Auftreten einer AS in Zusammenhang stehen. Darüber hinaus wurden die wichtigsten 10 Netzwerk-Hub-Gene einschließlich TYROBP, CSF1R, TLR2, CD14, CCL4, FCER1G, CD163, TREM1, PLEK und C5AR1 als wesentliche Schlüsselgene identifiziert. Außerdem wurden 4 Gene (TYROBP, CSF1R, FCGR1B und CD14) erfasst, die bedeutsam für die Diagnosestellung einer AS waren. Schließlich wurde bei dem Gen TYROBP festgestellt, dass es eine starke Korrelation mit immuninfiltrierenden Zellen aufweist.
Schlussfolgerung
In der vorliegenden Studie wurden 4 Gene entdeckt, die wirksame Biomarker für AS darstellen könnten und das Potenzial besitzen, zur klinischen Diagnose einer AS zu führen.











Similar content being viewed by others
References
Briggs RC, Atkinson JB, Miranda RN (2005) Variable expression of human myeloid specific nuclear antigen MNDA in monocyte lineage cells in atherosclerosis. J Cell Biochem 95:293–301. https://doi.org/10.1002/jcb.20435
Chang TT, Yang HY, Chen C, Chen JW (2020) CCL4 inhibition in atherosclerosis: effects on plaque stability, endothelial cell adhesiveness, and macrophages activation. Int J Mol Sci 21:6567. https://doi.org/10.3390/ijms21186567
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY (2014) cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8(Suppl 4):S11. https://doi.org/10.1186/1752-0509-8-s4-s11
Etzerodt A, Moestrup SK (2013) CD163 and inflammation: biological, diagnostic, and therapeutic aspects. Antioxid Redox Signal 18:2352–2363. https://doi.org/10.1089/ars.2012.4834
Geovanini GR, Libby P (2018) Atherosclerosis and inflammation: overview and updates. Clin Sci (Lond) 132:1243–1252. https://doi.org/10.1042/cs20180306
Guo L, Akahori H, Harari E, Smith SL, Polavarapu R et al (2018) CD163+ macrophages promote angiogenesis and vascular permeability accompanied by inflammation in atherosclerosis. J Clin Invest 128:1106–1124. https://doi.org/10.1172/jci93025
Hagan N, Kane JL, Grover D, Woodworth L, Madore C, Saleh J et al (2020) CSF1R signaling is a regulator of pathogenesis in progressive MS. Cell Death Dis 11:904. https://doi.org/10.1038/s41419-020-03084-7
Kim S, Karin M (2011) Role of TLR2-dependent inflammation in metastatic progression. Ann N Y Acad Sci 1217:191–206. https://doi.org/10.1111/j.1749-6632.2010.05882.x
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559. https://doi.org/10.1186/1471-2105-9-559
Liu H, Yang Y, Ge Y, Liu J, Zhao Y (2019) TERC promotes cellular inflammatory response independent of telomerase. Nucleic Acids Res 47:8084–8095. https://doi.org/10.1093/nar/gkz584
Liu Y, Liu N, Liu Q (2021) Constructing a ceRNA-immunoregulatory network associated with the development and prognosis of human atherosclerosis through weighted gene co-expression network analysis. Aging 13:3080–3100. https://doi.org/10.18632/aging.202486
McKinney C, Broen JC, Vonk MC, Beretta L, Hesselstrand R, Hunzelmann N et al (2012) Evidence that deletion at FCGR3B is a risk factor for systemic sclerosis. Genes Immun 13:458–460. https://doi.org/10.1038/gene.2012.15
Medina I, Cougoule C, Drechsler M, Bermudez B, Koenen RR, Sluimer J et al (2015) Hck/Fgr kinase deficiency reduces plaque growth and stability by blunting monocyte recruitment and intraplaque motility. Circulation 132:490–501. https://doi.org/10.1161/circulationaha.114.012316
Newman JL, Stone JR (2019) Immune checkpoint inhibition alters the inflammatory cell composition of human coronary artery atherosclerosis. Cardiovasc Pathol 43:107148. https://doi.org/10.1016/j.carpath.2019.107148
Roth GA, Mensah GA, Johnson CO, Addolorato G, Ammirati E, Baddour LM et al (2020) Global burden of cardiovascular diseases and risk factors, 1990–2019: update from the GBD 2019 study. J Am Coll Cardiol 76:2982–3021. https://doi.org/10.1016/j.jacc.2020.11.010
Soehnlein O, Libby P (2021) Targeting inflammation in atherosclerosis—From experimental insights to the clinic. Nat Rev Drug Discov. https://doi.org/10.1038/s41573-021-00198-1
Wolf D, Ley K (2019) Immunity and inflammation in atherosclerosis. Circ Res 124:315–327. https://doi.org/10.1161/circresaha.118.313591
Xue Y, Guo Y, Luo S, Zhou W, Xiang J, Zhu Y, Xiang Z, Shen J (2020) Aberrantly methylated-differentially expressed genes identify novel atherosclerosis risk subtypes. Front Genet 11:569572. https://doi.org/10.3389/fgene.2020.569572
Zhou J, Zhang C, Wu X, Xie Q, Li L, Chen Y, Yan H, Ren P, Huang X (2019) Identification of genes and pathways related to atherosclerosis comorbidity and depressive behavior via RNA-seq and bioinformation analysis in ApoE(-/-) mice. Ann Transl Med 7:733. https://doi.org/10.21037/atm.2019.11.118
Zhu Y, Xian X, Wang Z, Bi Y, Chen Q, Han X, Tang D, Chen R (2018) Research progress on the relationship between atherosclerosis and inflammation. Biomolecules 8:80. https://doi.org/10.3390/biom8030080
Author information
Authors and Affiliations
Contributions
JN, KH, and JX performed the literature search; JN and QL analyzed the data. This work was performed by JN and QL. The manuscript was written by JN and edited by CC.
Corresponding author
Ethics declarations
Conflict of interest
J. Ni, K. Huang, J. Xu, Q. Lu and C. Chen declare that they have no competing interests.
For this article no studies with human participants or animals were performed by any of the authors. All studies mentioned were in accordance with the ethical standards indicated in each case.
Rights and permissions
About this article
Cite this article
Ni, J., Huang, K., Xu, J. et al. Novel biomarkers identified by weighted gene co-expression network analysis for atherosclerosis. Herz (2023). https://doi.org/10.1007/s00059-023-05204-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00059-023-05204-3
Keywords
- Inflammation
- Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway
- Protein–protein interaction network
- Immune infiltration
- Clinical diagnosis