Abstract
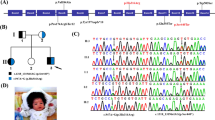
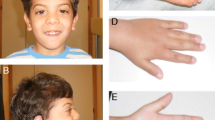
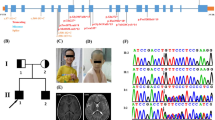
Sphingomyelin phosphodiesterase 4 (SMPD4) encodes a member of the Mg2+-dependent, neutral sphingomyelinase family that catalyzes the hydrolysis of the phosphodiester bond of sphingomyelin to form phosphorylcholine and ceramide. Recent studies have revealed that biallelic loss-of-function variants of SMPD4 cause syndromic neurodevelopmental disorders characterized by microcephaly, congenital arthrogryposis, and structural brain anomalies. In this study, three novel loss-of-function SMPD4 variants were identified using exome sequencing (ES) in two independent patients with developmental delays, microcephaly, seizures, and brain structural abnormalities. Patient 1 had a homozygous c.740_741del, p.(Val247Glufs*21) variant and showed profound intellectual disability, hepatomegaly, a simplified gyral pattern, and a thin corpus callosum without congenital dysmorphic features. Patient 2 had a compound heterozygous nonsense c.2124_2125del, p.(Phe709*) variant and splice site c.1188+2dup variant. RNA analysis revealed that the c.1188+2dup variant caused exon 13 skipping, leading to a frameshift (p.Ala406Ser*6). In vitro transcription analysis using minigene system suggested that mRNA transcribed from mutant allele may be degraded by nonsense-mediated mRNA decay system. He exhibited diverse manifestations, including growth defects, muscle hypotonia, respiratory distress, arthrogryposis, insulin-dependent diabetes mellitus, sensorineural hearing loss, facial dysmorphism, and various brain abnormalities, including cerebral atrophy, hypomyelination, and cerebellar hypoplasia. Here, we review previous literatures and discuss the phenotypic diversity of SMPD4-related disorders.



Similar content being viewed by others
Data Availability
The exome sequencing data of this study is not publicly available.
References
Wu BX, Clarke CJ, Hannun YA (2010) Mammalian neutral sphingomyelinases: regulation and roles in cell signaling responses. Neuromolecular Med 12(4):320–330. https://doi.org/10.1007/s12017-010-8120-z
Krut O, Wiegmann K, Kashkar H, Yazdanpanah B, Kronke M (2006) Novel tumor necrosis factor-responsive mammalian neutral sphingomyelinase-3 is a C-tail-anchored protein. J Biol Chem 281(19):13784–13793. https://doi.org/10.1074/jbc.M511306200
Peeters BWA, Piet ACA, Fornerod M (2022) Generating membrane curvature at the nuclear pore: a lipid point of view. Cells-Basel 11(3):469. https://doi.org/10.3390/cells11030469
Cheng LC, Baboo S, Lindsay C, Brusman L, Martinez-Bartolome S, Tapia O et al (2019) Identification of new transmembrane proteins concentrated at the nuclear envelope using organellar proteomics of mesenchymal cells. Nucleus-Phila 10(1):126–143. https://doi.org/10.1080/19491034.2019.1618175
Atilla-Gokcumen GE, Muro E, Relat-Goberna J, Sasse S, Bedigian A, Coughlin ML et al (2014) Dividing cells regulate their lipid composition and localization. Cell 156(3):428–439. https://doi.org/10.1016/j.cell.2013.12.015
Corcoran CA, He Q, Ponnusamy S, Ogretmen B, Huang Y, Sheikh MS (2008) Neutral sphingomyelinase-3 is a DNA damage and nongenotoxic stress-regulated gene that is deregulated in human malignancies. Mol Cancer Res 6(5):795–807. https://doi.org/10.1158/1541-7786.MCR-07-2097
Magini P, Smits DJ, Vandervore L, Schot R, Columbaro M, Kasteleijn E et al (2019) Loss of SMPD4 causes a developmental disorder characterized by microcephaly and congenital arthrogryposis. Am J Hum Genet. 105(4):689–705. https://doi.org/10.1016/j.ajhg.2019.08.006
Bijarnia-Mahay S, Somashekar PH, Kaur P, Kulshrestha S, Ramprasad VL, Murugan S et al (2021) Growth and neurodevelopmental disorder with arthrogryposis, microcephaly and structural brain anomalies caused by bi-allelic partial deletion of SMPD4 gene. J Hum Genet. https://doi.org/10.1038/s10038-021-00981-3
Yamada M, Suzuki H, Shima T, Uehara T, Kosaki K (2022) A patient with compound heterozygosity of SMPD4: another example of utility of exome-based copy number analysis in autosomal recessive disorders. Am J Med Genet A 188(2):613–617. https://doi.org/10.1002/ajmg.a.62535
Ravenscroft G, Clayton JS, Faiz F, Sivadorai P, Milnes D, Cincotta R et al (2021) Neurogenetic fetal akinesia and arthrogryposis: genetics, expanding genotype-phenotypes and functional genomics. J Med Genet 58(9):609–618. https://doi.org/10.1136/jmedgenet-2020-106901
Ji W, Kong X, Yin H, Xu J, Wang X (2022) Case report: novel biallelic null variants of SMPD4 confirm its involvement in neurodevelopmental disorder with microcephaly, arthrogryposis, and structural brain anomalies. Front Genet 13:872264. https://doi.org/10.3389/fgene.2022.872264
Smits DJ, Schot R, Krusy N, Wiegmann K, Utermohlen O, Mulder MT et al (2023) SMPD4 regulates mitotic nuclear envelope dynamics and its loss causes microcephaly and diabetes. Brain. https://doi.org/10.1093/brain/awad033
Monies D, Abouelhoda M, Assoum M, Moghrabi N, Rafiullah R, Almontashiri N et al (2019) Lessons learned from large-scale, first-tier clinical exome sequencing in a highly consanguineous population. Am J Hum Genet 105(4):879. https://doi.org/10.1016/j.ajhg.2019.09.019
Theresia KJ, Wolfgang H, Gundula G, Michael E, Alexander W, Caroline G et al (2023) Prenatal diagnosis of SMPD4 loss - a neurodevelopmental disorder with microcephaly, arthrogryposis and structural brain anomalies. Prenat Diagn 43(3):284–287. https://doi.org/10.1002/pd.6324
Watanabe K, Nakashima M, Kumada S, Mashimo H, Enokizono M, Yamada K et al (2021) Identification of two novel de novo TUBB variants in cases with brain malformations: case reports and literature review. J Hum Genet 66(12):1193–1197. https://doi.org/10.1038/s10038-021-00956-4
Miyamoto S, Nakashima M, Ohashi T, Hiraide T, Kurosawa K, Yamamoto T et al (2019) A case of de novo splice site variant in SLC35A2 showing developmental delays, spastic paraplegia, and delayed myelination. Mol Genet Genomic Med 7(8):e814. https://doi.org/10.1002/mgg3.814
Olsen ASB, Faergeman NJ (2017) Sphingolipids: membrane microdomains in brain development, function and neurological diseases. Open Biol 7(5). https://doi.org/10.1098/rsob.170069
Hussain G, Wang J, Rasul A, Anwar H, Imran A, Qasim M et al (2019) Role of cholesterol and sphingolipids in brain development and neurological diseases. Lipids Health Dis 18(1):26. https://doi.org/10.1186/s12944-019-0965-z
Ferreira LF, Moylan JS, Gilliam LA, Smith JD, Nikolova-Karakashian M, Reid MB (2010) Sphingomyelinase stimulates oxidant signaling to weaken skeletal muscle and promote fatigue. Am J Physiol Cell Physiol 299(3):C552–C560. https://doi.org/10.1152/ajpcell.00065.2010
De Larichaudy J, Zufferli A, Serra F, Isidori AM, Naro F, Dessalle K et al (2012) TNF-alpha- and tumor-induced skeletal muscle atrophy involves sphingolipid metabolism. Skelet Muscle 2(1):2. https://doi.org/10.1186/2044-5040-2-2
Cowart LA (2010) A novel role for sphingolipid metabolism in oxidant-mediated skeletal muscle fatigue. Focus on “Sphingomyelinase stimulates oxidant signaling to weaken skeletal muscle and promote fatigue”. Am J Physiol Cell Physiol 299(3):C549–C551. https://doi.org/10.1152/ajpcell.00236.2010
Moylan JS, Smith JD, Wolf Horrell EM, McLean JB, Deevska GM, Bonnell MR et al (2014) Neutral sphingomyelinase-3 mediates TNF-stimulated oxidant activity in skeletal muscle. Redox Biol 2:910–920. https://doi.org/10.1016/j.redox.2014.07.006
Chaurasia B, Summers SA (2015) Ceramides - lipotoxic inducers of metabolic disorders. Trends Endocrinol Metab 26(10):538–550. https://doi.org/10.1016/j.tem.2015.07.006
Borodzicz S, Czarzasta K, Kuch M, Cudnoch-Jedrzejewska A (2015) Sphingolipids in cardiovascular diseases and metabolic disorders. Lipids Health Dis 14:55. https://doi.org/10.1186/s12944-015-0053-y
Coblentz PD, Ahn B, Hayward LF, Yoo JK, Christou DD, Ferreira LF (2019) Small-hairpin RNA and pharmacological targeting of neutral sphingomyelinase prevent diaphragm weakness in rats with heart failure and reduced ejection fraction. Am J Physiol Lung Cell Mol Physiol 316(4):L679–LL90. https://doi.org/10.1152/ajplung.00516.2018
Zhang C, Qiao S, Wu J, Xu W, Ma S, Zhao B et al (2021) A new insulin-sensitive enhancer from Silene viscidula, WPTS, treats type 2 diabetes by ameliorating insulin resistance, reducing dyslipidemia, and promoting proliferation of islet beta cells. Pharmacol Res 165:105416. https://doi.org/10.1016/j.phrs.2020.105416
Schuchman EH, Desnick RJ (2017) Types A and B Niemann-Pick disease. Mol Genet Metab 120(1-2):27–33. https://doi.org/10.1016/j.ymgme.2016.12.008
Hwang SY, Kim TH, Lee HH (2015) Neutral sphingomyelinase and breast cancer research. J Menopausal Med 21(1):24–27. https://doi.org/10.6118/jmm.2015.21.1.24
Hofmann K, Tomiuk S, Wolff G, Stoffel W (2000) Cloning and characterization of the mammalian brain-specific, Mg2+-dependent neutral sphingomyelinase. Proc Natl Acad Sci U S A 97(11):5895–5900. https://doi.org/10.1073/pnas.97.11.5895
Clarke CJ, Wu BX, Hannun YA (2011) The neutral sphingomyelinase family: identifying biochemical connections. Adv Enzyme Regul 51(1):51–58. https://doi.org/10.1016/j.advenzreg.2010.09.016
Tomiuk S, Zumbansen M, Stoffel W (2000) Characterization and subcellular localization of murine and human magnesium-dependent neutral sphingomyelinase. J Biol Chem 275(8):5710–5717. https://doi.org/10.1074/jbc.275.8.5710
Acknowledgements
We would like to thank the patient’s family for participating in this work.
Funding
This study was funded by the Japan Society for the Promotion of Science, KAKENHI (Grant number JP 20H03641 and 23H02875, JP20K08236 and JP21K06819), the Japan Agency for Medical Research and Development (AMED) (JP23ek0109549 and 23ek01099674, and 23ek0109637), the Takeda Science Foundation, and HUSM Grant-in-Aid from Hamamatsu University School of Medicine.
Author information
Authors and Affiliations
Contributions
HS and MK contributed to the conception and design of the study. Material preparation, the acquisition, and analysis of ES data were performed by SA, KW, and MN. Medical examinations were carried out by YK, KK, and EK. SA, MK, MN, and HS contributed to the drafting of the text and preparing the figures. All authors revised and commented on the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Experimental protocols were approved by the Institutional Review Board of Hamamatsu University School of Medicine and Showa University Faculty of Medicine. Written informed consent was obtained from all participating families.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aoki, S., Watanabe, K., Kato, M. et al. Two novel cases of biallelic SMPD4 variants with brain structural abnormalities. Neurogenetics 25, 3–11 (2024). https://doi.org/10.1007/s10048-023-00737-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-023-00737-5