Abstract
Background
Reconstruction of amino acid sequences from assembled transcriptome is of interest in personalized medicine, for example, to predict drug-target (or protein-protein) interaction considering individual’s genomic variations. Most of the existing transcriptome assemblers, however, seems not well suited for this purpose.
Methods
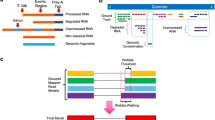
In this work, we present StringFix, an annotation guided transcriptome assembly and protein sequence reconstruction software tool that takes genome-aligned reads and the annotations associated to the reference genome as input. The tool ‘fixes’ the pre-annotated transcript sequence by taking small variations into account, finally to produce possible amino acid sequences that are likely to exist in the test tissue.
Results
The results show that, using outputs from existing reference-based assemblers as the input GTF-guide, StringFix could reconstruct amino acid sequences more precisely with higher sensitivity than direct generation using the recovered transcripts from all the assemblers we tested.
Conclusion
By using StringFix with the existing reference-based assemblers, one can recover not only a novel transcripts and isoforms but also the possible amino acid sequence stemming from them.




Similar content being viewed by others
Data and Software availability
The python code was deposited to PyPI and Github, respectively. Installation instruction, usage and example codes can be found at https://github.com/combio-dku/ (Project name: StringFix, license: GPL 3.0, Operating system(s): Platform independent, Programming language: python 3, other requirements: None). The datasets used in this work can be freely downloaded from the gene expression omnibus (GEO) at https://www.ncbi.nlm.nih.gov/geo/ using their accession number.
References
Adam G et al (2020) Machine learning approaches to drug response prediction: challenges and recent progress. NPJ Precis Oncol 4:19
Ahmadi Moughari F, Eslahchi C (2021) A computational method for drug sensitivity prediction of cancer cell lines based on various molecular information. PLoS ONE 16(4):e0250620
Alser M et al (2021) Technology dictates algorithms: recent developments in read alignment. Genome Biol 22(1):249
Bhatti H et al (2021) Recent advances in biological nanopores for nanopore sequencing, sensing and comparison of functional variations in MspA mutants. RSC Adv 11(46):28996–29014
Camacho C et al (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421
Chang Z et al (2015) Bridger: a new framework for de novo transcriptome assembly using RNA-seq data. Genome Biol 16(1):30
Chin CS et al (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10(6):563–569
Danecek P et al (2021) Twelve years of SAMtools and BCFtools. Gigascience, 10(2)
Dobin A et al (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1):15–21
Emdadi A, Eslahchi C (2020) DSPLMF: a method for Cancer Drug Sensitivity Prediction using a Novel Regularization Approach in Logistic Matrix Factorization. Front Genet 11:75
Feng J, Li W, Jiang T (2011) Inference of isoforms from short sequence reads. J Comput Biol 18(3):305–321
Firtina C et al (2020) Apollo: a sequencing-technology-independent, scalable and accurate assembly polishing algorithm. Bioinformatics 36(12):3669–3679
Fu Y et al (2021) Vulcan: improved long-read mapping and structural variant calling via dual-mode alignment. Gigascience, 10(9)
Gatter T, Stadler PF (2019) Ryuto: network-flow based transcriptome reconstruction. BMC Bioinformatics 20(1):190
Grabherr MG et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29(7):644–652
Griebel T et al (2012) Modelling and simulating generic RNA-Seq experiments with the flux simulator. Nucleic Acids Res 40(20):10073–10083
Guttman M et al (2010) Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol 28(5):503–510
Koren S et al (2017) Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res 27(5):722–736
Li W, Feng J, Jiang T (2011) IsoLasso: a LASSO regression approach to RNA-Seq based transcriptome assembly. J Comput Biol 18(11):1693–1707
Liu R, Dickerson J (2017) Strawberry: fast and accurate genome-guided transcript reconstruction and quantification from RNA-Seq. PLoS Comput Biol 13(11):e1005851
Liu J et al (2016a) TransComb: genome-guided transcriptome assembly via combing junctions in splicing graphs. Genome Biol 17(1):213
Liu J et al (2016b) BinPacker: packing-based De Novo Transcriptome Assembly from RNA-seq data. PLoS Comput Biol 12(2):e1004772
Loman NJ, Quick J, Simpson JT (2015) A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat Methods 12(8):733–735
Maitra RD, Kim J, Dunbar WB (2012) Recent advances in nanopore sequencing. Electrophoresis 33(23):3418–3428
Mao S et al (2020) RefShannon: a genome-guided transcriptome assembler using sparse flow decomposition. PLoS ONE 15(6):e0232946
Martin JA, Wang Z (2011) Next-generation transcriptome assembly. Nat Rev Genet 12(10):671–682
Mir K et al (2012) Predicting statistical properties of open reading frames in bacterial genomes. PLoS ONE 7(9):e45103
Peng Y et al (2013) IDBA-tran: a more robust de novo de bruijn graph assembler for transcriptomes with uneven expression levels. Bioinformatics 29(13):i326–i334
Pertea G, Pertea M (2020) GFF Utilities: GffRead and GffCompare F1000Res, 9
Pertea M et al (2015) StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33(3):290–295
Robertson G et al (2010) De novo assembly and analysis of RNA-seq data. Nat Methods 7(11):909–912
Sachdev K, Gupta MK (2019) A comprehensive review of feature based methods for drug target interaction prediction. J Biomed Inform 93:103159
Schulz MH et al (2012) Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28(8):1086–1092
Song L, Sabunciyan S, Florea L (2016) CLASS2: accurate and efficient splice variant annotation from RNA-seq reads. Nucleic Acids Res 44(10):e98
Stransky N et al (2015) Pharmacogenomic agreement between two cancer cell line data sets. Nature 528(7580):84–
Trapnell C et al (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28(5):511–515
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a Revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63
Wang L et al (2020) Incorporating chemical sub-structures and protein evolutionary information for inferring drug-target interactions. Sci Rep 10(1):6641
Wei D et al (2019) Comprehensive anticancer drug response prediction based on a simple cell line-drug complex network model. BMC Bioinformatics 20(1):44
Xie Y et al (2014) SOAPdenovo-Trans: de novo transcriptome assembly with short RNA-Seq reads. Bioinformatics 30(12):1660–1666
Yoon S et al (2018) TraRECo: a greedy approach based de novo transcriptome assembler with read error correction using consensus matrix. BMC Genomics 19(1):653
Acknowledgements
The authors gratefully acknowledge the Center for Bio-Medical Engineering Core Facility at Dankook University.
Funding
This work was supported by the research fund of Dankook university in 2022.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interest
The authors declare that they have no competing interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, J., Kim, M., Han, K. et al. StringFix: an annotation-guided transcriptome assembler improves the recovery of amino acid sequences from RNA-Seq reads. Genes Genom 45, 1599–1609 (2023). https://doi.org/10.1007/s13258-023-01458-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01458-7