Abstract
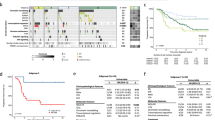
Although immune checkpoint inhibitors have led to durable clinical response in multiple cancers, only a small proportion of patients respond to this treatment. Therefore, we aim to develop a predictive model that utilizes gene mutation profiles to accurately identify the survival of pan-cancer patients with immunotherapy. Here, we develop and evaluate three different nomograms using two cohorts containing 1,594 cancer patients whose mutation profiles are obtained by MSK-IMPACT sequencing and 230 cancer patients receiving whole-exome sequencing, respectively. Using eighteen genes (SETD2, BRAF, NCOA3, LATS1, IL7R, CREBBP, TET1, EPHA7, KDM5C, MET, KMT2D, RET, PAK7, CSF1R, JAK2, FAT1, ASXL1 and SPEN), the first nomogram stratifies patients from both cohorts into High-Risk and Low-Risk groups. Pan-cancer patients in the High-Risk group exhibit significantly shorter overall survival and progression-free survival than patients in the Low-Risk group in both cohorts. Meanwhile, the first nomogram also accurately identifies the survival of patients with melanoma or lung cancer undergoing immunotherapy, or pan-cancer patients treated with anti-PD-1/PD-L1 inhibitor or anti-CTLA-4 inhibitor. The model proposed is not a prognostic model for the survival of pan-cancer patients without immunotherapy, but a simple, effective and robust predictive model for pan-cancer patients’ survival under immunotherapy, and could provide valuable assistance for clinical practice.






Similar content being viewed by others
Data Availability
The datasets of patients treated with ICIs used in our study are downloaded from the Memorial Sloan Kettering Cancer Center (MSKCC) dataset and the Dana-Farber Cancer Institute. The information of MSK non-ICIs patients is also downloaded from the MSKCC dataset. The TCGA dataset is downloaded from the UCSC XENA database. The data used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
Naidoo J, Nishino M, Patel SP, Shankar B, Rekhtman N, Illei P, Camus P (2020) Immune-related pneumonitis after chemoradiotherapy and subsequent immune checkpoint blockade in unresectable stage III non-small-cell Lung Cancer. Clin Lung Cancer 21(5):e435–e444. https://doi.org/10.1016/j.cllc.2020.02.025
Topalian SL, Taube JM, Anders RA, Pardoll DM (2016) Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat Rev Cancer 16:275–287. https://doi.org/10.1038/nrc.2016.36
Hodi FS, O’Day SJ, McDermott DF, Weber RW, Sosman JA, Haanen JB, Gonzalez R, Robert C, Schadendorf D, Hassel JC et al (2010) Improved survival with ipilimumab in patients with metastatic Melanoma. N Engl J Med 363:711–723. https://doi.org/10.1056/NEJMoa1003466
Robert C, Schachter J, Long GV, Arance A, Grob JJ, Mortier L, Daud A, Carlino MS, McNeil C, Lotem M et al (2015) Pembrolizumab versus Ipilimumab in Advanced Melanoma. N Engl J Med 372:2521–2532. https://doi.org/10.1056/NEJMoa1503093
Wolchok JD, Chiarion-Sileni V, Gonzalez R, Rutkowski P, Grob JJ, Cowey CL, Lao CD, Wagstaff J, Schadendorf D, Ferrucci PF et al (2017) Overall survival with combined Nivolumab and Ipilimumab in Advanced Melanoma. N Engl J Med 377:1345–1356. https://doi.org/10.1056/NEJMoa1709684
Reck M, Rodríguez-Abreu D, Robinson AG, Hui R, Csőszi T, Fülöp A, Gottfried M, Peled N, Tafreshi A, Cuffe S et al (2016) Pembrolizumab versus Chemotherapy for PD-L1-Positive non-small-cell Lung Cancer. N Engl J Med 375:1823–1833. https://doi.org/10.1056/NEJMoa1606774
Seiwert TY, Burtness B, Mehra R, Weiss J, Berger R, Eder JP, Heath K, McClanahan T, Lunceford J, Gause C et al (2016) Safety and clinical activity of pembrolizumab for treatment of recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-012): an open-label, multicentre, phase 1b trial. Lancet Oncol 17:956–965. https://doi.org/10.1016/s1470-2045(16)30066-3
Feng Y, Wang Y, Guo K, Feng J, Shao C, Pan M, Ding P, Liu H, Duan H, Lu D et al (2022) The value of WNT5A as prognostic and immunological biomarker in pan-cancer. Annals of Translational Medicine 10:466. https://doi.org/10.21037/atm-22-1317
Patel SP, Kurzrock R (2015) PD-L1 expression as a predictive biomarker in Cancer Immunotherapy. Mol Cancer Ther 14:847–856. https://doi.org/10.1158/1535-7163.mct-14-0983
Bai X, Wu DH, Ma SC, Wang J, Tang XR, Kang S, Fu QJ, Cao CH, Luo HS, Chen YH et al (2020) Development and validation of a genomic mutation signature to predict response to PD-1 inhibitors in non-squamous NSCLC: a multicohort study. Journal for immunotherapy of cancer 8. https://doi.org/10.1136/jitc-2019-000381
Jardim DL, Goodman A, de Melo Gagliato D, Kurzrock R (2021) The challenges of Tumor Mutational Burden as an Immunotherapy Biomarker. Cancer Cell 39:154–173. https://doi.org/10.1016/j.ccell.2020.10.001
Büttner R, Longshore JW, López-Ríos F, Merkelbach-Bruse S, Normanno N, Rouleau E, Penault-Llorca F (2019) Implementing TMB measurement in clinical practice: considerations on assay requirements. ESMO open 4:e000442. https://doi.org/10.1136/esmoopen-2018-000442
Samstein RM, Lee C-H, Shoushtari AN, Hellmann MD, Shen R, Janjigian YY, Barron DA, Zehir A, Jordan EJ, Omuro A et al (2019) Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet 51:202–206. https://doi.org/10.1038/s41588-018-0312-8
Subbiah V, Solit D, Chan T, Kurzrock R (2020) The FDA approval of pembrolizumab for adult and pediatric patients with Tumor mutational burden (TMB) ≥ 10: a decision centered on empowering patients and their physicians. Ann Oncol 31:1115–1118
Strickler JH, Hanks BA, Khasraw M (2021) Tumor Mutational Burden as a predictor of Immunotherapy Response: is more always better? Clinical cancer research. Official J Am Association Cancer Res 27:1236–1241. https://doi.org/10.1158/1078-0432.ccr-20-3054
Miao D, Margolis CA, Vokes NI, Liu D, Taylor-Weiner A, Wankowicz SM, Adeegbe D, Keliher D, Schilling B, Tracy A et al (2018) Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat Genet 50:1271–1281. https://doi.org/10.1038/s41588-018-0200-2
Xu S, Liu D, Chang T, Wen X, Ma S, Sun G, Wang L, Chen S, Xu Y, Zhang H (2022) Cuproptosis-Associated lncRNA establishes New Prognostic Profile and predicts Immunotherapy Response in Clear Cell Renal Cell Carcinoma. Front Genet 13:938259. https://doi.org/10.3389/fgene.2022.938259
Formica V, Sera F, Cremolini C, Riondino S, Morelli C, Arkenau HT, Roselli M (2022) KRAS and BRAF mutations in stage II and III Colon Cancer: a systematic review and Meta-analysis. J Natl Cancer Inst 114:517–527. https://doi.org/10.1093/jnci/djab190
Liu K, Wang JF, Zhan Y, Kong DL, Wang C (2021) Prognosis model of Colorectal cancer patients based on NOTCH3, KMT2C, and CREBBP mutations. J Gastrointest Oncol 12:79–88. https://doi.org/10.21037/jgo-21-28
Jiao S, Zhang X, Wang D, Fu H, Xia Q (2022) Genetic alteration and their significance on clinical events in small cell Lung Cancer. Cancer Manage Res 14:1493–1505. https://doi.org/10.2147/cmar.s356037
Funding
This study was supported by National Natural Science Foundation of China (project number 81973149, 82304250 and 81773551).
Author information
Authors and Affiliations
Contributions
Study design: Z-LC, LK, CL. Data collection: Z-LC, W-YY. Data analysis and interpretation: Z-LC, W-YY. Writing of the manuscript: Z-LC, W-YY. Revision of the manuscript: CL, W-LY, WM, LS, HJ, J-JX. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Competing interests
The authors declare that they have no conflicts of interest with the publication of the manuscript. This article does not contain any studies with human or animal subjects.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Wang, Y., Wang, L. et al. Identifying survival of pan-cancer patients under immunotherapy using genomic mutation signature with large sample cohorts. J Mol Med 102, 69–79 (2024). https://doi.org/10.1007/s00109-023-02398-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-023-02398-1