Abstract
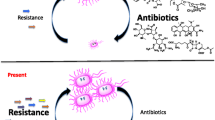
Antibiotics have been used in clinical treatment since the mid-20th century. With the use of antibiotics, infant mortality has decreased and average life has increased by about 20 years. However, since the early years of antibiotic use, bacteria have begun to change to reduce or eliminate the effectiveness of antibiotics. Antibiotic resistance is spreading rapidly, but the mechanisms of resistance acquisition, how resistance arises, and how it can be prevented are not clearly understood. The purpose of this review is to compile information on studies of antibiotic resistance and the prevention of resistance acquisition. Spontaneous mutations are an important cause of resistance acquisition in the presence of antibiotics. One of the most commonly used methods to study how these mutations arise is adaptive resistance experiments. Using the information obtained from these experiments, it has been determined that the SOS response plays an important role in the acquisition of resistance mutations. Therefore, the SOS response could be an important target for inhibiting the acquisition of antibiotic resistance.
Similar content being viewed by others
REFERENCES
Abat, C., Fournier, P.E., Jimeno, M.T., Rolain, J.M., and Raoult, D., Extremely and pandrug-resistant bacteria extra-deaths: Myth or reality?, Eur. J. Clin. Microbiol. Infect. Dis., 2018, vol. 37, no. 9, pp. 1687–1697. https://doi.org/10.1007/s10096-018-3300-0
Agyare, C., Etsiapa Boamah, V., Ngofi Zumbi, C., and Boateng Osei, F., Antibiotic use in poultry production and its effects on bacterial resistance, IntechOpen, 2018. https://doi.org/10.5772/intechopen.73725
Aminov, R.I., A brief history of the antibiotic era: Lessons learned and challenges for the future, Front. Microbiol., 2010, vol. 1, p. 134. https://doi.org/10.3389/fmicb.2010.00134
Appelbaum, P.C., The emergence of vancomycin-intermediate and vancomycin-resistant Staphylococcus aureus, Clin. Microbiol. Infect., 2006, vol. 12, no. 1, pp. 16–23. https://doi.org/10.1111/j.1469-0691.2006.01344.x
Arias, C.A., Panesso, D., McGrath, D.M., Qin, X., et al., Genetic basis for in vivo daptomycin resistance in enterococci, N. Engl. J. Med., 2011, vol. 365, no. 10, pp. 892–900. https://doi.org/10.1056/nejmoa1011138
Arnold, B.J., Huang, I.T., and Hanage, W.P., Horizontal gene transfer and adaptive evolution in bacteria, Nat. Rev. Microbiol., 2021, vol. 20, pp. 206–218. https://doi.org/10.1038/s41579-021-00650-4
Barr, V., Barr, K., Millar, M.R., and Lacey, R.W., β-Lactam antibiotics increase the frequency of plasmid transfer in Staphylococcus aureus, J. Antimicrob. Chemother., 1986, vol. 17, no. 4, pp. 409–413. https://doi.org/10.1093/jac/17.4.409
Baym, M., Lieberman, T.D., Kelsic, E.D., Chait, R., et al., Spatiotemporal microbial evolution on antibiotic landscapes, Science, 2016, vol. 353, no. 6304, pp. 1147–1151. https://doi.org/10.1126/science.aag0822
Blair, J.M., Webber, M.A., Baylay, A.J., and Ogbolu, D.O., and Piddock, L.J., Molecular mechanisms of antibiotic resistance, Nat. Rev. Microbiol., 2015, vol. 13, pp. 42–51. https://doi.org/10.1038/nrmicro3380
Blázquez, J., Couce, A., Rodríguez-Beltrán, J., and Rodríguez-Rojas, A., Antimicrobials as promoters of genetic variation, Curr. Opin. Microbiol., 2012, vol. 15, no. 5, pp. 561–569. https://doi.org/10.1016/j.mib.2012.07.007
Bore, E., Hébraud, M., Chafsey, I., Chambon, C., et al., Adapted tolerance to benzalkonium chloride in Escherichia coli K-12 studied by transcriptome and proteome analyses, Microbiology, 2007, vol. 153, no. 4, p. 935–946. https://doi.org/10.1099/mic.0.29288-0
Briales, A., Rodriguez-Martinez, J.M., Velasco, C., Machuca, J., et al., Exposure to diverse antimicrobials induces the expression of qnrB1, qnrD and smaqnr genes by SOS-dependent regulation, J. Antimicrob. Chemother., 2012, vol. 67, no. 12, p. 2854–2859. https://doi.org/10.1093/jac/dks326
Brito, I.L., Examining horizontal gene transfer in microbial communities, Nat. Rev. Microbiol., 2021, vol. 19, no 7, p. 442–453. https://doi.org/10.1038/s41579-021-00534-7
Burnham, J.P., Olsen, M.A., and Kollef, M.H., Re-estimating annual deaths due to multidrug-resistant organism infections, Infect. Control Hosp. Epidemiol., 2019, vol. 40, no. 1, pp. 112–113. https://doi.org/10.1017/ice.2018.304
Chambers, H.F. and DeLeo, F.R., Waves of resistance: Staphylococcus aureus in the antibiotic era, Nat. Rev. Microbiol., 2009, vol. 7, no. 9, pp. 629–641. https://doi.org/10.1038/nrmicro2200
Chellat, M.F., Raguž, L., and Riedl, R., Targeting antibiotic resistance, Angew. Chem., Int. Ed., 2016, vol. 55, no. 23, pp. 6600–6626. https://doi.org/10.1002/anie.201506818
Crane, J.K., Alvarado, C.L., and Sutton, M.D., Role of the SOS response in the generation of antibiotic resistance in vivo, Antimicrob. Agents Chemother., 2021, vol. 65, no. 7, pp. 1–17. https://doi.org/10.1128%2FAAC.00013-21
Cully, M., Public health: The politics of antibiotics, Nature, 2014, vol. 509, pp. 16–17. https://doi.org/10.1038/509S16a
Darcan, C. and Kahyaoglu, M., The effect of some boron derivatives on Kanamycin Resistance and survival of E. coli and P. aeruginosa in Lake Water, Biomed. Environ. Sci., 2012, vol. 25, no. 4, pp. 476–482. https://doi.org/10.3967/0895-3988.2012.04.014
Delcour, A.H., Outer membrane permeability and antibiotic resistance, Biochim. Biophys. Acta, 2009, vol. 1794, no. 5, pp. 808–816. https://doi.org/10.1016/j.bbapap.2008.11.005
Dersch, P., Khan, M.A., Mühlen, S., and Görke, B., Roles of regulatory RNAs for antibiotic resistance in bacteria and their potential value as novel drug targets, Front. Microbiol., 2017, vol. 8, p. 803. https://doi.org/10.3389/fmicb.2017.00803
Diaz-Diaz, S. and Recacha, E., MacHuca, J., Garcia-Duque, A., et al., Synergistic quinolone sensitization by targeting the recA SOS response gene and oxidative stress, Antimicrob. Agents Chemother., 2021, vol. 65, no. 4, pp. 1–11. https://doi.org/10.1128/aac.02004-20
Dong, T. and Schellhorn, H.E., Role of RpoS in virulence of pathogens, Infect. Immun., 2010, vol. 78, no. 3, pp. 887–897. https://doi.org/10.1128/iai.00882-09
Doucet-Populaire, F., Trieu-Cuot, P., Dosbaa, I., Andremont, A., et al., Inducible transfer of conjugative transposon Tn1545 from Enterococcus faecalis to Listeria monocytogenes in the digestive tracts of gnotobiotic mice, Antimicrob. Agents Chemother., 1991, vol. 35, no. 1, p. 185–187. https://doi.org/10.1128/aac.35.1.185
Du, B., Olson, C.A., and Sastry, A., v., Fang, X., et al., Adaptive laboratory evolution of Escherichia coli under acid stress, Microbiology, 2020, vol. 166, no. 2, pp. 141–148. https://doi.org/10.1099/mic.0.000867
Garau, J., Xercavins, M., Rodríguez-Carballeira, M., Gómez-Vera, J.R., et al., Emergence and dissemination of quinolone-resistant Escherichia coli in the community, Antimicrob. Agents Chemother., 1999, vol. 43, no. 11, pp. 2736–2741. https://doi.org/10.1128/aac.43.11.2736
Gutierrez, A., Laureti, L., Crussard, S., Abida, H., et al., β‑Lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity, Nat. Commun., 2013, vol. 4, pp. 1–9. https://doi.org/10.1038/ncomms2607
Gutiérrez, R., Ram, Y., Berman, J., de Sousa, K.C.M., et al., Adaptive resistance mutations at suprainhibitory concentrations independent of SOS mutagenesis, Mol. Biol. Evol., 2021, vol. 38, no. 10, pp. 4095–4115. https://doi.org/10.1093/molbev/msab196
Hoeksema, M., Jonker, M.J., and Brul, S., ter Kuile, B. H., Effects of a previously selected antibiotic resistance on mutations acquired during development of a second resistance in Escherichia coli, BMC Genomics, 2019, vol. 20, p. 1–14. https://doi.org/10.1186%2Fs12864-019-5648-7
Hosain, Z., Kabir, S.M.L., and Kamal, M., Antimicrobial uses for livestock production in developing countries, Vet. World, 2021, vol. 14, no. 1, pp. 210–221. https://doi.org/10.14202/vetworld.2021.210-221
Howden, B.P., McEvoy, C.R.E., Allen, D.L., Chua, K., et al., Evolution of multidrug resistance during Staphylococcus aureus infection involves mutation of the essential two component regulator WalKR, PLoS Pathog., 2011, vol. 7, no. 11, pp. 1–15. https://doi.org/10.1371/journal.ppat.1002359
Huang, F., Motlekar, N.A., Burgwin, C.M., Napper, A.D., et al., Identification of specific inhibitors of human, RAD51 recombinase using high-throughput screening, ACS Chem. Biol., 2011, vol. 6, no. 6, p. 628–635. https://doi.org/10.1021/cb100428c
Hutchings, M., Truman, A., Wilkinson, B., et al., Antibiotics: Past, present and future, Curr. Opin. Microbiol., 2019, vol. 51, pp. 72–80. https://doi.org/10.1016/j.mib.2019.10.008
Jevons, M.P., “Celbenin”—resistant Staphylococci, Br. Med. J., 1961, vol. 1, no. 5219, pp. 124–125.
Jo, S.B., Shin, C.H., Shin, Y.J., Kim, P.H., et al., Heavy metal and antibiotic co-resistance in Vibrio parahaemolyticus isolated from shellfish, Mar. Pollut. Bull., 2020, vol. 156, p. 111246. https://doi.org/10.1016/j.marpolbul.2020.111246
Jutkina, J., Marathe, N.P., Flach, C.F., and Larsson, D.G.J., Antibiotics and common antibacterial biocides stimulate horizontal transfer of resistance at low concentrations, Sci. Total Environ., 2018, vols. 616–617, pp. 172–178. https://doi.org/10.1016/j.scitotenv.2017.10.312
Kang, W., Zhang, Y.-J., Shi, X., He, J.-Z., and Hu, H.-W., Short-term copper exposure as a selection pressure for antibiotic resistance and metal resistance in an agricultural soil, Environ. Sci. Pollut. Res., 2018, vol. 25, pp. 29314–29324. https://doi.org/10.1007/s11356-018-2978-y
Knöppel, A., Näsvall, J., and Andersson, D.I., Evolution of antibiotic resistance without antibiotic exposure, Antimicrob. Agents Chemother., 2017, vol. 61, no. 11, pp. 1–5. https://doi.org/10.1128/aac.01495-17
Kohanski, M.A., DePristo, M.A., and Collins, J.J., Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis, Mol. Cell, 2010, vol. 37, no. 3, pp. 311–320. https://doi.org/10.1016/j.molcel.2010.01.003
Kucukyildirim, S., Whole-population genomic sequencing reveals the mutational profiles of the antibiotic-treated Escherichia coli populations, Biologia, 2022, vol. 77, pp. 525–531. https://doi.org/10.1007/s11756-021-00959-8
Kuile, B.H., Kraupner, N., and Brul, S., The risk of low concentrations of antibiotics in agriculture for resistance in human health care, FEMS Microbiol. Lett., 2016, vol. 363, no. 19, p. 210. https://doi.org/10.1093/femsle/fnw210
Kurenbach, B., Hill, A.M., Godsoe, W., van Hamelsveld, S., and Heinemann, J.A., Agrichemicals and antibiotics in combination increase antibiotic resistance evolution, PeerJ, 2018, vol. 6, p. e5801. https://doi.org/10.7717%2Fpeerj.5801
Lalaouna, D., Eyraud, A., Chabelskaya, S., Felden, B., and Massé, E., Regulatory RNAs involved in bacterial antibiotic resistance, PLoS Pathog., 2014, vol. 10, p. e1004299. https://doi.org/10.1371/journal.ppat.1004299
Lamrabet, O., Martin, M., Lenski, R.E., and Schneider, D., Changes in intrinsic antibiotic susceptibility during a long-term evolution experiment with escherichia coli, MBio, 2019, vol. 10, no. 2, pp. 1–12. https://doi.org/10.1128/mbio.00189-19
Lázár, V., Pal Singh, G., Spohn, R., Nagy, I., et al., Bacterial evolution of antibiotic hypersensitivity, Mol. Syst. Biol., 2013, vol. 9, p. 700. https://doi.org/10.1038/msb.2013.57
Lee, J.-Y., Seo, J., Kim, E.-S., Lee, H.-S., and Kim, P., Adaptive evolution of Corynebacterium glutamicum resistant to oxidative stress and its global gene expression profiling, Biotechnol. Lett., 2013, vol. 35, no. 5, pp. 709–717. https://doi.org/10.1007/s10529-012-1135-9
Léger, L., Budin-Verneuil, A., Cacaci, M., Benachour, A., et al., β-Lactam exposure triggers reactive oxygen species formation in Enterococcus faecalis via the respiratory chain component DMK, Cell Rep., 2019, vol. 29, no. 8, pp. 2184–2191. https://doi.org/10.1016/j.celrep.2019.10.080
Lerminiaux, N.A. and Cameron, A.D.S., Horizontal transfer of antibiotic resistance genes in clinical environments, Can. J. Microbiol., 2019, vol. 65, pp. 34–44. https://doi.org/10.1139/cjm-2018-0275
Li, G.Q., Quan, F., Qu, T., Lu, J., et al., Sublethal vancomycin-induced ROS mediating antibiotic resistance in Staphylococcus aureus, Biosci. Rep., 2015, vol. 35, no. 6, p. 279. https://doi.org/10.1042/bsr20140167
Li, M., Liu, Q., Teng, Y., Ou, L., et al., The resistance mechanism of Escherichia coli induced by ampicillin in laboratory, Infect. Drug Resist., 2019, vol. 12, pp. 2853–2863. https://doi.org/10.2147%2FIDR.S221212
Li, X., Gu, A.Z., Zhang, Y., Xie, B., et al., Sub-lethal concentrations of heavy metals induce antibiotic resistance via mutagenesis, J. Hazard. Mater., 2019, vol. 369, pp. 9–16. https://doi.org/10.1016/j.jhazmat.2019.02.006
Lindsey, H.A., Gallie, J., Taylor, S., and Kerr, B., Evolutionary rescue from extinction is contingent on a lower rate of environmental change, Nature, 2013, vol. 494, pp. 463–467. https://doi.org/10.1038/nature11879
Lowy, F.D., Antimicrobial resistance: The example of Staphylococcus aureus, J. Clin. Invest., 2003, vol. 111, no. 9, pp. 1265–1273. https://doi.org/10.1172/JCI18535
MacLean, R.C. and Millan, A.S., The evolution of antibiotic resistance, Science, 2019, vol. 365, no. 6458, pp. 1082–1083. https://doi.org/10.1126/science.aax3879
Maeda, T., Horinouchi, T., Sakata, N., Sakai, A., and Chikara, F., High-throughput identification of the sensitivities of an Escherichia coli recA mutant strain to various chemical compounds, J. Antibiot., 2019, vol. 72, pp. 566–573. https://doi.org/10.1038/s41429-019-0160-5
Maeda, T., Iwasawa, J., Kotani, H., Sakata, N., et al., High-throughput laboratory evolution reveals evolutionary constraints in Escherichia coli, Nat. Commun., 2020, vol. 11, no, 11, p. 1–13. https://doi.org/10.1038/s41467-020-19713-w
Malhotra-Kumar, S., Xavier, B.B., Das, A.J., Lammens, C., et al., Colistin resistance gene mcr-1 harboured on a multidrug resistant plasmid, Lancet Infect. Dis., 2016, vol. 16, no. 3, p. 283–284. https://doi.org/10.1016/S1473-3099(16)00012-8
Mandin, P. and Gottesman, S., Integrating anaerobic/aerobic sensing and the general stress response through the ArcZ small RNA, EMBO J., 2010, vol. 29, no. 18, pp. 3094–3107. https://doi.org/10.1038/emboj.2010.179
Manna, M.S., Tamer, Y.T., Gaszek, I., Poulides, N., et al., A trimethoprim derivative impedes antibiotic resistance evolution, Nat. Commun., 2021, vol. 12, pp. 1–10. https://doi.org/10.1038/s41467-021-23191-z
Mans, R., Daran, J.M.G., and Pronk, J.T., Under pressure: Evolutionary engineering of yeast strains for improved performance in fuels and chemicals production, Curr. Opin. Biotechnol., 2018, vol. 50, pp. 47–56. https://doi.org/10.1016/j.copbio.2017.10.011
Marti, E., Variatza, E., and Balcazar, J.L., The role of aquatic ecosystems as reservoirs of antibiotic resistance, Trends Microbiol., 2014, vol. 22, no. 1, pp. 36–41. https://doi.org/10.1016/j.tim.2013.11.001
Martinez, J.L. and Baquero, F., Mutation frequencies and antibiotic resistance, Antimicrob. Agents Chemother., 2000, vol. 44, no. 7, pp. 1771–1777. https://doi.org/10.1128/aac.44.7.1771-1777.2000
Martínez, J.L. and Rojo, F., Metabolic regulation of antibiotic resistance, FEMS Microbiol. Rev., 2011, vol. 35, no. 5, pp. 768–789. https://doi.org/10.1111/j.1574-6976.2011.00282.x
Mo, C.Y., Manning, S.A., Roggiani, M., Culyba, M.J., et al., Systematically altering bacterial SOS activity under stress reveals therapeutic strategies for potentiating antibiotics, MSphere, 2016, vol. 1, no. 4, pp. 1–15. https://doi.org/10.1128/msphere.00163-16
Nishino, K., Yamasaki, S., Hayashi-Nishino, M., and Yamaguchi, A., Effect of overexpression of small non-coding DsrA RNA on multidrug efflux in Escherichia coli, J. Antimicrob. Chemother., 2011, vol. 66, no. 2, pp. 291–296. https://doi.org/10.1093/jac/dkq420
Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations, WHO, 2014. https://www.who.int/news/item/29-04-2019-new-report-calls-for-urgent-action-to-avert-antimicrobial-resistance-crisis
Pereira, R., Wei, Y., Mohamed, E., Radi, M., et al., Adaptive laboratory evolution of tolerance to dicarboxylic acids in Saccharomyces cerevisiae, Metab. Eng., 2019, vol. 56, pp. 130–141. https://doi.org/10.1016/j.ymben.2019.09.008
Pinilla-Redondo, R., Cyriaque, V., Jacquiod, S., Sørensen, S.J., and Riber, L., Monitoring plasmid-mediated horizontal gene transfer in microbiomes: Recent advances and future perspectives, Plasmid, 2018, vol. 99, pp. 56–67. https://doi.org/10.1016/j.plasmid.2018.08.002
Recacha, E., Machuca, J., Díaz-Díaz, S., García-Duque, A., et al., Suppression of the SOS response modifies spatiotemporal evolution, post-antibiotic effect, bacterial fitness and biofilm formation in quinolone-resistant Escherichia coli, J. Antimicrob. Chemother., 2019, vol. 74, no. 1, pp. 66–73. https://doi.org/10.1093/jac/dky407
Reygaert, W.C., An overview of the antimicrobial resistance mechanisms of bacteria, AIMS Microbiol., 2018, vol. 4, no. 3, pp. 482–501. https://doi.org/10.3934/microbiol.2018.3.482
Rodríguez-Rosado, A.I., Valencia, E.Y., Rodríguez-Rojas, A., Costas, C., et al., N-acetylcysteine blocks SOS induction and mutagenesis produced by fluoroquinolones in Escherichia coli, J. Antimicrob. Chemother., 2019, vol. 74, no. 8, pp. 2188–2196.https://doi.org/10.1093/jac/dkz210
Romandini, A., Pani, A., Schenardi, P.A., Angela, G., et al., Antibiotic resistance in pediatric infections: Global emerging threats, predicting the near future, Antibiotics, 2021, vol. 10, no. 4, p. 393. https://doi.org/10.3390/antibiotics10040393
Sahni, A., Hajjari, M., Raheb, J., Foroughmand, A.M., and Asgari, M., The non-coding RNA rprA can increase the resistance to ampicillin in Escherichia coli, Microb. Pathog., 2019, vol. 129, pp. 266–270. https://doi.org/10.1016/j.micpath.2019.02.021
Sandberg, T.E., Salazar, M.J., Weng, L.L., Palsson, B.O., and Feist, A.M., The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology, Metab. Eng., 2019, vol. 56, pp. 1–16. https://doi.org/10.1016/j.ymben.2019.08.004
Sandegren, L., Low sub-minimal inhibitory concentrations of antibiotics generate new types of resistance, Sustainable Chem. Pharm., 2019, vol. 11, pp. 46–48. https://doi.org/10.1016/j.scp.2018.12.006
Saunders, N.J., Trivedi, U.H., Thomson, M.L., Doig, C., et al., Deep resequencing of serial sputum isolates of Mycobacterium tuberculosis during therapeutic failure due to poor compliance reveals stepwise mutation of key resistance genes on an otherwise stable genetic background, J. Infect., 2011, vol. 62, no. 3, pp. 212–217. https://doi.org/10.1016/j.jinf.2011.01.003
Serwecińska, L., Antimicrobials and antibiotic-resistant bacteria: A risk to the environment and to public health, Water, 2020, vol. 12, no. 12, pp. 1–17. https://doi.org/10.3390/w12123313
Sevik, H., Cetin, M., Ozel, H.B., and Akarsu, H., Zeren Cetin, I., Analyzing of usability of tree-rings as biomonitors for monitoring heavy metal accumulation in the atmosphere in urban area: A case study of cedar tree (Cedrus sp.), Environ. Monit. Assess, 2020, vol. 192, no. 1, pp. 1–11. https://doi.org/10.1007/s10661-019-8010-2
Summers, A.O., Wireman, J., Vimy, M.J., Lorscheider, F.L., et al., Mercury released from dental “silver” fillings provokes an increase in mercury- and antibiotic-resistant bacteria in oral and intestinal floras of primates, Antimicrob. Agents Chemother., 1993, vol. 37, no. 4, pp. 825–834. https://doi.org/10.1128/aac.37.4.825
Sun, D., Jeannot, K., Xiao, Y., and Knapp, C.W., Editorial: Horizontal gene transfer mediated bacterial antibiotic resistance, Front. Microbiol., 2019, vol. 10, p. 1933. https://doi.org/10.3389/fmicb.2019.01933
Suzuki, S., Horinouchi, T., and Furusawa, C., Suppression of antibiotic resistance acquisition by combined use of antibiotics, J. Biosci. Bioeng., 2015, vol. 120, no. 4, pp. 467–469. https://doi.org/10.1016/j.jbiosc.2015.02.003
Suzuki, S., Horinouchi, T., and Furusawa, C., Acceleration and suppression of resistance development by antibiotic combinations, BMC Genomics, 2017, vol. 18, no. 1, pp. 1–10. https://doi.org/10.1186/s12864-017-3718-2
Tang, J., Zhang, J., Ren, L., Zhou, Y., et al., Diagnosis of soil contamination using microbiological indices: A review on heavy metal pollution, J. Environ. Manage, 2019, vol. 242, pp. 121–130. https://doi.org/10.1016/j.jenvman.2019.04.061
Toprak, E., Veres, A., Michel, J.-B., Chait, R., et al., Evolutionary paths to antibiotic resistance under dynamically sustained drug selection, Nat. Genet., 2012, vol. 44, no. 1, pp. 101–105. https://doi.org/10.1038%2Fng.1034
Valencia, A.O., Braz, V.S., Magalhães, M., and Galhardo, R.S., Role of error-prone DNA polymerases in spontaneous mutagenesis in Caulobacter crescentus, Genet. Mol. Biol., 2020, vol. 43, no. 1, p. e20180283. https://doi.org/10.1590/1678-4685-gmb-2018-0283
Viswanathan, V.K., Off-label abuse of antibiotics by bacteria, Gut Microbes, 2014, vol. 5, no. 1, pp. 3–4. https://doi.org/10.4161%2Fgmic.28027
Wang, Z., Zhang, P., Ding, X., Wang, J., et al., Co-delivery of ampicillin and β-lactamase inhibitor by selenium nanocomposite to achieve synergistic anti-infective efficiency through overcoming multidrug resistance, Chem. Eng. J., 2021, vol. 414, p. 128908. https://doi.org/10.1016/j.cej.2021.128908
Webber, M.A. and Piddock, L.J.V., The importance of efflux pumps in bacterial antibiotic resistance, J. Antimicrob. Chemother., 2003, vol. 51, no. 1, p. 9–11. https://doi.org/10.1093/jac/dkg050
Wright, G.D., Bacterial resistance to antibiotics: Enzymatic degradation and modification, Adv. Drug Delivery Rev., 2005, vol. 57, no. 10, pp. 1451–1470. https://doi.org/10.1016/j.addr.2005.04.002
Xiao, X., Zeng, F., Li, R., Liu, Y., and Wang, Z., Subinhibitory concentration of colistin promotes the conjugation frequencies of Mcr-1- and blaNDM-5-positive plasmids, Microbiol. Spectrum, 2022, vol. 10, no. 2, p. 1–9. https://doi.org/10.1128/spectrum.02160-21
Yakimov, A., Bakhlanova, I., and Baitin, D., Targeting evolution of antibiotic resistance by SOS response inhibition, Comput. Struct. Biotechnol. J., 2021, vol. 19, pp. 777–783. https://doi.org/10.1016/j.csbj.2021.01.003
Yu, S., Wang, Y., Shen, F., Fang, H., and Yu, Y., Copper-based fungicide copper hydroxide accelerates the evolution of antibiotic resistance via gene mutations in Escherichia coli, Sci. Total Environ., 2022, vol. 815, p. 152885. https://doi.org/10.1016/j.scitotenv.2021.152885
Zhang, Q., Lambert, G., Liao, D., Kim, H., et al., Acceleration of emergence of bacterial antibiotic resistance in connected microenvironments, Science, 2011, vol. 333, no. 6050, pp. 1764–1767. https://doi.org/10.1126/science.1208747
Zhuang, M., Achmon, Y., Cao, Y., Liang, X., et al., Distribution of antibiotic resistance genes in the environment, Environ. Pollut., 2021, vol. 285, no. 2, p. 117402. https://doi.org/10.1016/j.envpol.2021.117402
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
The authors declare that they have no conflict of interest in the publication.
The manuscript does not involve any animal study. Therefore, this study does not require ethical approval.
AUTHOR CONTRIBUTION
All authors contributed to the study conception and design. Literature review, writing, review, and editing were performed by OT and CD. The first draft of the manuscript was written by OT and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Rights and permissions
About this article
Cite this article
Osman Türkyılmaz, Cihan Darcan The Emergence and Preventability of Globally Spreading Antibiotic Resistance: A Literature Review. Biol Bull Rev 13, 578–589 (2023). https://doi.org/10.1134/S2079086423060154
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S2079086423060154