Abstract
Background
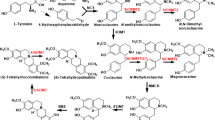
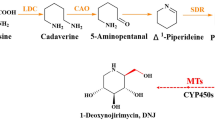
Secondary metabolites such as benzylisoquinoline alkaloids (BIA) have attracted considerable attention because of their pharmacological properties and potential therapeutic applications. Methyltransferases (MTs) can add methyl groups to alkaloid molecules, altering their physicochemical properties and bioactivity, stability, solubility, and recognition by other cellular components. Five types of O-methyltransferases and two types of N-methyltransferases are involved in BIA biosynthesis.
Objective
Since MTs may be the source for the discovery and development of novel biomedical, agricultural, and industrial compounds, we performed extensive molecular and phylogenetic analyses of O- and N-methyltransferases in BIA-producing plants.
Methods
MTs involved in BIA biosynthesis were isolated from transcriptomes of Berberis koreana and Caulophyllum robustum. We also mined the methyltransferases of Coptis japonica, Papaver somniferum, and Nelumbo nucifera from the National Center for Biotechnology Information protein database. Then, we analyzed the functional motifs and phylogenetic analysis.
Result
We mined 42 O-methyltransferases and 8 N-methyltransferases from the five BIA-producing plants. Functional motifs for S-adenosyl-L-methionine-dependent methyltransferases were retained in most methyltransferases, except for the three O-methyltransferases from N. nucifera. Phylogenetic analysis revealed that the methyltransferases were grouped into four clades, I, II, III and IV. The clustering patterns in the phylogenetic analysis suggested a monophyletic origin of methyltransferases and gene duplication within species. The coexistence of different O-methyltransferases in the deep branch subclade might support some cases of substrate promiscuity.
Conclusions
Methyltransferases may be a source for the discovery and development of novel biomedical, agricultural, and industrial compounds. Our results contribute to further understanding of their structure and reaction mechanisms, which will require future functional studies.

(Modified from Hagel et al. (2015)). Abbreviation: 3'-OHase, 3'-hydroxylase; 3'OMT, 3'-O-methyltransferase; 4HPPDC, 4-hydroxyphenylpyruvate decarboxylase; 4'OMT, 3'-hydroxyl-N-methylcoclaaurine 4'-O-methyltransferase; 6OMT, nocolaurine 6-O-methyltransferase, AT1, 1,13-dihydroxy-N-methycanadine 13-O-acetyltransferase; BBE, berberine bridge enzyme; BS, berbamunine synthase; CAS, canadine synthase; CNMT, coclaurine N-methyltransferase; CODN, codein O-demethylase; CoOMT, columbamine O-methyltransferase; COR, codeinone reductase; CTS, corytuberine synthase; CYP82X1, 1-hydroxyl-13-O-acetyl-N-methetylcanadine 8-hydroxylase; CYP82X2, 1-hydroxyl-N-methylcanadine 13-dehydroxylase; CYP82Y1, N-methylcanadine 1-hydroxylase; CDBOX, dihydrobenzophenanthridine oxidase; CXE, 3-O-acetypapaveroxine carboxylase; DBOX, dihydrobenzophenathridine oxidase; LdM, laudanisine demethylase; MSH, N-methylstylopine hydroxylase; N7OMT, norreticuline 7-O-methyltransferase; NCS, norcoclaurine synthase; NMCanH,N-methylcanadine 1-hydroxylase; NMCH, N-methylcocaurine 3'hydroxylase; NOS, noscapine synthase; P6H, protopine 6-hydroxylase; REPI, reticuline epimerase; SalAT, salutaridiol 7-O-acetyltransferase; SalR, salutaridine reductase; SalSyn, salutaridine synthase; SanR, Sanguinarine reductase; SOMT, scoulerine 9-O-methy;transferase; SPS, stylopine synthase; STOX, (S)-tetrahydroprotoberberine oxidase; T6ODM, thebaine 6-O-demethylase; TNMT, tetrahydroprotoberberine N-methyltransferase; TYDC, tyrosine decarboxylase; TyrAT, tyrosine aminotransferase




Similar content being viewed by others
References
Abdelraheem E, Thair B, Valera RF, Jockmann E, Popadic D, Hailes HC, Ward JM, Iribarren AM, Lewkowicz Andexer JN et al (2022) Methyltransferase: Function andapplications. Chem Bio Chem 23:e2022200212. https://doi.org/10.1002/cbic.202200212
Battersby AR, Binks R, Francis RJ, McCaldin DJ, Ramuz H (1964) Alkaloid biosynthesis. Part IV. 1-Benzylisoquinolines as precursors of thebaine, codeine, and morphine. J Chem Soc. https://doi.org/10.1039/jr9640003600
Beaudoin GAW, Facchini PJ (2014) Bezylisoqionoline alkaloid biosynthesis in opium poppy. Planta 240:19–32. https://doi.org/10.1007/s00425-014-2056-8
Bu J, Zhang X, Li Q, Ma Y, Yang J, Liu X, Wang R, Jiao X, Chen T, Lai C et al (2022) Catalytic promiscuity of O-methyltransferases from Corydalis yanhusuo leading to the structural diversity of benzylisoquinoline alkaloids. Hortic Res. https://doi.org/10.1093/hr/uhac152
Deng X, Zhao L, Fang T, Xiong Y, Ogutu C, Yang D, Vimolmangang S, Liu Y, Han Y (2018) Investigation of benzylisoquinoline biosynthetic pathway and its transcriptional regulation in lotus. Hortic Res 5:29. https://doi.org/10.1038/s41438-018-0035-0
Facchini PJ, Bird DA (1998) Development of regulation of benzylisoquinoline alkaloid biosynthesis in opium poppy plants and tissue cultures. In Vitro Cell Dev Biol 34:69–79. https://doi.org/10.1007/BF02823126
Hagel JM, Facchini PT (2013) Benzylisoquinoline alkaloid metabolism: a century of discovery and a brave new world. Plant Cell Physiol 54:647–672. https://doi.org/10.1093/pcp/pct020
Hagel JM, Morris JS, Lee E-J, Desgagne-Penix I, Bross CD, Chang L, Chen X, Farrow SC, Zhang Y, Soh J, Sensen CW, Facchini PJ (2015) Transcriptome analysis of 20 taxonomically related benzylisoquinoline alkaloid-producing plants. BMC Plant Biol 15:227. https://doi.org/10.1186/s12870-015-0596-0
Han X, Lamshoft M, Grobe N, Ren X, Fist AJ, Kutchan TM, Spiteller M, Zenk MH (2010) The biosynthesis of papaverine proceeds vis (S)-reticuline. Phytochemistry 71:1305–1312. https://doi.org/10.1016/j.phytochem.2010.04.022
He SM, Liang YL, Cong G, Chen G, Zhao QM, Zhang J-J, Wang X et al (2018) Identification and characterization of genes involved in benzylisoquinoline alkaloid biosynthesis in Coptis species. Front Plant Sci 9:2018. https://doi.org/10.3389/fpls.2018.00731
Höberg J, Saenz-Mendez P, Eriksson L (2018) QM/MM studies of Dph5 – A promiscuous methyltransferase in eukaryotic biosynthetic pathway of diphthamide. J Chem Inf Model 58:1406–1414. https://doi.org/10.1021/acs.jcim.8b00217
Ibrahim RK, Bruneau A, Bantignies B (1998) Plant O-methyltransferases: molecular analysis, common signature, and classification. Plant Mol Biol 36:1–10. https://doi.org/10.1023/a:1005939803300
Koirala N, Nguyen HT, Gopal PG, Duong VT, Sohng JK (2016) Methylation of flavonoids: Chemical structures, bioactivities, progress and perspectives for biotechnological production. Enzym Microb Tech 86:103–116. https://doi.org/10.1016/j.enzmictec.2016.02.003
Kozbial PZ, Mushegian AR (2005) Natural history of S-adenylmethionine-binding proteins. BMC Struct Biol 5:19. https://doi.org/10.1186/1472-76807-5-19
Kumar S, Stecher G, Li M, Knyaz C, Tamura K, Battistuzzi FU (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lam KC, Ibrahim RK, Behdad B, Dayanadan S (2007) Structure, function, and evolution of O-methyltrasferases. Genome 50:1001–1013. https://doi.org/10.1139/G07-077
Lankau T, Ken HC, Chang HS, Yu CH (2022) A computational study of promiscuity of SAM-dependent methyltransferase AtGTMT1. ACS Oemga 7(15):12753–12764. https://doi.org/10.1021/acsomega.1c07327
Liscombe DK, Macleod BP, Lpukanina N, Nandi OL, Fachinii PJ (2005) Evidence for the monophyletic evolution of benzylisoquinoline alkaloid biosynthesis in angiosperm. Phytochemistry 66:1374–1393. https://doi.org/10.1016/j.phytochem.2005.04029
Liscombe DK, Lousie GV, Noel JP (2012) Architecture, mechanisms and molecular evolution of natural product methyltransferases. Nat Prod Rep 29:1238–1250. https://doi.org/10.1039/c2np20029e
Menéndez-Perdomo IM, Facchini PJ (2018) Benzylisoquinoline alkaloids biosynthesis in sacred lotus. Molecules 23:2899. https://doi.org/10.3390/molecules23112899
Menéndez-Perdomo LM, Facchini PJ (2020) Isolation and characterization of two O-methyltransferases involved in benzylisoquinoline alkaloid biosynthesis in sacred lotus (Nelumbo nucifera). J Biol Chem 295:1598–1612. https://doi.org/10.1074/jbc.RA119.011547
Park N-I, Roy NS, Park Y, Choi B-S, Jeon MJ, Oh JY, Kim B-Y, Kim Y-D et al (2023) Isolation and characterization of the genes Involved in the berberine synthesis pathway in Asian blue cohosh. Caulophyllum Robustum Plants 12:1483. https://doi.org/10.3390/plants12071483
Petrovska BB (2012) Historical review of medicinal plants’ usage. Pharmacogn Rev 6(11):1–5. https://doi.org/10.4103/0973-7847.95849
Roy NS, Choi IY, Um TY, Jeon MJ, Kim BY, Kim YD, Yu JK, Kim S (2021) Kim NS (2021) Gene expression and isoform identification of PacBio full length cDNA sequences for Berberine biosynthesis in Berberis Koreana. Plants 10:1314. https://doi.org/10.3390/plants1007131
Roy NS, Park NI, Kim NS, Park Y, Kim BY, Kim YD, Yu JK, Kim YI, Um T, Kim S, Choi IK (2022) Comparative transcriptomics for genes related to berberine and berbamine biosynthesis in Berberidaceae. Plants 11:2676. https://doi.org/10.3390/plants11202676
Rubio-Pina J, Vazquez-Flota F (2013) Pharmaceutical applications of the benzylisoquinoline alkaloids from Argemone mxicana L. Curr Top Med Chem 13:2200–2217. https://doi.org/10.2174/15680266113139990152
Waddell TG, Eilders LL, Patel BP, Sims M (2000) Prebiotic methylation and the evolution of methyl transfer reactions in living cells. Org Life Evol Biosph 30:539–548. https://doi.org/10.1023/a:1026523222285
Weid M, Ziegler J, Kutchan T (2004) The role of latex and the vascular bundles in morphine biosynthesis in the opium poppy Papaver somniferum. Proc Nat Amer Sci USA 101:13957–13962. https://doi.org/10.1073/pnas.0405704101
Wlodarski T, Kutner J, Towpik J, Knizewski L, Rychlewski L, Kudlicki A, Rowicka M, Dziembowski A, Ginalski K (2011) Comprehensive structural and substrate specificity classification of the Saccharomyces cereviceae methyltransferome. PLoS ONE 68:e23168. https://doi.org/10.1371/journal.pone.0023168
Wolf YI, Brenner SE, Bash PA, Koonin EV (1999) Distribution pf protein folds in the three suprekingdoms of life. Genome Res 9:17–26. https://doi.org/10.1101/gr.9.1.17
Wong JM, Eirin-Lopez JM (2021) Evolution of methylatransferase-like (METTL) proteins in metazoan: A complex gene family involved in epitranscrioptomic regulation and other epigenetic processes. Mol Biol Evol 38:5309–5327. https://doi.org/10.1093/molbev/msab267
Zhong F, Huang L, Qi L, Ma Y, Yan Z (2020) Full-length transcriptome analysis of Coptis deltoids and identification of putative genes involved in benzylisoquinoline alkaloids biosynthesis based on combined sequencing platforms. Plant Mol Bio 102:477–499. https://doi.org/10.1007/s11103-019-00959-y
Acknowledgements
This work was supported by the National Institute of Biological Resources (NIBR202322101).
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
I.-Y.C. and S.L. conceived and designed the project and edited the manuscript. B.-S.C. and N.-S.K. contributed to the data analysis and drafted the manuscript. N.-I.P., Y.P., K.-C.P., E.S.K. and Y.K.S. prepared the sample materials and conducted an experiment of substance content. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare that they have no competing of interests.
Ethical approval
Not applicable.
Consent statement
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, S., Park, NI., Park, Y. et al. O- and N-Methyltransferases in benzylisoquinoline alkaloid producing plants. Genes Genom 46, 367–378 (2024). https://doi.org/10.1007/s13258-023-01477-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01477-4