Abstract
Background
Ferroptosis is an iron-dependent non-apoptotic programmed cell death. However, the regulatory mechanism of ferroptosis in colorectal cancer (CRC) is still unclear.
Objective
The aim of this study was to investigate the role and mechanism of enhancer-controlled genes in ferroptosis in CRC.
Methods
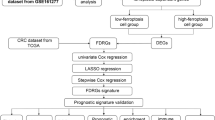
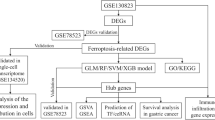
Dimensionality reduction and differentially expressed genes (DEGs) identification were conducted using Seurat algorithm based on single-cell RNA sequencing (scRNA-seq) data from the GSE200997 dataset. Ferroptosis-related pathway enrichment analysis was performed using the FerrDb V2 database. Enhancers were identified using HOMER algorithm based on H3K27ac ChIP-seq data from the GSE166254 dataset. Kaplan-Meier Plotter online tool was used to analyze prognosis and gene expression correlation. Transcription factors were predicted using the transcription factor affinity prediction web tool. The binding of enhancer to transcription factor and H3K27ac enrichment were detected by ChIP-qPCR. RSL3 was used to induce ferroptosis in CRC cells. Gene transcription was detected by qRT-PCR. Cell proliferation was detected by CCK8 assay.
Results
Nine cell clusters including T cells, natural killer cells, macrophages, mast cells, epithelial cells, fibroblasts, goblet cells, B cells and dendritic cells were identified in CRC and normal colonic tissue samples. Compared to normal colonic tissue-derived epithelial cells, 1075 DEGs were screened in CRC tissue-derived epithelial cells. Ferroptosis-related pathway enrichment suggested that DEGs were associated with the regulation of ferroptosis. DPEP1, ETV4, CEBPG, TIMP1, DUOX2 and LCN2 were identified as the significantly upregulated genes enriched in the “ferroptosis regulator” term, and their H3K27ac signals were significantly higher in CRC tissues than in normal colonic tissues. Of these, only the expression of TIMP1 predicted a poor prognosis of CRC patients. Transcription factor SPI1 drove TIMP1 transcription by binding to its enhancer. Overexpression of TIMP1 significantly promoted the resistance to ferroptosis induced by RSL3 in CRC cells, which was partially restored by SPI1 knockdown.
Conclusion
Transcription of TIMP1 was driven by transcription factor SPI1 in combination with its enhancer, consequently promoting CRC cells against ferroptosis. The SPI1/TIMP1 axis confers ferroptosis resistance in CRC, and thus has the potential to be the molecular targets for CRC treatment.






Similar content being viewed by others
Data Availability
The data generated or analyzed during this study can be obtained by contacting the corresponding author under reasonable request.
References
Banskota S, Regmi SC, Kim JA (2015) NOX1 to NOX2 switch deactivates AMPK and induces invasive phenotype in colon Cancer cells through overexpression of MMP-7. Mol Cancer 14:123
Becht E, McInnes L, Healy J, Dutertre CA, Kwok IWH, Ng LG, Ginhoux F, Newell EW (2018) Dimensionality reduction for visualizing single-cell data using UMAP. Nat Biotechnol
Cancer Genome Atlas N (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487:330–337
Chen X, Li J, Kang R, Klionsky DJ, Tang D (2021) Ferroptosis: machinery and regulation. Autophagy 17:2054–2081
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA et al (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A 107:21931–21936
Cui HY, Wei W, Qian MR, Tian RF, Fu X, Li HW, Nan G, Yang T, Lin P, Chen X et al (2022) PDGFA-associated protein 1 is a novel target of c-Myc and contributes to Colorectal cancer initiation and progression. Cancer Commun (Lond) 42:750–767
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE, Patel DN, Bauer AJ, Cantley AM, Yang WS et al (2012) Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149:1060–1072
Donati B, Lorenzini E, Ciarrocchi A (2018) BRD4 and Cancer: going beyond transcriptional regulation. Mol Cancer 17:164
Estevao D, da Cruz-Ribeiro M, Cardoso AP, Costa AM, Oliveira MJ, Duarte TL, da Cruz TB (2023) Iron metabolism in Colorectal cancer: a balancing act. Cell Oncol (Dordr
Gao N, Ye B (2020) SPI1-induced upregulation of lncRNA SNHG6 promotes non-small cell Lung cancer via miR-485-3p/VPS45 axis. Biomed Pharmacother 129:110239
Gao W, Huang Z, Duan J, Nice EC, Lin J, Huang C (2021) Elesclomol induces copper-dependent ferroptosis in Colorectal cancer cells via degradation of ATP7A. Mol Oncol 15:3527–3544
Guo L, Wang Y, Yang W, Wang C, Guo T, Yang J, Shao Z, Cai G, Cai S, Zhang L et al (2023) Molecular Profiling Provides Clinical Insights Into Targeted and Immunotherapies as Well as Colorectal Cancer Prognosis. Gastroenterology
Han Y, Gao X, Wu N, Jin Y, Zhou H, Wang W, Liu H, Chu Y, Cao J, Jiang M et al (2022) Long noncoding RNA LINC00239 inhibits ferroptosis in Colorectal cancer by binding to Keap1 to stabilize Nrf2. Cell Death Dis 13:742
He Q, Ye A, Ye W, Liao X, Qin G, Xu Y, Yin Y, Luo H, Yi M, Xian L et al (2021) Cancer-secreted exosomal mir-21-5p induces angiogenesis and vascular permeability by targeting KRIT1. Cell Death Dis 12:576
He Y, Lin L, Ou Y, Hu X, Xu C, Wang C (2022) Endothelial cell-specific molecule 1 (ESM1) promoted by transcription factor SPI1 acts as an oncogene to modulate the malignant phenotype of endometrial cancer. Open Med (Wars) 17:1376–1389
Heinz S, Romanoski CE, Benner C, Glass CK (2015) The selection and function of cell type-specific enhancers. Nat Rev Mol Cell Biol 16:144–154
Hou G, Zuo H, Shi J, Dai D, Wang H, Song X, Xu G, Tao G (2023) EIF4A3 induced circABCA5 promotes the gastric cancer progression by SPI1 mediated IL6/JAK2/STAT3 signaling. Am J Cancer Res 13:602–622
Ju HQ, Lu YX, Chen DL, Tian T, Mo HY, Wei XL, Liao JW, Wang F, Zeng ZL, Pelicano H et al (2016) Redox Regulation of stem-like cells though the CD44v-xCT Axis in Colorectal Cancer: mechanisms and therapeutic implications. Theranostics 6:1160–1175
Kaikkonen MU, Spann NJ, Heinz S, Romanoski CE, Allison KA, Stender JD, Chun HB, Tough DF, Prinjha RK, Benner C et al (2013) Remodeling of the enhancer landscape during macrophage activation is coupled to enhancer transcription. Mol Cell 51:310–325
Koppula P, Zhuang L, Gan B (2021) Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell 12:599–620
Lao VV, Grady WM (2011) Epigenetics and Colorectal cancer. Nat Rev Gastroenterol Hepatol 8:686–700
Levine M (2010) Transcriptional enhancers in animal development and evolution. Curr Biol 20:R754–763
Li X, Liang L, Huang L, Ma X, Li D, Cai S (2015) High expression of protein phosphatase 4 is associated with the aggressive malignant behavior of colorectal carcinoma. Mol Cancer 14:95
Liu B, Liu X, Han L, Chen X, Wu X, Wu J, Yan D, Wang Y, Liu S, Shan L et al (2022a) BRD4-directed super-enhancer organization of transcription repression programs links to chemotherapeutic efficacy in Breast cancer. Proc Natl Acad Sci U S A 119
Liu MY, Li HM, Wang XY, Xia R, Li X, Ma YJ, Wang M, Zhang HS (2022b) TIGAR drives Colorectal cancer ferroptosis resistance through ROS/AMPK/SCD1 pathway. Free Radic Biol Med 182:219–231
Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, Wang L, Lu T, Zhang Y, Sun Z et al (2022c) Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in Colorectal cancer. Nat Commun 13:816
Liu Y, Wan Y, Jiang Y, Zhang L, Cheng W (2023) GPX4: the hub of lipid oxidation, ferroptosis, Disease and treatment. Biochim Biophys Acta Rev Cancer 1878:188890
Luo D, Ge W (2020) MeCP2 promotes Colorectal Cancer Metastasis by modulating ZEB1 transcription. Cancers (Basel) 12
Nabet B, Reyes POB, Shieh JM, Lin K, Will CY, Popovic CM, Ezponda R, Bradner T, Golden JE AA et al (2015) Deregulation of the ras-Erk Signaling Axis modulates the enhancer Landscape. Cell Rep 12:1300–1313
Ostuni R, Piccolo V, Barozzi I, Polletti S, Termanini A, Bonifacio S, Curina A, Prosperini E, Ghisletti S, Natoli G (2013) Latent enhancers activated by stimulation in differentiated cells. Cell 152:157–171
Ries C (2014) Cytokine functions of TIMP-1. Cell Mol Life Sci 71:659–672
Schwalm MP, Knapp S (2022) BET bromodomain inhibitors. Curr Opin Chem Biol 68:102148
Shen L, Zhang J, Xu M, Zheng Y, Wang M, Yang S, Qin B, Li S, Dong L, Dai F (2022) DDX3 acts as a Tumor suppressor in Colorectal cancer as loss of DDX3 in advanced cancer promotes Tumor progression by activating the MAPK pathway. Int J Biol Sci 18:3918–3933
Shintoku R, Takigawa Y, Yamada K, Kubota C, Yoshimoto Y, Takeuchi T, Koshiishi I, Torii S (2017) Lipoxygenase-mediated generation of lipid peroxides enhances ferroptosis induced by erastin and RSL3. Cancer Sci 108:2187–2194
Sui X, Zhang R, Liu S, Duan T, Zhai L, Zhang M, Han X, Xiang Y, Huang X, Lin H et al (2018) RSL3 drives ferroptosis through GPX4 inactivation and ROS production in Colorectal Cancer. Front Pharmacol 9:1371
Thomas-Chollier M, Hufton A, Heinig M, O’Keeffe S, Masri NE, Roider HG, Manke T, Vingron M (2011) Transcription factor binding predictions using TRAP for the analysis of ChIP-seq data and regulatory SNPs. Nat Protoc 6:1860–1869
Wang L, Xu M, Kao CY, Tsai SY, Tsai MJ (2020) Small molecule JQ1 promotes Prostate cancer invasion via BET-independent inactivation of FOXA1. J Clin Invest 130:1782–1792
Wang Y, Zhang Z, Sun W, Zhang J, Xu Q, Zhou X, Mao L (2022) Ferroptosis in Colorectal cancer: potential mechanisms and effective therapeutic targets. Biomed Pharmacother 153:113524
Wang L, Wang J, Chen L (2023) TIMP1 represses sorafenib-triggered ferroptosis in Colorectal cancer cells by activating the PI3K/Akt signaling pathway. Immunopharmacol Immunotoxicol: 1–7
Wei R, Zhao Y, Wang J, Yang X, Li S, Wang Y, Yang X, Fei J, Hao X, Zhao Y et al (2021) Tagitinin C induces ferroptosis through PERK-Nrf2-HO-1 signaling pathway in Colorectal cancer cells. Int J Biol Sci 17:2703–2717
Yang WS, SriRamaratnam R, Welsch ME, Shimada K, Skouta R, Viswanathan VS, Cheah JH, Clemons PA, Shamji AF, Clish CB et al (2014) Regulation of ferroptotic cancer cell death by GPX4. Cell 156:317–331
Yang J, Mo J, Dai J, Ye C, Cen W, Zheng X, Jiang L, Ye L (2021) Cetuximab promotes RSL3-induced ferroptosis by suppressing the Nrf2/HO-1 signalling pathway in KRAS mutant Colorectal cancer. Cell Death Dis 12:1079
Yoshino S, Yokoyama T, Sunami Y, Takahara T, Nakamura A, Yamazaki Y, Tsutsumi S, Aburatani H, Nakamura T (2021) Trib1 promotes acute Myeloid Leukemia progression by modulating the transcriptional programs of Hoxa9. Blood 137:75–88
Young RS, Kumar Y, Bickmore WA, Taylor MS (2017) Bidirectional transcription initiation marks accessible chromatin and is not specific to enhancers. Genome Biol 18:242
Zhou N, Yuan X, Du Q, Zhang Z, Shi X, Bao J, Ning Y, Peng L (2023) FerrDb V2: update of the manually curated database of ferroptosis regulators and ferroptosis-disease associations. Nucleic Acids Res 51:D571–D582
Funding
This study was supported by the Science and Technology Extension Project of Hebei Provincial Health Commission Medical Science Research Fund Program (20220124).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M., Ni, Qy. & Yu, Sy. Integration of single-cell transcriptomics and epigenetic analysis reveals enhancer-controlled TIMP1 as a regulator of ferroptosis in colorectal cancer. Genes Genom 46, 121–133 (2024). https://doi.org/10.1007/s13258-023-01474-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01474-7