Abstract
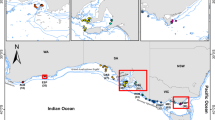
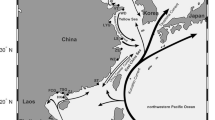
Understanding the genetic composition and regional adaptation of marine species under environmental heterogeneity and fishing pressure is crucial for responsible management. In order to understand the genetic diversity and adaptability of yellowfin seabream (Acanthopagrus latus) along southern China coast, this study was conducted a seascape genome analysis on yellowfin seabream from the ecologically diverse coast, spanning over 1600 km. A total of 92 yellowfin seabream individuals from 15 sites were performed whole-genome resequencing, and 4,383,564 high-quality single nucleotide polymorphisms (SNPs) were called. By conducting a genotype-environment association analysis, 29,951 adaptive and 4,328,299 neutral SNPs were identified. The yellowfin seabream exhibited two distinct population structures, despite high gene flow between sites. The seascape genome analysis revealed that genetic structure was influenced by a variety of factors including salinity gradients, habitat distance, and ocean currents. The frequency of allelic variation at the candidate loci changed with the salinity gradient. Annotation of these loci revealed that most of the genes are associated with osmoregulation, such as kcnab2a, kcnk5a, and slc47a1. These genes are significantly enriched in pathways associated with ion transport including G protein-coupled receptor activity, transmembrane signaling receptor activity, and transporter activity. Overall, our findings provide insights into how seascape heterogeneity affects adaptive evolution, while providing important information for regional management in yellowfin seabream populations.






Similar content being viewed by others
Data Availability
The DNA resequencing data from this study was deposited into NCBI Sequence Read Archive (SRA) under BioProject ID PRJNA996779 and BioSample IDs SAMN36636882-SAMN36636973.
References
Ackiss AS, Bird CE, Akita Y et al (2018) Genetic patterns in peripheral marine populations of the fusilier fish Caesio cuning within the Kuroshio Current. Ecol Evol 8:11875–11886
Alberts B, Johnson A, Lewis J et al (2002) Molecular Biology of the Cell vol 4th edition. Garland Science, New York. https://doi.org/10.1002/bmb.2003.494031049999
Alex Hesp S, Potter IC, Hall NG (2004) Reproductive biology and protandrous hermaphroditism in Acanthopagrus latus. Environ Biol Fishes 70:257–272
Alexander DH, Novembre J, Lange K (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Res 19:1655–1664
Antoniou A, Manousaki T, Ramírez F et al (2023) Sardines at a junction: seascape genomics reveals ecological and oceanographic drivers of variation in the NW Mediterranean Sea. Mol Ecol 32:1608–1628
Ashburner M, Ball CA, Blake JA et al (2000) Gene Ontology: tool for the unification of biology. Nat Genet 25:25–29
Assis J, Tyberghein L, Bosch S et al (2018) Bio-ORACLE v2.0: Extending marine data layers for bioclimatic modelling. Glob Ecol Biogeogr 27:277–284
Beaumont MA, Nichols RA (1996) Evaluating loci for use in the genetic analysis of population structure. Proc Biol Sci 263:1619–1626
Behrens KA, Girasek QL, Sickler A et al (2021) Regions of genetic divergence in depth-separated Sebastes rockfish species pairs: depth as a potential driver of speciation. Mol Ecol 30:4259–4275
Benestan L, Fietz K, Loiseau N et al (2021) Restricted dispersal in a sea of gene flow. Proc R Soc B Biol Sci 288:20210458
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Stat Soc: Ser B (methodol) 57:289–300
Bernal-Ramírez JH, Adcock GJ, Hauser L et al (2003) Temporal stability of genetic population structure in the New Zealand snapper, Pagrus auratus, and relationship to coastal currents. Mar Biol 142:567–574
Bonhomme M, Chevalet C, Servin B et al (2010) Detecting selection in population trees: the Lewontin and Krakauer Test Extended. Genetics 186:241–262
Bowen BW, Gaither MR, DiBattista JD et al (2016) Comparative phylogeography of the ocean planet. Proc Natl Acad Sci 113:7962–7969
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60
Capblancq T, Forester BR (2021) Redundancy analysis: a Swiss Army Knife for landscape genomics. Methods Ecol Evol 12:2298–2309
Capblancq T, Luu K, Blum MGB, Bazin E (2018) Evaluation of redundancy analysis to identify signatures of local adaptation. Mol Ecol Resour 18:1223–1233
Carroll SP, Jørgensen PS, Kinnison MT et al (2014) Applying evolutionary biology to address global challenges. Science 346:1245993
Caye K, Jumentier B, Lepeule J, François O (2019) LFMM 2: fast and accurate inference of gene-environment associations in genome-wide studies. Mol Biol Evol 36:852–860
Chen B, Bai Y, Wang J et al (2023a) Population structure and genome-wide evolutionary signatures reveal putative climate-driven habitat change and local adaptation in the large yellow croaker. Mar Life Sci Technol. https://doi.org/10.1007/s42995-023-00165-2
Chen B, Zhou Z, Shi Y et al (2023b) Genome-wide evolutionary signatures of climate adaptation in spotted sea bass inhabiting different latitudinal regions. Evol Appl 16:1029–1043
Chen C, Chen H, Zhang Y et al (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13:1194–1202
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Cingolani P, Platts A, Wang LL et al (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (austin) 6:80–92
Cnaani A, Grau G, Lerner D, Harpaz S (2014) Gastrointestinal osmoregulatory activity in Tilapia and its effects on growth, an opportunity for fish diet developments. https://doi.org/10.32747/2014.7594393.bard
Czesny S, Epifanio J, Michalak P (2012) Genetic divergence between freshwater and marine morphs of alewife (Alosa pseudoharengus): a ‘next-generation’ sequencing analysis. PLoS ONE 7:e31803
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158
Delerue-Ricard S, Darnaude AM, Raeymaekers JAM et al (2019) Extensive larval dispersal and restricted movement of juveniles on the nursery grounds of sole in the Southern North Sea. J Sea Res 155:101822
Diallinas G (2014) Understanding transporter specificity and the discrete appearance of channel-like gating domains in transporters. Front Pharmacol 5. https://doi.org/10.3389/fphar.2014.00207
Dormann CF, Elith J, Bacher S et al (2013) Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography 36:27–46
Drinan DP, Gruenthal KM, Canino MF et al (2018) Population assignment and local adaptation along an isolation-by-distance gradient in Pacific cod (Gadus macrocephalus). Evol Appl 11:1448–1464
Excoffier L, Hofer T, Foll M (2009) Detecting loci under selection in a hierarchically structured population. Heredity 103:285–298
Fariello MI, Boitard S, Naya H et al (2013) Detecting signatures of selection through haplotype differentiation among hierarchically structured populations. Genetics 193:929–941
Fitzpatrick MC, Chhatre VE, Soolanayakanahally RY, Keller SR (2021) Experimental support for genomic prediction of climate maladaptation using the machine learning approach Gradient Forests. Mol Ecol Resour 21:2749–2765
Foll M, Gaggiotti O (2008) A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180:977–993
Forester BR, Lasky JR, Wagner HH, Urban DL (2018) Comparing methods for detecting multilocus adaptation with multivariate genotype–environment associations. Mol Ecol 27:2215–2233
Galpern P, Peres-Neto PR, Polfus J, Manseau M (2014) MEMGENE: Spatial pattern detection in genetic distance data. Methods Ecol Evol 5:1116–1120
Grummer JA, Beheregaray LB, Bernatchez L et al (2019) Aquatic landscape genomics and environmental effects on genetic variation. Trends Ecol Evol 34:641–654
Hauser L, Carvalho GR (2008) Paradigm shifts in marine fisheries genetics: ugly hypotheses slain by beautiful facts. Fish Fish 9:333–362
Heino M, Díaz Pauli B, Dieckmann U (2015) Fisheries-induced evolution. Annu Rev Ecol Evol Syst 46:461–480
Hoey JA, Fodrie FJ, Walker QA et al (2020) Using multiple natural tags provides evidence for extensive larval dispersal across space and through time in summer flounder. Mol Ecol 29:1421–1435
Hoffmann AA, Willi Y (2008) Detecting genetic responses to environmental change. Nat Rev Genet 9:421–432
Iwatsuki Y (2013) Review of the Acanthopagrus latus complex (Perciformes: Sparidae) with descriptions of three new species from the Indo-West Pacific Ocean. J Fish Biol 83:64–95
Jiang DL, Gu XH, Li BJ et al (2019) Identifying a long QTL cluster across chrLG18 associated with salt tolerance in tilapia using GWAS and QTL-seq. Mar Biotechnol 21:250–261
Johannesson K, Leder EH, André C et al (2023) Ten years of marine evolutionary biology—Challenges and achievements of a multidisciplinary research initiative. Evol Appl 16:530–541
Junker J, Rick JA, McIntyre PB et al (2020) Structural genomic variation leads to genetic differentiation in Lake Tanganyika’s sardines. Mol Ecol 29:3277–3298
Koudrina N, Perry SF, Gilmour KM (2020) The role of TASK-2 channels in CO2 sensing in zebrafish (Danio rerio). Am J Physiol Regul Integr Comp Physiol 319:R329–R342
Kozak GM, Brennan RS, Berdan EL et al (2014) Functional and population genomic divergence within and between two species of killifish adapted to different osmotic niches. Evolution 68:63–80
Lasky JR, Des Marais DL, McKAY JK et al (2012) Characterizing genomic variation of Arabidopsis thaliana: the roles of geography and climate. Mol Ecol 21:5512–5529
Legendre P, Oksanen J, ter Braak CJF (2010) Testing the significance of canonical axes in redundancy analysis. Methods Ecol Evol 2:269–277
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics (oxford, England) 25:1754–1760
Li P, Liu H, Sun J et al (2023a) HiBrowser: an interactive and dynamic browser for synchronous Hi-C data visualization. Brief Bioinform bbad283. https://doi.org/10.1093/bib/bbad283
Li S, Lin G, Fang W et al (2020) Gonadal transcriptome analysis of sex-related genes in the protandrous yellowfin seabream (Acanthopagrus latus). Front Genet 11:709
Li X, Jiang S, Zheng H et al (2023b) Organophosphorus pesticides in southeastern China marginal seas: Land-based export and ocean currents redistribution. Sci Total Environ 858:160011
Lin G, Zheng M, Li S et al (2020) Response of gut microbiota and immune function to hypoosmotic stress in the yellowfin seabream (Acanthopagrus latus). Sci Total Environ 745:140976
Liu N, Wang Y, Li T, Feng X (2021) G-protein coupled receptors (GPCRs): signaling pathways, characterization, and functions in insect physiology and toxicology. Int J Mol Sci 22:5260
Lodish H, Berk A, Zipursky S et al (2000) Molecular cell biology. W.H. Freeman, New York
Lu J, Gao D, Sims Y et al (2022) Chromosome-level genome assembly of Acanthopagrus latus provides insights into salinity stress adaptation of Sparidae. Mar Biotechnol 24:655–660
Lu J, Huang P, Sun J, Liu J (2023) DupScan: predicting and visualizing vertebrate genome duplication database. Nucleic Acids Res 51:D906–D912
Lv J, Sun D, Yan D, et al (2019) Quantitative trait loci mapping and marker identification for low salinity tolerance trait in the swimming crab (Portunus trituberculatus). Front Genet 10. https://doi.org/10.3389/fgene.2019.01193
Marcel J (1948) G. Malecot. — Les Mathématiques de l’Hérédité. 66 p., 9 fig., Masson, Paris 1948. Publications De La Société Linnéenne De Lyon 17:203–203
Mattingsdal M, Jorde PE, Knutsen H et al (2020) Demographic history has shaped the strongly differentiated corkwing wrasse populations in Northern Europe. Mol Ecol 29:160–171
Mayer L, Jakobsson M, Allen G et al (2018) The Nippon Foundation-GEBCO Seabed 2030 Project: The Quest to See the World's Oceans Completely Mapped by 2030. Geosciences 8(2). https://doi.org/10.3390/geosciences8020063
McKenna A, Hanna M, Banks E et al (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303
Miraldo A, Li S, Borregaard MK et al (2016) An Anthropocene map of genetic diversity. Science 353:1532–1535
Naaykens T, D’Aloia CC (2022) Isolation-by-distance and genetic parentage analysis provide similar larval dispersal estimates. Mol Ecol 31:3072–3082
Norriss J, Jackson G (2007) A substantial recruitment year for the western yellowfin bream (Acanthopagrus latus, Sparidae) sustains years of high catch rates in the inner gulfs of Shark Bay, Western Australia. J R Soc West Aust 90:157–159. https://biostor.org/reference/256220
Oksanen J (2011) Vegan: Community Ecology Package. Available: https://CranR-Project.org/Web/Packages/Vegan/IndexHtml
Pan X, Ye Z, Xu B et al (2020) Population connectivity in a highly migratory fish, Japanese Spanish mackerel (Scomberomorus niphonius), along the Chinese coast, implications from otolith chemistry. Fish Res 231:105690
Petkova D, Novembre J, Stephens M (2016) Visualizing spatial population structure with estimated effective migration surfaces. Nat Genet 48:94–100
Pickrell JK, Pritchard JK (2012) Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 8:e1002967
Poloczanska ES, Brown CJ, Sydeman WJ et al (2013) Global imprint of climate change on marine life. Nature Clim Change 3:919–925
Pratt EAL, Beheregaray LB, Bilgmann K et al (2022) Seascape genomics of coastal bottlenose dolphins along strong gradients of temperature and salinity. Mol Ecol 31:2223–2241
Price N, Lopez L, Platts AE, Lasky JR (2020) In the presence of population structure: from genomics to candidate genes underlying local adaptation. Ecol Evol 10:1889–1904
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Qasemi A, Shadi A (2018) Population structure of Acanthopagrus latus from the northern Persian Gulf and Gulf of Oman based on microsatellite markers. Turk J Fish Aquat Sci 18:983–990
Revelle W (2023) Psych: procedures for psychological, psychometric, and personality research. Northwestern University, Evanston, Illinois. R package version 2.3.3. https://CRAN.R-project.org/package=psych
Rousset F (1997) Genetic differentiation and estimation of gene flow from F-statistics under isolation by distance. Genetics 145:1219–1228
Selmoni O, Lecellier G, Magalon H et al (2021) Seascape genomics reveals candidate molecular targets of heat stress adaptation in three coral species. Mol Ecol 30:1892–1906
Sork VL (2018) Genomic studies of local adaptation in natural plant populations. J Hered 109:3–15
Sun C-H, Gozlan RE, Wu T et al (2022) The role of ancestral seascape discontinuity and geographical distance in structuring rockfish populations in the Pacific Northwest. Front Mar Sci 9. https://doi.org/10.3389/fmars.2022.1018864
Tang G, He Z, Liu Y, et al (2023) Acanthopagrus latus migration patterns and habitat use in Wanshan Islands, Pearl River Estuary, determined using otolith microchemical analysis. Front Mar Sci 10. https://doi.org/10.3389/fmars.2023.1104528
Teske PR, Sandoval-Castillo J, Golla TR et al (2019) Thermal selection as a driver of marine ecological speciation. Proc R Soc B Biol Sci 286:20182023
Tiffin P, Ross-Ibarra J (2014) Advances and limits of using population genetics to understand local adaptation. Trends Ecol Evol 29:673–680
Tong C, Li M, Tang Y, Zhao K (2021) Genomic signature of shifts in selection and alkaline adaptation in highland fish. Genome Biol Evol 13:evab086
Tyberghein L, Verbruggen H, Pauly K et al (2012) Bio-ORACLE: a global environmental dataset for marine species distribution modelling. Glob Ecol Biogeogr 21:272–281
Van der Auwera GA, Carneiro MO, Hartl C, et al (2013) From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 43:11.10.1–11.10.33. https://doi.org/10.1002/0471250953.bi1110s43
van Oppen MJH, Coleman MA (2022) Advancing the protection of marine life through genomics. PLoS Biol 20:e3001801
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–935
Wang W, Dong H, Sun Y et al (2020) Immune and physiological responses of juvenile Chinese sea bass (Lateolabrax maculatus) to eugenol and tricaine methanesulfonate (MS-222) in gills. Aquac Rep 18
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wickham H (2016) ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag, New York. https://doi.org/10.1007/978-3-319-24277-4
Willi Y, Van Buskirk J, Schmid B, Fischer M (2007) Genetic isolation of fragmented populations is exacerbated by drift and selection. J Evol Biol 20:534–542
Wright S (1943) Isolation by distance. Genetics 28:114–138
Xia J-H, Huang J-H, Gong J-B, Jiang S-G (2008) Significant population genetic structure of yellowfin seabream Acanthopagrus latus in China. J Fish Biol 73:1979–1992
Xuan Z, Wang W-X (2023) Diversity of life history and population connectivity of threadfin fish Eleutheronema tetradactylum along the coastal waters of Southern China. Sci Rep 13:3976
Zhang P, Dai P, Zhang J et al (2021) Spatiotemporal variation, speciation, and transport flux of TDP in Leizhou Peninsula coastal waters, South China Sea. Mar Pollut Bull 167
Zhang Y, Wen H, Liu Y et al (2023) Gill histological and transcriptomic analysis provides insights into the response of spotted sea bass (Lateolabrax maculatus) to alkalinity stress. Aquaculture 563:738945
Zheng X, Asico LD, Ma X, Konkalmatt PR (2019) G protein-coupled receptor 37L1 regulates renal sodium transport and blood pressure. Am J Physiol Renal Physiol 316:F506–F516
Zheng Q, Fang G, Song YT (2006) Introduction to special section: dynamics and circulation of the Yellow, East, and South China Seas. J Geophys Res Oceans 111. https://doi.org/10.1029/2005JC003261
Acknowledgements
The authors would like to thank all laboratory members for their assistance during the experiments. We are immensely thankful to Zhentong Xu, Hezhao Yang, Junnan Lan, Gongyou Wang, Zhiqiang Chen, Fengliang Chen, Houmin chen, Sheng Long, Weibang Liang, Haiyan Su, Shigang Mo, Xiaoli Wu, and Fei An, for their irreplaceable assistance in obtaining the samples.
Funding
This study was supported by R&D Project for Jinwan Yellowfin Seabream Breeding System Construction (No. K20-42000-018), project supported by Innovation Group Project of Southern Marine Science and Engineering Guangdong Laboratory (Zhuhai) (No. 311021006), and the National Natural Science Foundation of China (No. 31902427).
Author information
Authors and Affiliations
Contributions
Wenhao Wang conceived and executed the study and wrote the paper. Junrou Huang called the SNP and performed gene flow and seascape genomic analyses. Yan Hu performed the population genetic analyses and carried out bioinformatic analysis. Jianxiang Feng provided expertise and commented on the paper. Dong Gao performed functional annotation and enrichment analyses. Wenyu Fang collected the samples and enrichment analyses. Meng Xu collected the samples and extracted DNA. Chunlei Ma produced environmental maps. Zhenqiang Fu provided expertise and bioinformatic analysis. Qinglong Chen provided bioinformatic analysis and helped for drawing. Xuanguang Liang provided suggestions and helped for drawing. Jianguo Lu conceived and supervised the study and wrote the paper. All authors approved the final version of the paper.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
All experimental procedures in our study with yellowfin seabream were approved by the Animal Care and Use Committee of the School of Marine Sciences, Sun Yat-sen University.
Competing Interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, W., Huang, J., Hu, Y. et al. Seascapes Shaped the Local Adaptation and Population Structure of South China Coast Yellowfin Seabream (Acanthopagrus latus). Mar Biotechnol 26, 60–73 (2024). https://doi.org/10.1007/s10126-023-10277-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-023-10277-6