Abstract
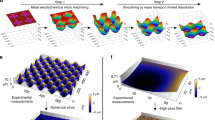
In vitro experiments have shown that cell scale curvatures influence cell migration; cells avoid convex hills and settle in concave valleys. However, it is not known whether dynamic changes in curvature can guide cell migration. This study extends a previous in-silico model to explore the effects over time of changing the substrate curvature on cell migration guidance. By simulating a dynamic surface curvature using traveling wave patterns, we investigate the influence of wave height and speed, and find that long-distance cell migration guidance can be achieved on specific wave patterns. We propose a mechanistic explanation of what we call dynamic curvotaxis and highlight those cellular features that may be involved. Our results open a new area of study for understanding cell mobility in dynamic environments, from single-cell in vitro experiments to multi-cellular in vivo mechanisms.







Similar content being viewed by others
Data availability
The datasets related to the codes that we used to generate the cell model and to launch cell migration simulations for the current study are available via the following persistent web link: https://amubox.univ-amu.fr/s/bKiWG6r87nazLP8. The large datasets of computational results that we obtained during the simulations and we analyzed for the current study are available from the corresponding author on reasonable request.
References
Arima S, Nishiyama K, Ko T, Arima Y, Hakozaki Y, Sugihara K, Koseki H, Uchijima Y, Kurihara Y, Kurihara H (2011) Angiogenic morphogenesis driven by dynamic and heterogeneous collective endothelial cell movement. Dev Camb Engl 138:4763–4776. https://doi.org/10.1242/dev.068023
Bailles A, Collinet C, Philippe J-M, Lenne P-F, Munro E, Lecuit T (2019) Genetic induction and mechano-chemical propagation of a morphogenetic wave. Nature 572:467–473. https://doi.org/10.1038/s41586-019-1492-9
Caballero D, Voituriez R, Riveline D (2015a) The cell ratchet: Interplay between efficient protrusions and adhesion determines cell motion. Cell Adhes Migr 9:327–334. https://doi.org/10.1080/19336918.2015.1061865
Caballero D, Comelles J, Piel M, Voituriez R, Riveline D (2015b) Ratchetaxis: long-range directed cell migration by local cues. Trends Cell Biol 25:815–827. https://doi.org/10.1016/j.tcb.2015.10.009
Callens SJP, Uyttendaele RJC, Fratila-Apachitei LE, Zadpoor AA (2020) Substrate curvature as a cue to guide spatiotemporal cell and tissue organization. Biomaterials 232:119739. https://doi.org/10.1016/j.biomaterials.2019.119739
Chandorkar Y, Castro Nava A, Schweizerhof S, Van Dongen M, Haraszti T, Köhler J, Zhang H, Windoffer R, Mourran A, Möller M, De Laporte L (2019) Cellular responses to beating hydrogels to investigate mechanotransduction. Nat Commun 10:4027. https://doi.org/10.1038/s41467-019-11475-4
Chelberg MK, Tsilibary EC, McCarthy HAR, JB, (1989) Type 1V collagen-mediated melanoma cell adhesion and migration: involvement of multiple distinct domains of the collagen molecule. Cancer Res 49:4796–4802
Chevalier NR (2018) The first digestive movements in the embryo are mediated by mechanosensitive smooth muscle calcium waves. Philos Trans r Soc B Biol Sci 373:20170322. https://doi.org/10.1098/rstb.2017.0322
Davidson PM, Cadot B (2021) Actin on and around the nucleus. Trends Cell Biol 31:211–223. https://doi.org/10.1016/j.tcb.2020.11.009
Deguchi S, Ohashi T, Sato M (2006) Tensile properties of single stress fibers isolated from cultured vascular smooth muscle cells. J Biomech 39:2603–2610. https://doi.org/10.1016/j.jbiomech.2005.08.026
Devreotes P, Janetopoulos C (2003) Eukaryotic chemotaxis: distinctions between directional sensing and polarization. J Biol Chem 278(23):20445–20448. https://doi.org/10.1074/jbc.R300010200
Dokukina IV, Gracheva ME (2010) A model of fibroblast motility on substrates with different rigidities. Biophys J 98:2794–2803. https://doi.org/10.1016/j.bpj.2010.03.026
Dubois F, Jean M (2006) The non smooth contact dynamic method: recent LMGC90 software developments and application. In: Wriggers P, Nackenhorst U (eds) Analysis and simulation of contact problems. Springer, Berlin, pp 375–378
Gittes F, Mickey B, Nettleton J, Howard J (1993) Flexural rigidity of microtubules and actin filaments measured from thermal fluctuations in shape. J Cell Biol 120:923–934. https://doi.org/10.1083/jcb.120.4.923
Han SJ, Bielawski KS, Ting LH, Rodriguez ML, Sniadecki NJ (2012) Decoupling substrate stiffness, spread area, and micropost density: a close spatial relationship between traction forces and focal adhesions. Biophys J 103:640–648. https://doi.org/10.1016/j.bpj.2012.07.023
Harmand N, Huang A, Hénon S (2021) 3D shape of epithelial cells on curved substrates. Phys Rev X 11:031028. https://doi.org/10.1103/PhysRevX.11.031028
Hersen P, Ladoux B (2011) Push it, pull it. Nature 470:340–341. https://doi.org/10.1038/470340a
Isenberg BC, DiMilla PA, Walker M, Kim S, Wong JY (2009) Vascular smooth muscle cell durotaxis depends on substrate stiffness gradient strength. Biophys J 97(5):1313–1322. https://doi.org/10.1016/j.bpj.2009.06.021
Jaffe LF, Créton R (1998) On the conservation of calcium wave speeds. Cell Calcium 24:1–8. https://doi.org/10.1016/S0143-4160(98)90083-5
Jaffe LF (2008) Calcium waves. Philos Trans r Soc B Biol Sci 363:1311–1317. https://doi.org/10.1098/rstb.2007.2249
Jean M (1999) The non-smooth contact dynamics method. Comput Methods Appl Mech Eng 177:235–257. https://doi.org/10.1016/S0045-7825(98)00383-1
Jiang X, Bruzewicz DA, Wong AP, Piel M, Whitesides GM (2005) Directing cell migration with asymmetric micropatterns. PNAS 102(4):975–978. https://doi.org/10.1073/pnas.0408954102
Kishi K, Onuma TA, Nishida H (2014) Long-distance cell migration during larval development in the appendicularian, Oikopleura dioica. Dev Biol 395:299–306. https://doi.org/10.1016/j.ydbio.2014.09.006
le Digabel J, Biais N, Fresnais J, Berret J-F, Hersen P, Ladoux B (2011) Magnetic micropillars as a tool to govern substrate deformations. Lab Chip 11:2630–2636. https://doi.org/10.1039/c1lc20263d
Lo CM, Wang HB, Dembo M, Wang YI (2000) Cell movement is guided by the rigidity of the substrate. Biophys J 79(1):144–152. https://doi.org/10.1016/S0006-3495(00)76279-5
Löber J, Ziebert F, Aranson IS (2014) Modeling crawling cell movement on soft engineered substrates. Soft Matter 10(9):1365–1373
Luxton GWG, Gomes ER, Folker ES, Vintinner E, Gundersen GG (2010) Linear arrays of nuclear envelope proteins harness retrograde actin flow for nuclear movement. Science 329:956–959. https://doi.org/10.1126/science.1189072
Malheiro V, Lehner F, Dinca V, Hoffmann P, Maniura-Weber K (2016) Convex and concave micro-structured silicone controls the shape, but not the polarization state of human macrophages. Biomater Sci 4:1562–1573. https://doi.org/10.1039/C6BM00425C
Marzban B, Kang J, Li N, Sun Y, Yuan H (2019) A contraction–reaction–diffusion model: integrating biomechanics and biochemistry in cell migration. Extreme Mech Lett 32:100566. https://doi.org/10.1016/j.eml.2019.100566
Mayor R, Etienne-Manneville S (2016) The front and rear of collective cell migration. Nat Rev Mol Cell Biol 17:97–109. https://doi.org/10.1038/nrm.2015.14
Milan J-L, Manifacier I, Beussman KM, Han SJ, Sniadecki NJ, About I, Chabrand P (2016) In silico CDM model sheds light on force transmission in cell from focal adhesions to nucleus. J Biomech. https://doi.org/10.1016/j.jbiomech.2016.05.031
Otrock ZK, Mahfouz RAR, Makarem JA, Shamseddine AI (2007) Understanding the biology of angiogenesis: Review of the most important molecular mechanisms. Blood Cells Mol Dis 39:212–220. https://doi.org/10.1016/j.bcmd.2007.04.001
Patan S (2004) Vasculogenesis and Angiogenesis. In: Kirsch M, Black PMCL (eds) Angiogenesis Brain Tumors. Springer, Boston, pp 3–32. https://doi.org/10.1007/978-1-4419-8871-3_1
Park JS, Kim DH, Levchenko A (2017) Topotaxis: a new mechanism of directed cell migration in topographic ECM gradients. Biophys J 114(6):1257–1263. https://doi.org/10.1016/j.bpj.2017.11.3813
Pieuchot L, Marteau J, Guignandon A, Dos Santos T, Brigaud I, Chauvy P-F, Cloatre T, Ponche A, Petithory T, Rougerie P, Vassaux M, Milan J-L, Tusamda Wakhloo N, Spangenberg A, Bigerelle M, Anselme K (2018) Curvotaxis directs cell migration through cell-scale curvature landscapes. Nat Commun 9:3995. https://doi.org/10.1038/s41467-018-06494-6
Raghavan S, Desai RA, Kwon Y, Mrksich M, Chen CS (2010) Micropatterned dynamically adhesive substrates for cell migration. Langmuir 26:17733–17738. https://doi.org/10.1021/la102955m
Reversat A, Gaertner F, Merrin J et al (2020) Cellular locomotion using environmental topography. Nature 582:582–585. https://doi.org/10.1038/s41586-020-2283-z
Ribeiro AJS, Denisin AK, Wilson RE, Pruitt BL (2016) For whom the cells pull: hydrogel and micropost devices for measuring traction forces. Methods. https://doi.org/10.1016/j.ymeth.2015.08.005
Rougerie P, Pieuchot L, dos Santos RS, Marteau J, Bigerelle M, Chauvy P-F, Farina M, Anselme K (2020) Topographical curvature is sufficient to control epithelium elongation. Sci Rep 10:14784. https://doi.org/10.1038/s41598-020-70907-0
Scarpa E, Mayor R (2016) Collective cell migration in development. J Cell Biol 212:143–155. https://doi.org/10.1083/jcb.201508047
Schamberger B, Ziege R, Anselme K, Ben Amar M, Bykowski M, Castro AP, Cipitria A, Coles RA, Dimova R, Eder M, Ehrig S (2023) Curvature in biological systems: its quantification, emergence and implications across the scales. Adv Mater. https://doi.org/10.1002/adma.202206110
SenGupta S, Parent CA, Bear JE (2021) The principles of directed cell migration. Nat Rev Mol Cell Biol 22:529–547. https://doi.org/10.1038/s41580-021-00366-6
Shellard A, Mayor R (2020) All roads lead to directional cell migration. Trends Cell Biol 30:852–868. https://doi.org/10.1016/j.tcb.2020.08.002
Shellard A, Szabó A, Trepat X, Mayor R (2018) Supracellular contraction at the rear of neural crest cell groups drives collective chemotaxis. Science 362:339–343. https://doi.org/10.1126/science.aau3301
Shellard A, Mayor R (2019) Supracellular migration – beyond collective cell migration. J Cell Sci. https://doi.org/10.1242/jcs.226142
Sokolow A, Toyama Y, Kiehart DP, Edwards GS (2012) Cell ingression and apical shape oscillations during dorsal closure in drosophila. Biophys J 102:969–979. https://doi.org/10.1016/j.bpj.2012.01.027
Szabó A, Melchionda M, Nastasi G, Woods ML, Campo S, Perris R, Mayor R (2016) In vivo confinement promotes collective migration of neural crest cells. J Cell Biol 213:543–555. https://doi.org/10.1083/jcb.201602083
Theveneau E, Mayor R (2012) Neural crest migration: interplay between chemorepellents, chemoattractants, contact inhibition, epithelial–mesenchymal transition, and collective cell migration, WIREs. Dev Biol 1:435–445. https://doi.org/10.1002/wdev.28
Tomba C, Petithory T, Pedron R, Airoudj A, Di Meglio I, Roux A, Luchnikov V (2019) In laser-assisted strain engineering of thin elastomer films to form variable wavy substrates for cell culture. Small 15(21):e1900162. https://doi.org/10.1002/smll.201900162
Tjhung E, Tiribocchi A, Marenduzzo D, Cates ME (2015) A minimal physical model captures the shapes of crawling cells. Nat Commun 6:5420. https://doi.org/10.1038/ncomms6420
Vassaux M, Pieuchot L, Anselme K, Bigerelle M, Milan J-L (2019) A biophysical model for curvature-guided cell migration. Biophys J 117:1136–1144. https://doi.org/10.1016/j.bpj.2019.07.022
Maxime Vassaux (2018) mvassaux/adhSC: Initial release of the adhesion cell model (v1.0.0). Zenodo. https://doi.org/10.5281/zenodo.1187087
Wakhloo NT, Anders S, Badique F, Eichhorn M, Brigaud I, Petithory T, Vassaux M, Milan JL, Freund JN, Rühe J, Davidson PM, Pieuchot L, Anselme K (2020) Actomyosin, vimentin and LINC complex pull on osteosarcoma nuclei to deform on micropillar topography. Biomaterials 234:119746
Winkler B, Aranson IS, Ziebert F (2019) Confinement and substrate topography control cell migration in a 3D computational model. Commun Phys 2:1–11. https://doi.org/10.1038/s42005-019-0185-x
Werner M, Petersen A, Kurniawan NA, Bouten CVC (2019) Cell-perceived substrate curvature dynamically coordinates the direction, speed, and persistence of stromal cell migration. Adv Biosyst 3:1900080. https://doi.org/10.1002/adbi.201900080
Zhang J, Wang Yl (2017) Centrosome defines the rear of cells during mesenchymal migration. Mol Biol Cell 28(23):3240–3251
Funding
Agence Nationale de la Recherche, ANR-22-CE45-0010. A CC-BY public copyright license has been applied by the authors to the Author Accepted Manuscript, in accordance with the grant’s open access conditions.
Author information
Authors and Affiliations
Contributions
IM, JLM, wrote the main manuscript text with substantial contributions from GC, MV, LP, VL and KA. IM, JLM, MV, GC and DL prepared and launched the in-silico simulations. IM prepared Fig. 1 and JLM the others. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (AVI 30498 KB)
Supplementary file2 (AVI 6110 KB)
Supplementary file3 (MP4 2533 KB)
Supplementary file4 (MP4 684 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Manifacier, I., Carlin, G., Liu, D. et al. In silico analysis shows that dynamic changes in curvature guide cell migration over long distances. Biomech Model Mechanobiol 23, 315–333 (2024). https://doi.org/10.1007/s10237-023-01777-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-023-01777-4