Abstract
Epoxides, aziridines, and cyclopropanes are found in various medicinal natural products, including polyketides, terpenes, peptides, and alkaloids. Many classes of biosynthetic enzymes are involved in constructing these ring structures during their biosynthesis. This review summarizes our current knowledge regarding how α-ketoglutarate-dependent nonheme iron enzymes catalyze the formation of epoxides, aziridines, and cyclopropanes in nature, with a focus on enzyme mechanisms.
Similar content being viewed by others
Introduction
Three-membered rings, such as epoxides, aziridines, and cyclopropanes, are ubiquitously found in natural products derived from various organisms, including plants and microorganisms [1]. These natural products often exhibit unique biological activities due to the distinctive chemical properties of the three-membered rings. The inherent large ring strain of these small cycles often accounts for their bioactivities via alkylation reactions when they interact with target molecules. For example, the epoxide-containing natural product fosfomycin serves as an inhibitor of MurA, which is involved in peptidoglycan synthesis, thereby exhibiting a potent antibiotic activity [2,3,4]. This inhibition results from the covalent bonding between fosfomycin and a cystine residue in MurA, triggered by the ring opening of the epoxide moiety [4]. The antitumor aziridine natural product mitomycin C is recognized for its ability to crosslink with DNA at specific sequences, forming covalent bonds following the opening of its aziridine ring [5,6,7]. Furthermore, the unique biological activities may also result from the distinct geometric properties of three-membered rings, such as the coplanarity of their three atoms and the rigidity of their ring conformation [8].
Due to the valuable medicinal properties and pharmaceutical applications of natural products containing three-membered rings, their biosynthetic mechanisms have garnered significant attention [1, 9, 10]. These small rings can be constructed through diverse enzymatic mechanisms, such as intramolecular nucleophilic substitution [11,12,13], atom transfer to alkenes [14], and cation-mediated cyclization [15,16,17]. Recent studies revealed that certain α-ketoglutarate (αKG)-dependent nonheme iron enzymes are also capable of catalyzing oxidative cyclization reactions to produce epoxides, aziridines, and cyclopropanes from the corresponding linear substrates, during the biosynthesis of various classes of natural products. Other αKG-dependent nonheme iron enzymes catalyze a diverse array of oxidative transformations, including hydroxylation, halogenation, and rearrangement reactions by cleaving a C–H bond in the substrate [18,19,20,21,22,23]. Although the mechanisms of the three-membered ring-forming iron enzymes are only partially understood, their reactions also appear to be initiated by H abstraction from unactivated carbon centers, in sharp contrast to the aforementioned, well-known cyclization mechanisms [18,19,20]. Since the highly strained rings are generally difficult to synthesize in a chemo- and stereo-selective manner by chemical methods, understanding the mechanisms of the nonheme iron cyclases will provide new opportunities to develop biocatalysts to access three-membered rings directly from the corresponding linear materials, via C–H bond activation. This review summarizes our understanding of the enzymatic chemistry of αKG-dependent nonheme iron enzymes involved in the natural production of epoxides, aziridines, and cyclopropanes, with a primary focus on enzyme mechanisms.
Epoxidation
Scopolamine (3) is a naturally occurring tropane alkaloid produced primarily by plants from the Solanaceae family, such as Datura, Hyoscyamus, and Atropa species (Fig. 1a) [24,25,26]. It is employed clinically for motion sickness and nausea prevention, as well as for managing conditions involving anxiety and agitation [24,25,26]. The molecular structure of scopolamine (3) is characterized by an 8-azabicyclo [3.2.1]octane core, derived from ornithine and two molecules of malonyl-CoA [27]. The αKG-dependent nonheme iron enzyme H6H catalyzes the final two steps of scopolamine biosynthesis from hyoscyamine (1) by installing the epoxide ring (Fig. 1a) [28,29,30,31,32,33,34,35]. The enzyme first introduces a hydroxyl group at C6 of hyoscyamine (1), as typically seen for this αKG-dependent nonheme iron enzyme family. The conversion of the hydroxylated intermediate 2 to scopolamine (3) was initially thought to involve 6,7-desaturated hyoscyamine, because it could be converted to scopolamine (3) in Datura ferox L [28]. The hypothetical biosynthetic intermediate, 6,7-desaturated hyoscyamine, is also accepted by H6H in vitro to produce scopolamine (3) [30]. However, a feeding experiment with [6-18O]6β-hydroxyhyoscyamine and Datura scions demonstrated that the transformation of 6β-hydroxyhyoscyamine (2) to scopolamine (3) does not involve a dehydration step [31]. This result was also consistent with the retention of the 18O atom when [6-18O]6β-hydroxyhyoscyamine was incubated with purified H6H [31]. The in vitro reaction of [7β-2H]6β-hydroxyhyoscyamine with H6H revealed the loss of the deuterium atom, indicating that the epoxidation reaction proceeds with the retention of the C7 stereoconfiguration [32].
Possible mechanisms of the epoxidation reaction catalyzed by H6H are depicted in Fig. 1b [36]. Since 6β-hydroxylation is the first step, the hydroxyl group in 2 may be located near the iron center in the H6H active site. Therefore, the iron center may be ligated by the C6-hydroxyl group in 6β-hydroxyhyoscyamine (2), similar to the proposed mechanism for the epoxidation reaction catalyzed by the αKG-independent nonheme iron enzyme HppE (Fig. 1e), although no evidence has been provided to prove this hypothesis. As a member of the αKG-dependent nonheme iron oxygenase family, an Fe(IV)-oxo species may be formed to abstract the H atom from the exo-C7 position in 6β-hydroxyhyoscyamine (2). The resulting radical intermediate 5 may react with the Fe(III)–OH to produce a 6,7-dihydroxylhyoscyamine intermediate (6 or 7), which might undergo ring closure via intramolecular substitution. Alternatively, a carbocation species 8 may be formed via single electron transfer from the radical intermediate 5 to the Fe(III)–OH complex.
To explore the potential involvement of the diol species, two possible isomers 6 and 7 were synthesized and subjected to testing with H6H (Fig. 1c) [36]. Only 7 was accepted by H6H, leading to the 7-keto product 9, suggesting that the cyclization mechanism involving a diol intermediate (6 or 7) is unlikely. Since H6H does not catalyze the hydroxylation of 6β-hydroxyhyoscyamine (2), it would be intriguing to determine what controls hydroxylation versus cyclization. It was hypothesized that the hydroxyl group adjacent to the site of H atom abstraction affects the course of the reaction. Consistent with this hypothesis, the synthetic 7β-hydroxylated compound (10) was transformed to scopolamine (3) by H6H, without producing the C6-hydroxylated product (Fig. 1c). In contrast, alcohol compounds containing a 9-azabicyclo[3.3.1]nonane core such as 12 and 13 were hydroxylated to generate the corresponding vicinal diols 14 and 15 when they were incubated with H6H (Fig. 1d). Thus, the ring size of the substrate (8-azabicyclo[3.2.1]octane versus 9-azabicyclo[3.3.1]nonane) appeared to correlate with the outcomes of the oxidation reactions by H6H. The small H–C6–C7–OH dihedral angle of 6β-hydroxyhyoscyamine (2) may be critical for the epoxidation activity. However, DFT calculations of radical intermediates derived from 6β-hydroxyhyoscyamine (2) and 13 revealed that the angle between the p orbital at C7 and the O–C6 bond would be small in both cases (less than 22 degrees). Thus, H6H may maintain the initial substrate geometry during the oxidation reactions. Another possible explanation is that a discrete radical species is only produced from a substrate with the staggered conformation (such as 12 and 13), and that H atom abstraction and C–O bond formation occur in a concerted manner during the cyclization of 6β-hydroxyhyoscyamine (2) to scopolamine (3) [36].
H6H abstracts an H atom from C6 for the first hydroxylation reaction, but activates C7 during the second epoxidation reaction. C6 may also be the preferred site of H atom abstraction during the epoxidation. Kinetic analyses of the H6H-catalyzed hydroxylation and epoxidation were conducted using deuterated substrates [37]. Interestingly, during hydroxylation, the regioselectivity of H abstraction from C6 versus C7 was approximately 85:1, but it was reversed to be approximately 1:16, indicating a more than thousand-fold difference. The significant change of the regioselectivity of H abstraction could be explain by the proposed direct coordination of the hydroxyl group in 2 during the cyclization reaction.
The regioselectivity of H abstraction during the hydroxylation step was also investigated, based on crystal structures and QM/MM calculations [38]. The X-ray crystal structure of H6H from Datura metel in complex with hyoscyamine and N-oxalylglycine (NOG) revealed that C6 in hyoscyamine (1) is located 4.8 Å from the metal center, while C7 is positioned at a distance of 4.5 Å (Fig. 2). However, based on the QM/MM calculations with a model including the Fe(IV)-oxo species, H atom abstraction from C6 is significantly favored. Detailed geometrical analysis indicated that the Fe–O–H angle in the transition state was 141°, presumably enabling the energetically preferred σ-channel H abstraction. In contrast, the angle was 110° in the transition state structure for the H abstraction from C7, implying that this step follows the energetically less favored π-channel pathway. In silico amino acid substitutions in H6H, followed by energy calculations, were also conducted to identity the active site residues that might be critical in determining the regioselectivity of H abstraction. The results suggested that Tyr326 stabilizes the transition state of C6–H abstraction and destabilizes that of C7–H abstraction by interacting with the phenyl ring of the tropate side chain of hyoscyamine (1). The Lys129 residue is also important for the regioselectivity because it fixes the position of hyoscyamine (1) in the active site through electrostatic interactions with Glu116 that forms a hydrogen bond with the ammonium moiety in the substrate (Fig. 2) [38].
Crystal structure of H6H from Datura metel in complex with hyoscyamine and NOG. Hyo = hyoscyamine (1) (PDB 6TTN) [38]
FfnD is a recently characterized cyclase that produces 17 from 16 during the biosynthesis of fosfonochlorin (Fig. 1e) [39]. In vitro enzyme assays with deuterated substrates revealed that the FfnD reaction retains the stereoconfiguration of the substrate. It is worth noting that this cyclization reaction bears similarities to the epoxidation reaction (from 18 to 19) catalyzed by HppE in fosfomycin biosynthesis [40, 41]. However, there are several fundamental distinctions between these two enzymes. FfnD requires αKG and O2 as co-substrates, whereas HppE utilizes either the combination of NADH/FMN/O2 or H2O2. In addition, HppE epoxidation results in an inversion of the configuration. Consequently, the mechanism employed by FfnD may differ from that of HppE [42].
Aziridination
Tryptoquialanine (23) is a peptide-based natural product produced by Penicillium aethiopicum (Fig. 3a) [43, 44]. Biosynthetic studies revealed that tryptoquialanine (23) is assembled by the non-ribosomal peptide synthase (NRPS) TqaA, using amino acid components such as l-alanine, anthranilic acid, l-tryptophan, and 2-aminoisobutyric aid (22) [45,46,47]. 2-Aminoisobutyric aid (22) is a nonproteinogenic amino acid that is also present in other various biologically active fungal natural compounds, including alamethicin [48,49,50], zervamicin [51, 52], and emericellipsin [53]. Gene deletion in P. aethiopicum and in vitro enzyme characterization demonstrated that three sequential enzymatic reactions by TqaL, TqaF, and TqaM convert l-valine to 2-aminoisobutyric acid (22) [47, 54]. TqaL is an αKG-dependent nonheme iron oxygenase that catalyzes the aziridination of l-valine (20) to pleurocybellaziridine (21) (Fig. 3a) [54]. An in vitro analysis of stereoselectively deuterated l-valine (20) suggested that the aziridination reaction catalyzed by TqaL is not stereospecific, and instead involves both C3 retention and inversion of stereochemical courses [55]. A possible reaction mechanism is shown in Fig. 3e. The aziridination may be initiated by H atom abstraction from C3 of l-valine by an Fe(IV)-oxo species (20). The radical species 32 may be cyclized via a radical-based mechanism. Alternatively, 32 could be oxidized by the Fe(III)–OH species to generate the carbocation intermediate 33 that undergoes C–N bond-forming ring closure via a polar mechanism. The C2–C3 bond may undergo rapid rotation at the radical intermediate (32) stage [55].
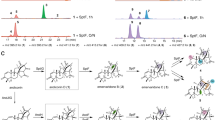
TqaL from Neurospora crassa (TqaL-nc) can also accept l-isoleucine (24) and l-allo-isoleucine (25), in addition to the native substrate l-valine (20) (Fig. 3b) [55]. These two non-native substrates were converted to the same diastereomeric pairs of the aziridine product with a constant ratio (26: 27 = 3: 2), implying the involvement of an enzyme-controlled stereoconvergent process. The molecular basis of substrate binding was studied, based on a crystal structure analysis and structure prediction by Alphafold2 (Fig. 4) [55]. Accordingly, it was proposed that the amino group of the substrate interacts with Glu251, while the carboxylate moiety forms an H bonding network with three arginine residues, Arg144, Arg178, and Arg185. Notably, Ile343 and Phe345 are in the proximity of the side chain of the substrate, raising the hypothesis that these two residues may control the stereoselectivity of the aziridination of l-isoleucine (24) and l-allo-isoleucine (25). Consistent with this hypothesis, individual substitutions of Ile343 and Phe345 to alanine improved the stereoselectivity of the aziridination (26: 27 = 3: 1) (Fig. 3c). These studies demonstrated that the stereoselectivity could be rationally controlled by structure-based protein engineering, by dictating the conformation of the unstable reaction intermediate.
Predicted structure of TqaL-nc docked with l-isoleucine (24) [55]
Several peptide natural products, such as atroviridins [56] and efrapeptins [57, 58] from fungal origins, contain isovaline moieties as an amino acid building block, implying that it is derived from l-isoleucine (24) via pathways analogous to that of 2-aminoisobutyric acid (22) in tryptoquialanine (23). A recent study indeed identified a homologue of TqaL (TqaL-ha) from the atroviridin-producing species Hypocrea atroviridis, which accepts l-isoleucine (24) to selectively produce 26 (Fig. 3d) [59]. Analysis of the TqaL-ha reaction by stopped-flow absorption spectroscopy using deuterated l-isoleucine (24) indicated that the Fe(IV)-oxo species is generated to abstract an H atom from the substrate. The results also suggested that a hydroxylated species is unlikely to be a reaction intermediate, because two isomers of β-hydroxylated isoleucines were not converted to the aziridine product 26 when incubated with TqaL-ha. Interestingly, when l-homoalanine (28) was employed as a possible substrate of TqaL-ha, the hydroxylated product 29 was produced without the formation of the aziridine product (Fig. 3d) [59]. This observed change of the reaction flux from aziridination to hydroxylation may be correlated with the stability of the cation intermediate. In the native reaction of 23 for TqaL-nc or 24 for TqaL-ha, following H abstraction by the Fe(IV)-oxo species, the radical intermediate may be oxidized to the tertiary carbocation via single electron transfer. In contrast, the corresponding radical intermediate derived from l-homoalanine (28) may undergo hydroxyl rebound with the Fe(III)-OH intermediate, because the hypothetical secondary carbocation that could have resulted from l-homoalanine (28) may be difficult to form, due to its decreased stability. The aziridination mechanism involving the carbocation intermediate was further supported by an experiment with 4-trifluorovaline (30), where the hydroxylated product 31 was predominantly generated with no formation of the corresponding aziridine product (Fig. 3d) [59].
Cyclopropanation
Cycloclavine (40) is a cyclopropane ring-containing ergot alkaloid from the filamentous fungus, Aspergillus japonicus (Fig. 5) [60,61,62,63]. The biosynthetic gene cluster responsible for cycloclavine biosynthesis encodes seven enzymes [64]. Expression of the biosynthetic genes in Saccharomyces cerevisiae revealed that the cyclopropyl amine moiety is constructed during the last two steps of cycloclavine biosynthesis [65]. Namely, the αKG-dependent nonheme iron oxygenase EasH catalyzes the cyclopropanation of 34/35 to generate 39, which is then reductively transformed to cycloclavine (40) by the NADPH-dependent reductase EasG. A related homologue of EasH from Claviceps purpurea catalyzes the C–O bond-forming cyclization of dihydroergotaman to dihydroergotamine [66]. Another homologue from an Epichlo sp. was also analyzed by heterologous gene expression, but its catalytic function remains unclear [67]. Two possible mechanisms for the EasH-catalyzed cyclopropanation are shown in Fig. 5 [65]. The substrate of EasH may exist as a mixture of the imine (34) and enamine (35) forms. An Fe(IV)-oxo species may abstract an H atom from C10 in 35, followed by hydroxyl rebound to generate 36. Then, C–C bond formation may occur with elimination of the newly installed hydroxyl group to produce 39. This intramolecular substitution process is hypothesized to require acid–base chemistry. However, the X-ray crystal structure of EasH from A. japonicus suggested that the protein active site is constructed primarily by hydrophobic residues [68]. Therefore, a non-polar mechanism that is not assisted by polar active site residues has also been proposed (Fig. 5). The radical in 37 may intramolecularly add across the double bond of the enamine moiety to generate 38, which is further oxidized into the imine product 39. The mechanism involving the radical-based C–C bond formation step has also been suggested by QM/MM calculations [69]. These studies characterized EasH as the first example of cyclopropanation catalysis via αKG-dependent nonheme iron chemistry, although further examination will be necessary to test the hypothetical mechanisms.
The second examples of αKG-dependent nonheme iron cyclopropanases were discovered in the biosynthetic pathways of belactosin A (41) and hormaomycin (43) (Fig. 6a). Belactosin A (41) and hormaomycin (43) are distinct types of peptide natural products from Streptomycetes [70,71,72,73,74]. Belactosin A (41) has a β-lactone warhead conjugated with a (1′R,2′S)-3-(2-aminocyclopropyl)alanine (42, Acpa) residue. The structure of hormaomycin (43) features several unusual (amino)acid units, including 5-chloro-1-hydroxypyrrole-2-carboxylic acid, 3-propenylproline, and (1′R,2′R)-3-(2-nitrocyclopropyl)alanine (44, Ncpa) [75, 76]. (1′R,2′S)-Acpa (42) and (1′R,2′R)-Ncpa (44) are similar to each other, but possess different nitrogen substituent oxidation states and cyclopropane ring stereoconfigurations. The resemblance between Acpa (42) and Ncpa (44) implied that they share a common biosynthetic pathway, in which l-lysine serves as a precursor [77, 78]. The biosynthetic gene clusters of belactosin A (41) and hormaomycin (43) were independently discovered [79, 80]. Gene analysis suggested that the peptide structure of hormaomycin (43) is assembled by NRPSs, while the peptide bonds in belactosin A (41) are constructed by ATP-grasp ligases. Notably, the two biosynthetic gene clusters contain three pairs of the homologous genes belK-hrmI, belL-hrmJ, and belM-hrmT. Gene deletions in the belactosin A-producing strain Streptomyces sp. UCK14, combined with in vitro enzyme assays, indicated that the heme oxygenase-like diiron enzymes BelK and HrmI have the same catalytic function to oxidize l-lysine and generate l-6-nitronorleucine (45) [81,82,83]. Subsequently, the nonheme iron enzymes BelL and HrmJ similarly catalyze the dehydrogenative cyclization of the nitroalkane moiety of l-6-nitronorleucine (45) to generate Ncpa (44) [81, 82]. Consistent with the stereoconfigurations found in belactosin A (41) and hormaomycin (43), BelL produces (1′S,2′S)-Ncpa, while HrmJ produces (1′R,2′R)-Ncpa despite its moderate sequence identity (49% identity). The homologous genes belM and hrmT may encode diaminopimelate epimerases that could supply l-lysine as a precursor for Acpa (42) and Ncpa (44). It should also be noted that, in belactosin A (41) biosynthesis, the nitro group in (1′S,2′S)-Ncpa ((1′S,2′S)-44) is predicted to be reduced by the molybdenum-dependent reductase BelN [84].
A recent bioinformatic analysis of the belL gene revealed that many bacterial strains of Actinomycetota and Proteobacteria have belL gene homologues [85]. In vitro analysis of the l-6-nitronorleucine (45) reaction with purified BelL homologues indicated that they also produce the cis isomers (Fig. 6a). For example, the BelL homologue from Streptomyces rimosus subsp. Paromomycinus NBRC 15454 (SrBelL) generates (1′S,2′R)-Ncpa ((1′S,2′R)-44), while that from Streptomyces cavourensis NBRC 13026 (ScBelL) generates (1′R,2′S)-Ncpa ((1′R,2′S)-44). Therefore, this group of nonheme iron cyclopropanases produces one of the four possible diastereomers of Ncpa (44) with a remarkable stereocontrol mechanism.
Possible mechanisms for the cyclopropanation of l-6-nitronorleucine (45) are shown in Fig. 6b [81, 82, 85]. The α-carbon (C6) of the nitro group in l-6-nitronorleucine (45) may be deprotonated by a base in the active site to generate the nitronate intermediate 46. Then, an Fe(IV)–oxo species abstracts an H atom from the C4 position to form the radical intermediate 47. Several different pathways can be envisioned. The radical 47 may undergo radical addition across the double bond of the nitronate moiety to form the cyclopropane ring with the nitro anion radical 48. Further oxidation of this species could afford the product Ncpa (44). An alternative mechanism involving the cation intermediate 49 should also be considered. It is also possible that the radical in 47 receives the hydroxyl group from the Fe(III) to form the C4–OH intermediate 50, which might be cyclized into 44 via an intramolecular substitution reaction. It was suggested that the C6-deprotonation of l-6-nitronorleucine (45) could occur non-enzymatically at pH 7.5 [85]. Nevertheless, the addition of BelL to l-6-nitronorleucine (45) accelerated the rate of C6-deprotonation. The deprotonation is critical for the cyclization activity, because the fluorinated substrate analogue 53 that could not be deprotonated was not accepted by HrmJ, while the substrate analogue 51 containing a carboxylate moiety as a mimic of the nitro group in 45 was only hydroxylated, without the formation of the cyclized product (Fig. 6c). In addition, the enzymatic cyclization of l-6-nitronorleucine (45) was promoted under basic conditions (pH 9–11), consistent with the C6-deprotonation as a critical step.
Similar to other αKG-dependent nonheme iron enzymes, BelL and its homologues utilize an Fe(IV)-oxo species to abstract an H atom from the substrate or 46, as confirmed by a transient kinetic analysis using stopped-flow absorption spectroscopy and Mössbauer spectroscopy [85]. The site of H abstraction from C4 was also determined to be the pro-S–H, based on the observed loss of the deuterium atom when (4S)-[4-2H]-45 was employed as the substrate of BelL (Fig. 6b) [81, 85]. Notably, all other tested homologues of BelL similarly abstract pro-S–H, regardless of the diastereoselectivity of the reactions they catalyze. The radical 47 is likely cyclized directly into 44 via either a cation- or radical-based mechanism, since the synthetic 4-hydroxyl compound 54 was not converted to Ncpa (44), effectively excluding the possibility of the involvement of 50 (and 54) as an on-pathway intermediate. DFT calculations suggested that the radical-based cyclization of 47 to 48 is energetically feasible, with an activation energy (ΔG‡) of 7.4 kcal mol–1. It is possible that the delocalization of the spin density over the three heteroatoms may contribute to the stabilization of 48, thereby overcoming the inherent ring strain of the cyclopropane ring.
BelL and its homologues exhibit remarkable stereochemical control during the dehydrogenative cyclization of l-6-nitronorleucine (45) (Fig. 6a). The contributions of the active site residues toward the stereochemical outcomes are partially understood, based on structural analyses of this class of enzymes [85]. The X-ray crystal structure of ScBelL suggested that it resembles l-isoleucine dioxygenase (IDO) from Bacillus thuringiensis, consisting of a jelly roll fold with two anti-parallel β-sheets. Sequence alignments and comparisons of the homology models among the tested cyclopropanases revealed that many of the active site residues are conserved, while a few amino acid residues differ. For example, BelL has His at position 136, while the corresponding residue in HrmJ is substituted with Arg (Fig. 7). These residues are critical in controlling the product stereochemistry, because the BelL-H136R variant produced (1′R,2′R)-Ncpa ((1′R,2′R)-44) without forming the original product (1′S,2′S)-Ncpa ((1′S,2′S)-44). As expected, the HrmJ-R136H variant generated (1′S,2′S)-Ncpa ((1′S,2′S)-44), although the variant retained some activity to produce (1′R,2′R)-Ncpa ((1′R,2′R)-44). When the amino acid substitution was rationally designed based on sequence comparisons with homologues that produce cis-Ncpa (44), BelL was engineered to produce cis-Ncpa (44). However, the rational substitution of the active site residues did not necessarily lead to the desired changes of the product stereochemistry [85]. Thus, further investigations will be needed to clarify how these cyclopropanases control selectivity with only subtle differences in their primary sequences.
Comparison of the predicted structures of a BelL and b HrmJ docked with the nitronate form of the substrate 46 [85]
Another αKG-dependent nonheme iron cyclopropanase was recently characterized in the biosynthetic pathway of breviones [86] (Fig. 8). Breviones are meroterpenoid natural products from fungal strains such as Penicillium brevicompactum and Penicillium bialowiezense CBS 227.28 [87]. Brevione B (55) is biosynthesized via combined polyketide and terpenoid biosynthetic machineries and is proposed to be a common intermediate for setosusin and brevione E, a highly functionalized derivative of breviones. The biosynthetic gene clusters of setosusin and breviones encode two related nonheme iron enzymes, SetK and BrvJ, respectively [86, 88]. Heterologous expression revealed that SetK catalyzes brevione B (55) hydroxylation at the C1 position, while BrvJ performs oxidative ring expansion of the same intermediate to generate brevione C (61) via brevione A (57) [86]. BrvJ further converted brevione C (61) into the cyclopropane-containing brevione W (64) under the tested in vitro conditions. Structure-based mutagenesis of the active site residues (F159L/A227L/L238M) converted BrvJ into a C1-hydroxylase like SetK that generates brevione S (56).
The ring expansion of brevione A (57) may be initiated by H atom abstraction from C18 (Fig. 8) [86]. The resulting radical 58 rearranges to 60 through the O-centered radical with a cyclopropane ring (59), followed by H atom abstraction from C18 to produce brevione C (61). In the next catalytic cycle, an H atom from C18 in brevione C (61) may be abstracted to generate 52, which undergoes radical addition onto the enone moiety to form the radical 63 with a cyclopropane ring. Finally, hydroxyl rebound with the Fe(III)-OH species may occur to produce 64. Alternatively, the introduction of the hydroxy group at C2 may be effected via the carbocation intermediate 65.
Conclusions
Recent advancements in genome sequencing and bioinformatic technologies have enabled the discovery of nonheme iron-dependent cyclases producing epoxides, aziridines, and cyclopropanes, with H6H from the 1980s serving as an exception. Accumulated mechanistic knowledge suggests that, following C–H bond heterolytic cleavage, the key ring-closure step is likely achieved through either radical addition across an unsaturated bond or cation-mediated bond formation. However, unresolved enzymological questions remain to be answered. For example, it is not fully understood how these enzymes catalyze selective cyclization while avoiding hydroxylation, which is the typical outcome of αKG-dependent nonheme iron enzyme catalysis. Further mechanistic investigations will create new opportunities to develop practical biocatalysts for challenging cyclization reactions beyond the capabilities of current organic synthesis technology.
References
Thibodeaux CJ, Chang W-c, Liu H-w (2012) Enzymatic chemistry of cyclopropane, epoxide, and aziridine biosynthesis. Chem Rev 112:1681–1709
Falagas ME, Vouloumanou EK, Samonis G, Vardakas KZ (2016) Fosfomycin. Clin Microbiol Rev 29:321–347
Michalopoulos AS, Livaditis IG, Gougoutas V (2011) The revival of fosfomycin. Int J Infect Dis 15:e732–e739
Silver LL (2017) Fosfomycin: mechanism and resistance. Cold Spring Harb Perspec Med 7:a025262
Crooke ST, Bradner WT (1976) Mitomycin C: a review. Cancer Treat Rev 3:121–139
Tomasz M (1995) Mitomycin C: small, fast and deadly (but very selective). Chem Biol 2:575–579
Suresh Kumar G, Lipman R, Cummings J, Tomasz M (1997) Mitomycin C−DNA adducts generated by DT-diaphorase. Revised mechanism of the enzymatic reductive activation of mitomycin C. Biochemistry 36:14128–14136
Talele TT (2016) The “cyclopropyl fragment” is a versatile player that frequently appears in preclinical/clinical drug molecules. J Med Chem 59:8712–8756
Ma S, Mandalapu D, Wang S, Zhang Q (2022) Biosynthesis of cyclopropane in natural products. Nat Prod Rep 39:926–945
Wessjohann LA, Brandt W, Thiemann T (2003) Biosynthesis and metabolism of cyclopropane rings in natural compounds. Chem Rev 103:1625–1648
Kelly WL, Boyne MT, Yeh E, Vosburg DA, Galonić DP, Kelleher NL, Walsh CT (2007) Characterization of the aminocarboxycyclopropane-forming enzyme CmaC. Biochemistry 46:359–368
Kurosawa S, Hasebe F, Okamura H, Yoshida A, Matsuda K, Sone Y, Tomita T, Shinada T, Takikawa H, Kuzuyama T, Kosono S, Nishiyama M (2022) Molecular basis for enzymatic aziridine formation via sulfate elimination. J Am Chem Soc 144:16164–16170
Zha L, Jiang Y, Henke MT, Wilson MR, Wang JX, Kelleher NL, Balskus EP (2017) Colibactin assembly line enzymes use S-adenosylmethionine to build a cyclopropane ring. Nat Chem Biol 13:1063–1065
Guengerich FP (2003) Cytochrome P450 oxidations in the generation of reactive electrophiles: epoxidation and related reactions. Arch Biochem Biophys 409:59–71
Savage TJ, Croteau R (1993) Biosynthesis of monoterpenes: regio-and stereochemistry of (+)-3-carene biosynthesis. Arch Biochem Biophys 305:581–587
Hoelscher DJ, Williams DC, Wildung MR, Croteau R (2003) A cDNA clone for 3-carene synthase from Salvia stenophylla. Phytochemistry 62:1081–1086
Fäldt J, Martin D, Miller B, Rawat S, Bohlmann J (2003) Traumatic resin defense in Norway spruce (Picea abies): Methyl jasmonate-induced terpene synthase gene expression, and cDNA cloning and functional characterization of (+)-3-carene synthase. Plant Mol Biol 51:119–133
Ushimaru R, Abe I (2022) Unusual dioxygen-dependent reactions catalyzed by nonheme iron enzymes in natural product biosynthesis. ACS Catal 13:1045–1076
Martinez S, Hausinger RP (2015) Catalytic mechanisms of Fe(II)- and 2-oxoglutarate-dependent oxygenases. J Biol Chem 290:20702–20711
Krebs C, Galonić Fujimori D, Walsh CT, Bollinger JM Jr (2007) Non-heme Fe(IV)–oxo intermediates. Acc Chem Res 40:484–492
Wu L-F, Meng S, Tang G-L (2016) Ferrous iron and α-ketoglutarate-dependent dioxygenases in the biosynthesis of microbial natural products. Biochim Biophys Acta Proteins Proteom 1864:453–470
Nakamura H, Matsuda Y, Abe I (2018) Unique chemistry of non-heme iron enzymes in fungal biosynthetic pathways. Nat Prod Rep 35:633–645
Gao S-S, Naowarojna N, Cheng R, Liu X, Liu P (2018) Recent examples of α-ketoglutarate-dependent mononuclear non-haem iron enzymes in natural product biosyntheses. Nat Prod Rep 35:792–837
Klinkenberg I, Blokland A (2010) The validity of scopolamine as a pharmacological model for cognitive impairment: a review of animal behavioral studies. Neurosci Biobehav Rev 34:1307–1350
Renner UD, Oertel R, Kirch W (2005) Pharmacokinetics and pharmacodynamics in clinical use of scopolamine. Ther Drug Monit 27:655–665
Kohnen-Johannsen KL, Kayser O (2019) Tropane alkaloids: chemistry, pharmacology, biosynthesis and production. Molecules 24:796
Huang J-P, Wang Y-J, Tian T, Wang L, Yan Y, Huang S-X (2021) Tropane alkaloid biosynthesis: a centennial review. Nat Prod Rep 38:1634–1658
Fodor G, Romeike A, Janzso G, Koczor I (1959) Epoxidation experiments in vivo with dehydrohyoscyamine and related compounds. Tetrahedron Lett 1:19–23
Hashimoto T, Yamada Y (1986) Hyoscyamine 6β-hydroxylase, a 2-oxoglutarate-dependent dioxygenase, in alkaloid-producing root cultures. Plant Physiol 81:619–625
Hashimoto T, Yamada Y (1987) Purification and characterization of hyoscyamine 6β-hydroxylase from root cultures of Hyoscyamus niger L. Hydroxylase and epoxidase activities in the enzyme preparation. Euro J Biochem 164:277–285
Hashimoto T, Kohno J, Yamada Y (1987) Epoxidation in vivo of hyoscyamine to scopolamine does not involve a dehydration step. Plant physiol 84:144–147
Hashimoto T, Kohno J, Yamada Y (1989) 6β-Hydroxyhyoscyamine epoxidase from cultured roots of Hyoscyamus niger. Phytochemistry 28:1077–1082
Matsuda J, Okabe S, Hashimoto T, Yamada Y (1991) Molecular cloning of hyoscyamine 6 beta-hydroxylase, a 2-oxoglutarate-dependent dioxygenase, from cultured roots of Hyoscyamus niger. J Biol Chem 266:9460–9464
Hashimoto T, Matsuda J, Yamada Y (1993) Two-step epoxidation of hyoscyamine to scopolamine is catalyzed by bifunctional hyoscyamine 6β-hydroxylase. FEBS lett 329:35–39
Li J, van Belkum MJ, Vederas JC (2012) Functional characterization of recombinant hyoscyamine 6β-hydroxylase from Atropa belladonna. Bioorg Med Chem 20:4356–4363
Ushimaru R, Ruszczycky MW, Chang W-c, Yan F, Liu Y-n, Liu H-w (2018) Substrate conformation correlates with the outcome of hyoscyamine 6β-hydroxylase catalyzed oxidation reactions. J Am Chem Soc 140:7433–7436
Ushimaru R, Ruszczycky MW, Liu H-w (2018) Changes in regioselectivity of H atom abstraction during the hydroxylation and cyclization reactions catalyzed by hyoscyamine 6β-hydroxylase. J Am Chem Soc 141:1062–1066
Kluza A, Wojdyla Z, Mrugala B, Kurpiewska K, Porebski PJ, Niedzialkowska E, Minor W, Weiss MS, Borowski T (2020) Regioselectivity of hyoscyamine 6β-hydroxylase-catalysed hydroxylation as revealed by high-resolution structural information and QM/MM calculations. Dalton Trans 49:4454–4469
Gama SR, Stankovic T, Hupp K, Al Hejami A, McClean M, Evans A, Beauchemin D, Hammerschmidt F, Pallitsch K, Zechel DL (2022) Biosynthesis of the fungal organophosphonate fosfonochlorin involves an iron (II) and 2-(oxo) glutarate dependent oxacyclase. ChemBioChem 23:e202100352
Liu P, Murakami K, Seki T, He X, Yeung S-M, Kuzuyama T, Seto H, Liu H-w (2001) Protein purification and function assignment of the epoxidase catalyzing the formation of fosfomycin. J Am Chem Soc 123:4619–4620
Wang C, Chang W-c, Guo Y, Huang H, Peck SC, Pandelia ME, Lin G-m, Liu H-w, Krebs C, Bollinger JM Jr (2013) Evidence that the fosfomycin-producing epoxidase, HppE, is a non–heme-iron peroxidase. Science 342:991–995
Zhou S, Pan J, Davis KM, Schaperdoth I, Wang B, Boal AK, Krebs C, Bollinger JM Jr (2019) Steric enforcement of cis-epoxide formation in the radical C-O-coupling reaction by which (S)-2-hydroxypropylphosphonate epoxidase (HppE) produces Fosfomycin. J Am Chem Soc 141:20397–20406
Clardy J, Springer JP, Buechi G, Matsuo K, Wightman R (1975) Tryptoquivaline and tryptoquivalone, two tremorgenic metabolites of Aspergillus clavatus. J Am Chem Soc 97:663–665
Ariza MR, Larsen TO, Petersen BO, Duus JØ, Barrero AF (2002) Penicillium digitatum metabolites on synthetic media and citrus fruits. J Agric Food Chem 50:6361–6365
Gao X, Haynes SW, Ames BD, Wang P, Vien LP, Walsh CT, Tang Y (2012) Cyclization of fungal nonribosomal peptides by a terminal condensation-like domain. Nat Chem Biol 8:823–830
Haynes SW, Ames BD, Gao X, Tang Y, Walsh CT (2011) Unraveling terminal C-domain-mediated condensation in fungal biosynthesis of imidazoindolone metabolites. Biochemistry 50:5668–5679
Gao X, Chooi Y-H, Ames BD, Wang P, Walsh CT, Tang Y (2011) Fungal indole alkaloid biosynthesis: genetic and biochemical investigation of the tryptoquialanine pathway in Penicillium aethiopicum. J Am Chem Soc 133:2729–2741
Leitgeb B, Szekeres A, Manczinger L, Vagvolgyi C, Kredics L (2007) The history of alamethicin: a review of the most extensively studied peptaibol. Chem Biodivers 4:1027–1051
Hall JE, Vodyanoy I, Balasubramanian T, Marshall GR (1984) Alamethicin. A rich model for channel behavior. Biophys J 45:233–247
Fox RO Jr, Richards FM (1982) A voltage-gated ion channel model inferred from the crystal structure of alamethicin at 1.5-Å resolution. Nature 300:325–330
Rinehart KL Jr, Gaudioso LA, Moore ML, Pandey RC, Cook JC Jr, Barber M, Sedgwick RD, Bordoli RS, Tyler AN, Green BN (1981) Structures of eleven zervamicin and two emerimicin peptide antibiotics studied by fast atom bombardment mass spectrometry. J Am Chem Soc 103:6517–6520
Karle IL, Flippen-Anderson J, Sukumar M, Balaram P (1987) Conformation of a 16-residue zervamicin IIA analog peptide containing three different structural features: 3 (10)-helix, alpha-helix, and beta-bend ribbon. Proc Natl Acad Sci USA 84:5087–5091
Rogozhin EA, Sadykova VS, Baranova AA, Vasilchenko AS, Lushpa VA, Mineev KS, Georgieva ML, Kul’ko AB, Krasheninnikov ME, Lyundup AV (2018) A novel lipopeptaibol emericellipsin A with antimicrobial and antitumor activity produced by the extremophilic fungus Emericellopsis alkalina. Molecules 23:2785
Bunno R, Awakawa T, Mori T, Abe I (2021) Aziridine formation by a FeII/α-ketoglutarate dependent oxygenase and 2-aminoisobutyrate biosynthesis in fungi. Angew Chem Int Ed 60:15847–15831
Tao H, Ushimaru R, Awakawa T, Mori T, Uchiyama M, Abe I (2022) Stereoselectivity and substrate specificity of the Fe (II)/α-ketoglutarate-dependent oxygenase TqaL. J Am Chem Soc 144:21512–21520
Oh S-U, Lee S-J, Kim J-H, Yoo I-D (2000) Structural elucidation of new antibiotic peptides, atroviridins A, B and C from trichoderma atroviride. Tetrahedron Lett 41:61–64
Abrahams JP, Buchanan SK, Van Raaij MJ, Fearnley IM, Leslie A, Walker JE (1996) The structure of bovine F1-ATPase complexed with the peptide antibiotic efrapeptin. Proc Natl Acad Sci USA 93:9420–9424
Cross RL, Kohlbrenner W (1978) The mode of inhibition of oxidative phosphorylation by efrapeptin (A23871). Evidence for an alternating site mechanism for ATP synthesis. J Biol Chem 253:4865–4873
Cha L, Paris JC, Zanella B, Spletzer M, Yao A, Guo Y, Chang W-c (2023) Mechanistic studies of aziridine formation catalyzed by mononuclear non-heme iron enzymes. J Am Chem Soc 145:6240–6246
Davis KA, Jones AM, Panaccione DG (2023) Two satellite gene clusters enhance ergot alkaloid biosynthesis capacity of Aspergillus leporis. Appl Environ Microbiol 89:e00793-e723
Pařenicová L, Skouboe P, Frisvad J, Samson RA, Rossen L, Mt H-S, Visser J (2001) Combined molecular and biochemical approach identifies Aspergillus japonicus and Aspergillus aculeatus as two species. Appl Environ Microbiol 67:521–527
Stauffacher D, Niklaus P, Tscherter H, Weber H, Hofmann A (1969) Cycloclavin, ein neues alkaloid aus Ipomoea hildebrandtii vatke—71: Mutterkornalkaloide. Tetrahedron 25:5879–5887
Incze M, Dörnyei G, Moldvai I, Temesvári-Major E, Egyed O, Szántay C (2008) New routes to clavine-type ergot alkaloids. Part 2: Synthesis of the last, so far not yet synthesized member of the clavine alkaloid family, (±)-cycloclavine. Tetrahedron 64:2924–2929
Wallwey C, Li S-M (2011) Ergot alkaloids: structure diversity, biosynthetic gene clusters and functional proof of biosynthetic genes. Nat Prod Rep 28:496–510
Jakubczyk D, Caputi L, Hatsch A, Nielsen CA, Diefenbacher M, Klein J, Molt A, Schröder H, Cheng JZ, Naesby M (2015) Discovery and reconstitution of the cycloclavine biosynthetic pathway—enzymatic formation of a cyclopropyl group. Angew Chem Int Ed 54:5117–5121
Havemann J, Vogel D, Loll B, Keller U (2014) Cyclolization of d-lysergic acid alkaloid peptides. Chem Biol 21:146–155
Robinson SL, Panaccione DG (2014) Heterologous expression of lysergic acid and novel ergot alkaloids in Aspergillus fumigatus. Appl Environ Microbiol 80:6465–6472
Jakubczyk D, Caputi L, Stevenson CE, Lawson DM, O’Connor SE (2016) Structural characterization of EasH (Aspergillus japonicus)—an oxidase involved in cycloclavine biosynthesis. Chem Commun 52:14306–14309
Yan L, Liu Y (2019) Insights into the mechanism and enantioselectivity in the biosynthesis of ergot alkaloid cycloclavine catalyzed by Aj_EasH from Aspergillus japonicus. Inorg Chem 58:13771–13781
Asai A, Hasegawa A, Ochiai K, Yamashita Y, Mizukami T (2000) Belactosin A, a novel antituor antibiotic acting on cyclin/CDK mediated cell cycle regulation, produced by Streptomyces sp. J Antibiot 53:81–83
Groll M, Larionov OV, Huber R, de Meijere A (2006) Inhibitor-binding mode of homobelactosin C to proteasomes: new insights into class I MHC ligand generation. Proc Natl Acad Sci USA 103:4576–4579
Kaysser L (2019) Built to bind: biosynthetic strategies for the formation of small-molecule protease inhibitors. Nat Prod Rep 36:1654–1686
Omura S, Mamada H, Wang N-J, Imamura N, Oiwa R, Iwai Y, Muto N (1984) Takaokamycin, a new peptide antibiotic produced by Streptomyces sp. J Antibiot 37:700–705
Andres N, Wolf H, Zähner H, Rössner E, Zeeck A, König WA, Sinnwell V (1989) Stoffwechselprodukte von Mikroorganismen. 253. Mitteilung. Hormaomycin, ein neues Peptid-lacton mit morphogener Aktivität auf Streptomyceten. Helv Chim Acta 72:426–437
Rössner E, Zeeck A, König WA (1990) Elucidation of the structure of hormaomycin. Angew Chem Int Ed 29:64–65
Zlatopolskiy BD, Loscha K, Alvermann P, Kozhushkov SI, Nikolaev SV, Zeeck A, de Meijere A (2004) Final elucidation of the absolute configuration of the signal metabolite hormaomycin. Chem Eur J 10:4708–4717
Brandl M, Kozhushkov SI, Zlatopolskiy BD, Alvermann P, Geers B, Zeeck A, de Meijere A (2005) The biosynthesis of 3‐(trans‐2‐nitrocyclopropyl) alanine, a constituent of the signal metabolite hormaomycin. Eur J Org Chem 123–135
Kozhushkov SI, Zlatopolskiy BD, Brandl M, Alvermann P, Radzom M, Geers B, de Meijere A, Zeeck A (2005) Hormaomycin analogues by precursor‐directed biosynthesis–synthesis of and feeding experiments with amino acids related to the unique 3-(trans‐2‐nitrocyclopropyl) alanine constituent. Eur J Org Chem 854–863
Wolf F, Bauer JS, Bendel TM, Kulik A, Kalinowski J, Gross H, Kaysser L (2017) Biosynthesis of the β-lactone proteasome inhibitors belactosin and cystargolide. Angew Chem Int Ed 56:6665–6668
Höfer I, Crüsemann M, Radzom M, Geers B, Flachshaar D, Cai X, Zeeck A, Piel J (2011) Insights into the biosynthesis of hormaomycin, an exceptionally complex bacterial signaling metabolite. Chem Biol 18:381–391
Shimo S, Ushimaru R, Engelbrecht A, Harada M, Miyamoto K, Kulik A, Uchiyama M, Kaysser L, Abe I (2021) Stereodivergent nitrocyclopropane formation during biosynthesis of belactosins and hormaomycins. J Am Chem Soc 143:18413–18418
Li X, Shimaya R, Dairi T, Wc C, Ogasawara Y (2022) Identification of cyclopropane formation in the biosyntheses of hormaomycins and belactosins: sequential nitration and cyclopropanation by metalloenzymes. Angew Chem Int Ed 61:e202113189
Pang L, Niu W, Duan Y, Huo L, Li A, Wu J, Zhang Y, Bian X, Zhong G (2022) In vitro characterization of a nitro-forming oxygenase involved in 3-(trans-2′-aminocyclopropyl)alanine biosynthesis. Eng Microbiol 2:100007
Engelbrecht A, Wolf F, Esch A, Kulik A, Kozhushkov SI, de Meijere A, Hughes CC, Kaysser L (2022) Discovery of a cryptic nitro intermediate in the biosynthesis of the 3-(trans-2′-aminocyclopropyl) alanine moiety of belactosin A. Org Lett 24:736–740
Ushimaru R, Cha L, Shimo S, Li X, Paris J, Mori T, Miyamoto K, Coffer L, Uchiyama M, Guo Y et al (2023) Mechanistic analysis of stereodivergent nitroalkane cyclopropanation catalyzed by nonheme iron enzymes. J Am Chem Soc. https://doi.org/10.1021/jacs.3c08413
Yan D, Matsuda Y (2022) Biosynthetic elucidation and structural revision of brevione E: characterization of the key dioxygenase for pathway branching from setosusin biosynthesis. Angew Chem Int Ed 61:e202210938
Macias FA, Varela RM, Simonet AM, Cutler HG, Cutler SJ, Dugan FM, Hill RA (2000) Novel bioactive breviane spiroditerpenoids from Penicillium brevicompactum Dierckx. J Org Chem 65:9039–9046
Wei X, Matsuyama T, Sato H, Yan D, Chan PM, Miyamoto K, Uchiyama M, Matsuda Y (2021) Molecular and computational bases for spirofuranone formation in setosusin biosynthesis. J Am Chem Soc 143:17708–17715
Acknowledgements
This work was supported by grants from the Ministry of Education, Culture, Sports, Science and Technology, Japan (JSPS KAKENHI Grant Number JP20KK0173, JP22H05123, JP23K13847, JP23H02641, and JP23H00393), Kobayashi Foundation, Koyanagi Foundation, Astellas Foundation, Mochida Memorial Foundation, Naito Foundation, Japan Foundation for Applied Enzymology, Amano Enzyme Foundation, Takeda Science Foundation, Sumitomo Foundation, and Suzuken Memorial Foundation.
Funding
Open access funding provided by The University of Tokyo.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing financial interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ushimaru, R. Three-membered ring formation catalyzed by α-ketoglutarate-dependent nonheme iron enzymes. J Nat Med 78, 21–32 (2024). https://doi.org/10.1007/s11418-023-01760-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11418-023-01760-4