Abstract
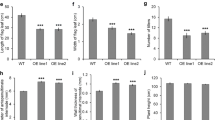
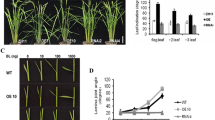
Pre-harvest sprouting (PHS) frequently occurs in rice due to the long spells of rainy weather, and causes severe yield loss and grain quality decrease. Here, we identified one PHS-related gene OsCNX1 cloned from rice PHS mutant, which encoded a molybdenum cofactor (MoCo) biosynthesis enzyme. Genetic complementation indicated OsCNX1 could rescue the PHS and seedling lethal phenotype of the mutant. Expression pattern showed that OsCNX1 was expressed in rice tissue including seedling shoot, culm, blade, and sheath of flag leaf, young panicle, and the seeds at different development stages. Overexpression of OsCNX1 significantly decreased the plant height, and the seed germination of the dormant seeds harvested from fresh panicles, comparing to the wild type (WT). In addition, 1492 differentially expressed genes (DEGs) were identified between OsCNX1-overexpressed line and WT by RNA-sequencing, which were mainly classified in plant-pathogen interaction, plant hormone signal transduction, and starch/sucrose metabolism. These results showed that OsCNX1 was not only necessary for rice seed germination, but also participated in plant development, indicating that OsCNX1 may be useful in rice breeding of PHS resistance and plant height.





Similar content being viewed by others
References
Chen W, Wang W, Lyu Y, Wu Y, Huang P, Hu S, Wei X, Jiao G, Sheng Z, Tang S (2021) OsVP1 activates Sdr4 expression to control rice seed dormancy via the ABA signaling pathway. Crop Journal 9(1):68–78
Chen Y, Xiang Z, Liu M, Wang S, Zhang L, Cai D, Huang Y, Mao D, Fu J, Chen L (2023) ABA biosynthesis gene OsNCED3 contributes to preharvest sprouting resistance and grain development in rice. Plant Cell Environ 46(4):1384–1401
Counce P, Keisling T, Mitchell A (2000) A uniform, objective, and adaptive system for expressing rice development. Crop Sci 40(2):436–443
Du L, Xu F, Fang J, Gao S, Tang J, Fang S, Wang H, Tong H, Zhang F, Chu J, Wang G, Chu C (2018) Endosperm sugar accumulation caused by mutation of PHS8/ISA1 leads to pre-harvest sprouting in rice. Plant J 95(3):545–556
Fang J, Chu C (2008) Abscisic acid and the pre-harvest sprouting in cereals. Plant Signal Behav 3(12):1046
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415
Gubler F, Millar AA, Jacobsen JV (2005) Dormancy release, ABA and pre-harvest sprouting. Curr Opin Plant Biol 8(2):183–187
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40:D109–D114
Koornneef M, Bentsink L, Hilhorst H (2002) Seed dormancy and germination. Curr Opin Plant Biol 5(1):33–36
Leng P, Yuan B, Guo Y (2014) The role of abscisic acid in fruit ripening and responses to abiotic stress. J Exp Bot 65(16):4577
Li X, Tian X, He M, Liu X, Li Z, Tang J, Mei E, Xu M, Liu Y, Wang Z, Guan Q, Meng W, Fang J, Zhang J, Bu Q (2022) bZIP71 delays flowering by suppressing Ehd1 expression in rice. J Integr Plant Biol 64(7):1352–1363
Liu X, Wang J, Yu Y, Kong L, Liu Y, Liu Z, Li H, Wei P, Liu M, Zhou H, Bu Q, Fang J (2019) Identification and characterization of the rice pre-harvest sprouting mutants involved in molybdenum cofactor biosynthesis. New Phytologist 222(1):275–285
Livak K, Schmittgen T (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4):402–408
Love M, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550
Mao X, Cai T, Olyarchuk J, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21(19):3787–3793
Mendel R, Hänsch R (2002) Molybdoenzymes and molybdenum cofactor in plants. J Exp Bot 53(375):1689–1698
Moullet O, Díaz Bermúdez G, Fossati D, Brabant C, Mascher F, Schori A (2022) Pyramiding wheat pre-harvest sprouting resistance genes in triticale breeding. Mol Breed 42(10):60
Nakamura S (2018) Grain dormancy genes responsible for preventing pre-harvest sprouting in barley and wheat. Breed Sci 68(3):295–304
Oliva M, Farcot E, Vernoux T (2013) Plant hormone signaling during development: insights from computational models. Curr Opin Plant Biol 16(1):19–24
Santner A, Calderon-Villalobos L, Estelle M (2009) Plant hormones are versatile chemical regulators of plant growth. Nat Chem Biol 5(5):301–307
Schwarz G, Mendel R (2006) Molybdenum cofactor biosynthesis and molybdenum enzymes. Annu Rev Plant Biol 57(1):623–647
Sohn S, Pandian S, Kumar T, Zoclanclounon Y, Muthuramalingam P, Shilpha J, Satish L, Ramesh M (2021) Seed dormancy and pre-harvest sprouting in rice-an updated overview. Int J Mol Sci 22(21):11804
Sugimoto K, Takeuchi Y, Ebana K, Miyao A, Hirochika H, Hara N, Ishiyama K, Kobayashi M, Ban Y, Hattori T, Yano M (2010) Molecular cloning of Sdr4, a regulator involved in seed dormancy and domestication of rice. Proc Natl Acad Sci USA 107(13):5792–5797
Tsuji H, Taoka K, Shimamoto K (2011) Regulation of flowering in rice: two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol 14(1):45–52
Verma V, Ravindran P, Kumar P (2016) Plant hormone-mediated regulation of stress responses. BMC Plant Biol 16(1):86
Vetch J, Stougaard R, Martin J, Giroux M (2019) Revealing the genetic mechanisms of pre-harvest sprouting in hexaploid wheat (Triticum aestivum L.). Plant Sci 281:180–185
Wang J, Deng Q, Li Y, Yu Y, Liu X, Han Y, Luo X, Wu X, Ju L, Sun J, Liu A, Fang J (2020a) Transcription factors Rc and OsVP1 Coordinately regulate preharvest sprouting tolerance in red pericarp rice. J Agric Food Chem 68(50):14748–14757
Wang J, Korkmaz U, Guo M, Pipatpongpinyo W, Gu X (2020b) Pyramiding seed dormancy genes to improve resistance of semi-dwarf varieties to pre-harvest sprouting in rice. Mol Breed 40(10):93
Xu F, Tang J, Gao S, Cheng X, Du L, Chu C (2019) Control of rice pre-harvest sprouting by glutaredoxin-mediated abscisic acid signaling. Plant J 100(5):1036–1051
Xu F, Tang J, Wang S, Cheng X, Wang H, Ou S, Gao S, Li B, Qian Y, Gao C, Chu C (2022) Antagonistic control of seed dormancy in rice by two bHLH transcription factors. Nat Genet 54(12):1972–1982
Ye H, Feng J, Zhang L, Zhang J, Mispan M, Cao Z, Beighley D, Yang J, Gu X (2015) Map-based cloning of seed dormancy1-2 identified a gibberellin synthesis gene regulating the development of endosperm-imposed dormancy in rice. Plant Physiol 169(3):2152–2165
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11(2):R14
Zhang C, Yuan W, Zhang Q (2012) RPL1, a gene involved in epigenetic processes regulates phenotypic plasticity in rice. Mol Plant 5(2):482–493
Zhao B, Zhang H, Chen T, Ding L, Zhang L, Ding X, Zhang J, Qian Q, Xiang Y (2022a) Sdr4 dominates pre-harvest sprouting and facilitates adaptation to local climatic condition in Asian cultivated rice. J Integr Plant Biol 64(6):1246–1263
Zhao G, Wang J, Chen X, Sha H, Liu X, Han Y, Qiu G, Zhang F, Fang J (2022b) OsASHL1 and OsASHL2, two members of the COMPASS-like complex, control floral transition and plant development in rice. J Genet Genomics 49(9):870–880
Acknowledgements
We are grateful to Prof. Jun Fang (Northeast Institute of Geography and Agroecology, Chinese Academy of Sciences) for supporting the early research and to Dr. Guangxin Zhao (Jilin Agricultural University) for providing helpful discussions and advisement.
Funding
This work was financially supported by the National Natural Science Foundation of China (31901524) and by the Heilongjiang Provincial Natural Science Foundation of China (YQ2022C008).
Author information
Authors and Affiliations
Contributions
XL and DZ performed most of the experiments; YZ, XL, and SW provided technical assistance. XL designed the research, performed parts of the experiments, and wrote the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, X., Zhang, D., Zhang, Y. et al. The molybdenum cofactor biosynthesis gene, OsCNX1, is essential for seedling development and seed germination in rice. Mol Breeding 43, 77 (2023). https://doi.org/10.1007/s11032-023-01424-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-023-01424-x