Abstract
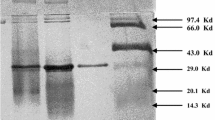
An extracellular cyclodextrin glucanotransferase (CGTase, EC 2.4.1.19) has been characterized for the first time in a strain of bacteria of the species Caldalkalibacillus mannanilyticus IB-OR17-B1. The enzyme was isolated from the culture supernatant using ultrafiltration and affinity adsorption on corn starch. The specific activity of the CGTase increased by 18 times as a result of purification with an enzyme yield of 56%. The molecular weight of the isolated enzyme was 70 kDa according to the denaturing electrophoresis in polyacrylamide gel. The CGTase of C. mannanilyticus IB-OR17-B1 demonstrated its maximum cyclizing activity at pH 8 and a temperature of 60°C, respectively, and it was stable in the pH range of 7–10 and temperatures of 70°C or less. The thermal stability of the enzyme under 70°C increased by 10–15% in the presence of 5–10 mM of calcium and magnesium salts. The cations of Ag+, Cu2+, Zn2+, Fe2+ and Fe3+ at a concentration of 5 mM inhibited CGTase activity by 90, 26, 23, 18, and 11%, respectively. Under the optimal conditions and enzyme-substrate ratio 1 U/g the isolated CGTase converted potato starch to mixture of α-, β-and γ-cyclodextrins with weight ratio 38.8 : 52.6 : 8.6 and a yield of 42% in 24 h.







Similar content being viewed by others
REFERENCES
Jemli, S., Messaoud, E., Ayadi-Zouari, D., Naili, B., Khemakhem, B., and Bejar, S., Biochem. Eng. J., 2007, vol. 34, no. 1, pp. 44–50. https://doi.org/10.1016/j.bej.2006.11.016
Aroob, I., Ahmad, N., and Rashid, N., Amylase, 2021, vol. 5, no. 1, pp. 23–37. https://doi.org/10.1515/amylase-2021-0003
Kurkov, S.V. and Loftsson, T., Int. J. Pharm., 2013, vol. 453, no. 1, pp. 167–180. https://doi.org/10.1016/j.ijpharm.2012.06.055
Astray, G., Gonzalez-Barreiro, C., Mejuto, J., Rial-Otero, R., and Simal-Gandara, J., Food Hydrocolloids, 2009, vol. 23, no. 7, pp. 1631–1640. https://doi.org/10.1016/j.foodhyd.2009.01.001
Abdel-Naby, M.A., El-Refai, H.A., and Abdel-Fattah, A.F., J. Appl. Microbiol., 2011, vol. 111, no. 5, pp. 1129–1137. https://doi.org/10.1111/j.1365-2672.2011.05136.x
Szejtli, J., Cyclodextrin Technology—Topics in Inclusion Science, Netherlands: Springer Science and Business Media, 2013. https://doi.org/10.1007/978-94-015-7797-7
Czinkoczky, R. and Nemeth, A., Hung. J. Ind. Chem., 2019, vol. 47, no. 2, pp. 5–10. https://doi.org/10.33927/hjic-2019-14
Hamoudi, M., Fattal, E., Gueutina, C., Nicolas, V., and Bochota, A., Int. J. Pharm., 2011, vol. 416, no. 2, pp. 507–514. https://doi.org/10.1016/j.ijpharm.2011.01.062
Marcon, F., Mathiron, D., Pilard, S., Lemaire-Hurtel, A., Dubaele, J., and Djedaini-Pilard, F., Int. J. Pharm., 2009, vol. 379, no. 2, pp. 244–250. https://doi.org/10.1016/j.ijpharm.2009.05.029
Sian, H.K., Said, M., Hassan, O., Kamaruddin, K., Ismail, A.F., Rahman, R., et al., Process Biochem., 2005, vol. 40, nos. 3–4, pp. 1101–1111. https://doi.org/10.1016/j.procbio.2004.03.018
Wang, J., Cao, Y., Sun, B., and Wang, C., Food Chem., 2011, vol. 127, no. 4, pp. 1680–1685. https://doi.org/10.1016/j.foodchem.2011.02.036
Lim, C.H., Rasti, B., Sulistyo, J., and Hamid, M.A., Heliyon, 2021, vol. 7, p. e06305. https://doi.org/10.1016/j.heliyon.2021.e06305
Saini, K., Pathak, V.M., Tyagi, A., and Gupta, R., Catal. Res., 2022, vol. 2, no. 3. https://doi.org/10.21926/cr.2203029
Zhao, F., Li, Y., Li, C., Ban, X., Gu, Z., and Li, Z., Food Hydrocolloids, 2022, vol. 133, no. 1, p. 107951. https://doi.org/10.1016/j.foodhyd.2022.107951
Zhou, J., Feng, Z., Liu, S., Wei, F., Shi, Y., Zhao, L., et al., Mol. Plant Pathol., 2021, vol. 22, no. 1, pp. 130–144. https://doi.org/10.1111/mpp.13014
Biwer, A., Antranikian, G., and Heinzle, E., Appl. Microbiol. Biotechnol., 2002, vol. 59, no. 6, pp. 609–617. https://doi.org/10.1007/s00253-002-1057-x
Zheng, M., Endo, T., and Zimmermann, W., Aust. J. Chem., 2002, vol. 55, no. 2, pp. 39–48. https://doi.org/10.1071/CH01189
Li, C., Ahn, H.J., Kim, J.H., and Kim, Y.W., Carbohydr. Res., 2014, vol. 99, pp. 39–46. https://doi.org/10.1016/j.carbpol.2013.08.056
Saini, K., Pathak, V.M., Tyagi, A., and Gupta, R., Catal. Res., 2022, vol. 2, no. 3: 029, pp. 1–56. https://doi.org/10.21926/cr.2203029
Melentiev, A.I., Galimzianova, N.F., Gilvanova, E.A., Shchelchkova, E.A., Kuzmina, L.Yu., Boyko, T.F., et al., Adv. Microbiol., 2014, vol. 4, no. 8, pp. 455–464. https://doi.org/10.4236/aim.2014.48050
Gupta, R.S., Patel, S., Saini, N., and Chen, S., Int. J. Syst. Evol. Microbiol., 2020, vol. 70, no. 11, pp. 5753–5798. https://doi.org/10.1099/ijsem.0.004475
Yoon, S.H., Ha, S.M., Kwon, S., Lim, J., Kim, Y., Seo, H., et al., Int. J. Syst. Evol. Microbiol., 2017, vol. 67, no. 5, pp. 1613–1617. https://doi.org/10.1099/ijsem.0.001755
Adebule, A.P., J. Adv. Med. Life Sci., 2018, vol. 6, no. 3, pp. 1–3. https://doi.org/10.5281/zenodo.1198928
Kumar, S., Stecher, G., Li, M., Knyaz, C., and Tamura, K., Mol. Biol. Evol., 2018, vol. 35, no. 6, pp. 1547–1549. https://doi.org/10.1093/molbev/msy096
Felsenstein, J., Evolution, 1985, vol. 39, no. 4, pp. 783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Martins, R.F. and Hatti-Kaul, R., Enzyme Microb. Technol., 2002, vol. 30, no. 1, pp. 116–124. https://doi.org/10.1016/S0141-0229(01)00461-6
Usanov, N.G., Gil’vanova, E.A., Eli’zarev, P.A., Prutsakova, E.A., and Melent’ev, A.I., Appl. Biochem. Microbiol., 2007, vol. 43, no. 1, pp. 105–110. https://doi.org/10.1134/S000368380701019X
Tilden, E.B. and Hudson, G.S., J. Bacteriol., 1942, vol. 43, no. 4, pp. 527–544. https://doi.org/10.1128/jb.43.4.527-544.1942
Neuhoff, V., Arold, N., Taube, D., and Ehrhardt, W., Electrophoresis, 1988, vol. 9, no. 6, pp. 255–262. https://doi.org/10.1002/elps.1150090603
Dawson, R., Elliott, D., Elliott, W., and Jones, K., Data for Biochemical Research, 3rd ed., Oxford: Clarendon, 1986.
Nogi, Y., Takami, H., and Horikoshi, K., Int. J. Syst. Evol. Microbiol., 2005, vol. 55, no. 6, pp. 2309–2315. https://doi.org/10.1099/ijs.0.63649-0
Xue, Y., Zhang, X., Zhou, C., Zhao, Y., Cowan, D.A., Heaphy, S., et al., Int. J. Syst. Evol. Microbiol., 2006, vol. 56, no. 6, pp. 1217–1221. https://doi.org/10.1099/ijs.0.64105-0
Zhao, W., Zhang, C.L., Romanek, C.S., and Wiegel, J., Int. J. Syst. Evol. Microbiol., 2008, vol. 58, no. 5, pp. 1106–1108. https://doi.org/10.1099/ijs.0.65363-0
de Jong, S.I., Broek, M.A., Merkel, A.Y., de la Torre, CortesP., Kalamorz, F., Cook, G.M., et al., Extremophiles, 2020, vol. 24, no. 6, pp. 923–935. https://doi.org/10.1007/s00792-020-01205-w
Yampayont, P., Iizuka, M., Ito, K., and Limpaseni, T., J. Incl. Phenom. Macrocycl. Chem., 2006, vol. 56, nos. 1–2, pp. 203–207. https://doi.org/10.1007/s10847-006-9084-3
Alves-Prado, H.F., Carneiro, A.A.J., Pavezzi, F.C., Gomes, E., Boscolo, M., Franco, C.M.L., et al., Appl. Biochem. Biotechnol., 2008, vol. 146, nos. 1–3, pp. 3–13. https://doi.org/10.1007/s12010-007-8093-z
Savergave, L.S., Dhule, S.S., Jogdand, V.V., Nene, S.N., and Gadre, R.V., Biochem. Eng. J., 2008, vol. 39, no. 3, pp. 510–515. https://doi.org/10.1016/j.bej.2007.09.020
More, S.S., Niraja, R., Evelyn, C., Byadgi, A.M., Shweta, V., and Mangaraj, S.D., Croat. J. Food Technol. Biotechnol. Nutr., 2012, vol. 7, nos. 1–2, pp. 90–97.
Reddy, S.V., More, S.S., and Annappa, G.S., J. Basic Microbiol., 2017, vol. 57, no. 11, pp. 974–981. https://doi.org/10.1002/jobm.201700270
Atanasova, N., Kitayska, T., Bojadjieva, I., Yankov, D., and Tonkova, A., Process Biochem., 2011, vol. 46, no. 1, pp. 116–122. https://doi.org/10.1016/j.procbio.2010.07.027
Cao, X., Jin, Z., Chen, F., and Wang, X., J. Food Biochem., 2005, vol. 28, no. 6, pp. 463–475. https://doi.org/10.1111/j.1745-4514.2004.04603.x
Kitayska, T., Petrova, P., Ivanova, V., and Tonkova, A., Appl. Biochem. Biotechnol., 2011, vol. 165, nos. 5–6, pp. 1285–1295. https://doi.org/10.1007/s12010-011-9346-4
Fujita, Y., Tsubouchi, H., Inagi, Y., Tomita, K., Ozaki, A., and Nakanishi, K., J. Ferment. Bioeng., 1990, vol. 70, no. 3, pp. 150–154. https://doi.org/10.1016/0922-338X(90)90174-U
Yim, D.G., Sato, H.H., Park, Y.H., and Park, Y.K., J. Ind. Microbiol. Biotechnol., 1997, vol. 18, no. 6, pp. 402–405. https://doi.org/10.1038/sj.jim.2900400
Higuti, I.H., Grande, S.W., Sacco, R., and Jose do Nascimento, A., Braz. Arch. Biol. Technol., 2003, vol. 46, no. 2, pp. 183–186. https://doi.org/10.1590/S1516-89132003000200007
Li, C., Chen, S., Gu, Z., Hong, Y., Cheng, L., and Li, Z., Food Biosci., 2018, vol. 26, pp. 139–144. https://doi.org/10.1016/j.fbio.2018.10.006
Chung, H-J., Yoon, S-H., Kim, M-J., Kweon, K-S., Lee, I-W., Kim, J-W., et al., J. Agric. Food Chem., 1998, vol. 46, no. 3, pp. 952–959. https://doi.org/10.1021/jf970707d
Jia, X., Ye, X., Chen, J., Lin, X., Vasseur, L., and You, M., Starch–Starke, 2017, vol. 70, nos. 1–2. https://doi.org/10.1002/star.201700016
ACKNOWLEDGMENTS
During the research, the equipment of the “Agidel” Central Common Use of the Ufa Federal Research Center of the Russian Academy of Sciences was used.
Funding
The work was carried out within the framework of the state assignment of the Ministry of Education and Science of the Russian Federation on topic No. 220131100163-4 “Interspecific interactions in microbial communities and plant-microbial associations of natural and technogenic ecosystems (genetic, biochemical and biotechnological aspects)”.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflicts of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Milman, P.Y., Gilvanova, E.A. & Aktuganov, G.E. Cyclodextringlucanotransferase of the Alkalophilic Strain Caldalkalibacillus mannanilyticus IB-OR17-B1. Appl Biochem Microbiol 59, 570–579 (2023). https://doi.org/10.1134/S0003683823050125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683823050125