Abstract
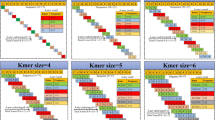
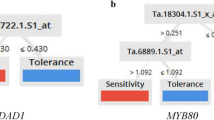
Abiotic stresses are major limiting factors for maize growth. Therefore, exploration of the mechanisms underlying the response to abiotic stress in maize is of great interest. Toward this end, we performed integration of the feature selection method into the meta-analysis of microarray gene expression. Following extraction of raw data, normalization, and batch effect removal, the data were merged into one expression profile. Differentially expressed genes (DEGs) between control and abiotic conditions were used for the feature selection algorithm to find the minimum features for high-performance classification. Feature selection was performed using a correlation-based feature selection (CFS) algorithm, considering features with a coefficient of 0.7 to 1. Different algorithms of Bayes, Functions, Lazy, Meta, Rules, and Trees were then tested in order to classify the samples and find the best performance classifier in each group. Moreover, the biological pathways and promoter motif analysis of selected genes were identified. The superior and overall performance of classification using all features (DEGs) were 98.86% (Multilayer Perceptron) and 81.25%, respectively. Classification based on feature selection resulted in an average accuracy of 94.69% and 93.56% with 33 and 12 features, respectively. Subsequently, gene ontology and promoter analysis were performed for the 12 selected biomarker genes. Five of them were downregulated and 7 were upregulated. ABRE, unnamed-1, G-box, and G-Box are motifs related to genes involved in several abiotic stress responses and are located upstream of at least nine probes in our study. This study revealed key genes associated with tolerance to abiotic stress in maize.
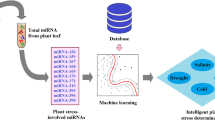
Graphical abstract






Similar content being viewed by others
References
Ashrafi-Dehkordi E, Alemzadeh A, Tanaka N, et al. 2018 Meta-analysis of transcriptomic responses to biotic and abiotic stress in tomato. PeerJ 6 e4631
Bai J, Mao J, Yang H, et al. 2017 Sucrose non-ferment 1 related protein kinase 2 (SnRK2) genes could mediate the stress responses in potato (Solanum tuberosum L.). BMC Genet. 18 41
Buske FA, Bodén M, Bauer DC, et al. 2010 Assigning roles to DNA regulatory motifs using comparative genomics. Bioinformatics 26 860–866
Carranza-Rojas J, Goeau H, Bonnet P, et al. 2017 Going deeper in the automated identification of Herbarium specimens. BMC Evol. Bio. 17 181
Durand M, Porcheron B, Hennion N, et al. 2016 Water deficit enhances C export to the roots in Arabidopsis thaliana plants with contribution of sucrose transporters in both shoot and roots. Plant Physiol. 170 1460–1479
Gautier L, Cope L, Bolstad B, et al. 2004 affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20 307–315
Godfray HCJ, Beddington JR, Crute IR, et al. 2010 Food security: the challenge of feeding 9 billion people. Science 327 812–818
Huang C-K, Shen Y-L, Huang L-F, et al. 2016 The DEAD-box RNA helicase AtRH7/PRH75 participates in pre-rRNA processing, plant development and cold tolerance in Arabidopsis. Plant Cell Physiol. 57 174–191
Irizarry RA, Bolstad BM, Collin F, et al. 2003 Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31 e15
Jain M and Khurana JP 2009 Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J. 276 3148–3162
Ji J, Yang L, Fang Z, et al. 2022 Plant SWEET family of sugar transporters: structure, evolution and biological functions. Biomolecules 12 205
Ji J, Zheng L, Yue J, et al. 2017 Identification of two CiGADs from Caragana intermedia and their transcriptional responses to abiotic stresses and exogenous abscisic acid. PeerJ 5 e3439
Kulik A, Wawer I, Krzywińska E, et al. 2011 SnRK2 protein kinases—key regulators of plant response to abiotic stresses. Omics 15 859–872
Leek JT, Johnson WE, Parker HS, et al. 2012 The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28 882–883
Lescot M, Déhais P, Thijs G, et al. 2002 PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30 325–327
Mahalingam R 2015 Consideration of combined stress: a crucial paradigm for improving multiple stress tolerance in plants; in Combined stresses in plants (Springer) pp 1–25
Mallikarjuna MG, Thirunavukkarasu N, Sharma R, et al. 2020 Comparative transcriptome analysis of iron and zinc deficiency in maize (Zea mays L.). Plants 9 1812
Mao X, Zhang H, Tian S, et al. 2010 TaSnRK2. 4, an SNF1-type serine/threonine protein kinase of wheat (Triticum aestivum L.), confers enhanced multistress tolerance in Arabidopsis. J. Exp. Bot. 61 683–696
Mattiello L, Begcy K, Da Silva FR, et al. 2014 Transcriptome analysis highlights changes in the leaves of maize plants cultivated in acidic soil containing toxic levels of Al3+. Mol. Biol. Rep. 41 8107–8116
Mattiello L, Kirst M, da Silva FR, et al. 2010 Transcriptional profile of maize roots under acid soil growth. BMC Plant Biol. 10 196
Mochida K, Koda S, Inoue K, et al. 2018 Statistical and machine learning approaches to predict gene regulatory networks from transcriptome datasets. Front. Plant Sci. 9 1770
Nakamura T, Muramoto Y, Yokota S, et al. 2004 Structural and transcriptional characterization of a salt-responsive gene encoding putative ATP-dependent RNA helicase in barley. Plant Sci. 167 63–70
Nawaz G and Kang H 2017 Chloroplast- or mitochondria-targeted DEAD-box RNA helicases play essential roles in organellar RNA metabolism and abiotic stress responses. Front. Plant Sci. 8 871
Nguyen LV, Seok H-Y, Woo D-H, et al. 2018 Overexpression of the DEAD-box RNA helicase gene AtRH17 confers tolerance to salt stress in Arabidopsis. Int. J. Mol. Sci. 19 3777
Phukan UJ, Mishra S, Timbre K, et al. 2014 Mentha arvensis exhibit better adaptive characters in contrast to Mentha piperita when subjugated to sustained waterlogging stress. Protoplasma 251 603–614
Pillemer D and Light R 1980 Synthesizing outcomes: How to use research evidence from many studies. Harvard Edu. Rev. 50 176–195
Ramasamy A, Mondry A, Holmes CC, et al. 2008 Key issues in conducting a meta-analysis of gene expression microarray datasets. PLoS Med. 5 e184
Rung J and Brazma A 2013 Reuse of public genome-wide gene expression data. Nat. Rev. Genet. 14 89–99
Sabanci K, Ünlerşen MF and Polat K 2016 Classification of different forest types with machine learning algorithms. Res. Rural Dev. 1 254–260
Sharma R, De Vleesschauwer D, Sharma MK, et al. 2013 Recent advances in dissecting stress-regulatory crosstalk in rice. Mol. Plant 6 250–260
Shen P-C, A-L Hour and L-YD Liu 2017 Microarray meta-analysis to explore abiotic stress-specific gene expression patterns in Arabidopsis. Bot. Stud. 58 22
Shen Q, Lin Y, Li Y, et al. 2021 Dynamics of H3K27me3 modification on plant adaptation to environmental cues. Plants 10 1165
Singh A, Ganapathysubramanian B, Singh AK, et al. 2016 Machine learning for high-throughput stress phenotyping in plants. Trends Plant Sci. 21 110–124
Tahmasebi A, Ashrafi-Dehkordi E, Shahriari AG, et al. 2019 Integrative meta-analysis of transcriptomic responses to abiotic stress in cotton. Prog. Biophys. Mol. Biol. 146 112–122
Tahmasebi A and Niazi A 2021 Comparison of transcriptional response of C3 and C4 plants to drought stress using meta-analysis and systems biology approach. Front. in Plant Sci. 1295
Thirunavukkarasu N, Hossain F, Mohan S, et al. 2013 Genome-wide expression of transcriptomes and their co-expression pattern in subtropical maize (Zea mays L.) under waterlogging stress. PLoS One 8 e70433
Tuteja N, Sahoo RK, Garg B, et al. 2013 Os SUV 3 dual helicase functions in salinity stress tolerance by maintaining photosynthesis and antioxidant machinery in rice (Oryza sativa L. cv. IR 64). Plant J. 76 115–127
van Ijzendoorn DG, Szuhai K, Briaire-de Bruijn IH, et al. 2019 Machine learning analysis of gene expression data reveals novel diagnostic and prognostic biomarkers and identifies therapeutic targets for soft tissue sarcomas. PLoS Comput. Biol. 15 e1006826
Wang L, Yao L, Hao X, et al. 2018a Tea plant SWEET transporters: expression profiling, sugar transport, and the involvement of CsSWEET16 in modifying cold tolerance in Arabidopsis. Plant Mol. Biol. 96 577–592
Wang M, Xu Z and Kong Y 2018b The tubby-like proteins kingdom in animals and plants. Gene 642 16–25
Wang Y, Li T, Sun Z, et al. 2022 Comparative transcriptome meta-analysis reveals a set of genes involved in the responses to multiple pathogens in maize. Front. Plant Sci. 13 971371
Xu C and Jackson SA 2019 Machine learning and complex biological data (Springer)
Yamada K, Osakabe Y, Mizoi J, et al. 2010 Functional analysis of an Arabidopsis thaliana abiotic stress-inducible facilitated diffusion transporter for monosaccharides. J. Biol. Chem. 285 1138–1146
Yu J, Schumann AW, Cao Z, et al. 2019 Weed detection in perennial ryegrass with deep learning convolutional neural network. Front. Plant Sci. 10 1422
Zhang J and Li S 2017 A review of machine learning based species' distribution modelling; in 2017 International Conference on Industrial Informatics-Computing Technology, Intelligent Technology, Industrial Information Integration (ICIICII) (IEEE) pp 199–206
Zhang W, Wang S, Yu F, et al. 2019 Genome-wide characterization and expression profiling of SWEET genes in cabbage (Brassica oleracea var. capitata L.) reveal their roles in chilling and clubroot disease responses. BMC Genomics 20 93
Zheng J, Fu J, Gou M, et al. 2010 Genome-wide transcriptome analysis of two maize inbred lines under drought stress. Plant Mol. Biol. 72 407–421
Zhou A, Ma H, Feng S, et al. 2018 A novel sugar transporter from Dianthus spiculifolius, DsSWEET12, affects sugar metabolism and confers osmotic and oxidative stress tolerance in Arabidopsis. Int. J. Mol. Sci. 19 497
Zinati Z, Sazegari S, Tahmasebi A, et al. 2021 A comprehensive meta-analysis to identify transcriptional signatures of abiotic stress responses in barley (Hordeum vulgare). Cereal Res. Commun. 49 385–391
Zuluaga AP, Bidzinski P, Chanclud E, et al. 2020 The rice DNA-binding protein ZBED controls stress regulators and maintains disease resistance after a mild drought. Front. Plant Sci. 11 1265
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
LN and ZZ contributed to the conception and design of study, performing machine learning and bioinformatics analysis, and writing the submitted manuscript. PB contributed to the conception of study, writing, and editing of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Corresponding editor: Jyothilakshmi Vadaserry
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nazari, L., Zinati, Z. & Bagnaresi, P. Identification of biomarker genes from multiple studies for abiotic stress in maize through machine learning. J Biosci 49, 1 (2024). https://doi.org/10.1007/s12038-023-00392-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-023-00392-w