Abstract
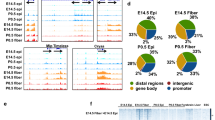
Differentiation of lens fiber cells involves a complex interplay of signals from growth factors together with tightly regulated gene expression via transcriptional and post-transcriptional regulators. Various studies have demonstrated that RNA-binding proteins, functioning in ribonucleoprotein granules, have important roles in regulating post-transcriptional expression during lens development. In this study, we examined the expression and localization of two members of the BTG/TOB family of RNA-binding proteins, TOB1 and TOB2, in the developing lens and examined the phenotype of mice that lack Tob1. By RT-PCR, both Tob1 and Tob2 mRNA were detected in epithelial and fiber cells of embryonic and postnatal murine lenses. In situ hybridization showed Tob1 and Tob2 mRNA were most intensely expressed in the early differentiating fibers, with weaker expression in anterior epithelial cells, and both appeared to be downregulated in the germinative zone of E15.5 lenses. TOB1 protein was detected from E11.5 to E16.5 and was predominantly detected in large cytoplasmic puncta in early differentiating fiber cells, often co-localizing with the P-body marker, DCP2. Occasional nuclear puncta were also observed. By contrast, TOB2 was detected in a series of interconnected peri-nuclear granules, in later differentiating fiber cells of the inner cortex. TOB2 did not appear to co-localize with DCP2 but did partially co-localize with an early stress granule marker (EIF3B). These data suggest that TOB1 and TOB2 are involved with different aspects of the mRNA processing cycle in lens fiber cells. In vitro experiments using rat lens epithelial explants treated with or without a fiber differentiating dose of FGF2 showed that both TOB1 and TOB2 were up-regulated during FGF-induced differentiation. In differentiating explants, TOB1 also co-localized with DCP2 in large cytoplasmic granules. Analyses of Tob1-/- mice revealed relatively normal lens morphology but a subtle defect in cell cycle arrest of some cells at the equator and in the lens fiber mass of E13.5 embryos. Overall, these findings suggest that TOB proteins play distinct regulatory roles in RNA processing during lens fiber differentiation.








Similar content being viewed by others
References
Anand D, Kakrana A, Siddam AD, Huang H, Saadi I, Lachke SA (2018) RNA sequencing-based transcriptomic profiles of embryonic lens development for cataract gene discovery. Hum Genet 137(11–12):941–954. https://doi.org/10.1007/s00439-018-1958-0
Anderson P, Kedersha N (2009) RNA granules: post-transcriptional and epigenetic modulators of gene expression. Nat Rev Mol Cell Biol 10(6):430–436. https://doi.org/10.1038/nrm2694
Basu S, Rajakaruna S, De Arcangelis A, Zhang L, Georges-Labouesse E, Menko AS (2014a) α6 integrin transactivates insulin-like growth factor receptor-1 (IGF-1R) to regulate caspase-3-mediated lens epithelial cell differentiation initiation. J Biol Chem 289(7):3842–3855. https://doi.org/10.1074/jbc.M113.515254
Basu S, Rajakaruna S, Reyes B, Van Bockstaele E, Menko AS (2014b) Suppression of MAPK/JNK-MTORC1 signaling leads to premature loss of organelles and nuclei by autophagy during terminal differentiation of lens fiber cells. Autophagy 10(7):1193–1211. https://doi.org/10.4161/auto.28768
Boswell BA, Musil LS (2015) Synergistic interaction between the fibroblast growth factor and bone morphogenetic protein signaling pathways in lens cells. Mol Biol Cell 26(13):2561–2572. https://doi.org/10.1091/mbc.E15-02-0117
Boswell BA, Overbeek PA, Musil LS (2008) Essential role of BMPs in FGF-induced secondary lens fiber differentiation. Dev Biol 324(2):202–212. https://doi.org/10.1016/j.ydbio.2008.09.003
Buchan JR (2014) mRNP granules: Assembly, function, and connections with Disease. RNA Biol 11:8
Cain S, Martinez G, Kokkinos MI, Turner K, Richardson RJ, Abud HE, Huelsken J, Robinson ML, de Iongh RU (2008) Differential requirement for beta-catenin in epithelial and fiber cells during lens development. Dev Biol 321(2):420–433
Cavalheiro GR, Matos-Rodrigues GE, Gomes AL, Rodrigues PM, Martins RA (2014) c-Myc regulates cell proliferation during lens development. PLoS ONE 9(2):e87182. https://doi.org/10.1371/journal.pone.0087182
Chen CA, Strouz K, Huang KL, Shyu AB (2020) Tob2 phosphorylation regulates global mRNA turnover to reshape transcriptome and impact cell proliferation. RNA (New York NY) 26(9):1143–1159. https://doi.org/10.1261/rna.073528.119
Choi JJ, Ting CT, Trogrlic L, Milevski SV, Familari M, Martinez G, de Iongh RU (2014) A role for smoothened during murine lens and cornea development. PLoS ONE 9(9):e108037. https://doi.org/10.1371/journal.pone.0108037
Collart MA, Panasenko OO (2012) The Ccr4–not complex. Gene 492(1):42–53. https://doi.org/10.1016/j.gene.2011.09.033
Collart MA, Panasenko OO, Nikolaev SI (2013) The Not3/5 subunit of the Ccr4-Not complex: a central regulator of gene expression that integrates signals between the cytoplasm and the nucleus in eukaryotic cells. Cell Signal 25(4):743–751. https://doi.org/10.1016/j.cellsig.2012.12.018
Cvekl A, Ashery-Padan R (2014) The cellular and molecular mechanisms of vertebrate lens development. Development 141(23):4432–4447. https://doi.org/10.1242/dev.107953
Cvekl A, Zhang X (2017) Signaling and Gene Regulatory Networks in mammalian Lens Development. Trends Genet 33(10):677–702. https://doi.org/10.1016/j.tig.2017.08.001
Cvekl A, McGreal R, Liu W (2015) Lens Development and crystallin gene expression. Prog Mol Biol Transl Sci 134:129–167. https://doi.org/10.1016/bs.pmbts.2015.05.001
D’Ambrogio A, Nagaoka K, Richter JD (2013) Translational control of cell growth and malignancy by the CPEBs. Nat Rev Cancer 13(4):283–290. https://doi.org/10.1038/nrc3485
Dash S, Dang CA, Beebe DC, Lachke SA (2015) Deficiency of the RNA binding protein caprin2 causes lens defects and features of Peters anomaly. Dev Dynamics: Official Publication Am Association Anatomists 244(10):1313–1327. https://doi.org/10.1002/dvdy.24303
de Iongh RU, Duncan MK (2014) Growth Factor Signaling in Lens Fiber Differentiation. In: Saika S, Werner L, Lovicu FJ (eds) Lens Epithelium and Posterior Capsular Opacification. Springer Japan, pp 81–104. https://doi.org/10.1007/978-4-431-54300-8_5
de Iongh RU, Lovicu FJ, Overbeek PA, Schneider MD, Joya J, Hardeman ED, McAvoy JW (2001) Requirement for TGFbeta receptor signaling during terminal lens fiber differentiation. Development 128(20):3995–4010
Decker CJ, Parker R (2012) P-bodies and stress granules: possible roles in the control of translation and mRNA degradation. Cold Spring Harb Perspect Biol 4(9):a012286. https://doi.org/10.1101/cshperspect.a012286
Doidge R, Mittal S, Aslam A, Winkler GS (2012a) The anti-proliferative activity of BTG/TOB proteins is mediated via the Caf1a (CNOT7) and Caf1b (CNOT8) deadenylase subunits of the Ccr4-not complex. PLoS ONE 7(12):e51331. https://doi.org/10.1371/journal.pone.0051331
Doidge R, Mittal S, Aslam A, Winkler GS (2012b) Deadenylation of cytoplasmic mRNA by the mammalian Ccr4-Not complex. Biochem Soc Trans 40(4):896–901. https://doi.org/10.1042/bst20120074
Ezzeddine N, Chang TC, Zhu W, Yamashita A, Chen CY, Zhong Z, Yamashita Y, Zheng D, Shyu AB (2007) Human TOB, an antiproliferative transcription factor, is a poly(A)-binding protein-dependent positive regulator of cytoplasmic mRNA deadenylation. Mol Cell Biol 27(22):7791–7801. https://doi.org/10.1128/mcb.01254-07
Ezzeddine N, Chen CY, Shyu AB (2012) Evidence providing new insights into TOB-promoted deadenylation and supporting a link between TOB’s deadenylation-enhancing and antiproliferative activities. Mol Cell Biol 32(6):1089–1098. https://doi.org/10.1128/mcb.06370-11
Faulkner-Jones B, Zandy AJ, Bassnett S (2003) RNA stability in terminally differentiating fibre cells of the ocular lens. Exp Eye Res 77(4):463–476
Hawse JR, DeAmicis-Tress C, Cowell TL, Kantorow M (2005) Identification of global gene expression differences between human lens epithelial and cortical fiber cells reveals specific genes and their associated pathways important for specialized lens cell functions. Mol Vis 11:274–283
Helms MW, Kemming D, Contag CH, Pospisil H, Bartkowiak K, Wang A, Chang SY, Buerger H, Brandt BH (2009) TOB1 is regulated by EGF-dependent HER2 and EGFR signaling, is highly phosphorylated, and indicates poor prognosis in node-negative Breast cancer. Cancer Res 69(12):5049–5056. https://doi.org/10.1158/0008-5472.can-08-4154
Ho KJ, Do NL, Otu HH, Dib MJ, Ren X, Enjyoji K, Robson SC, Terwilliger EF, Karp SJ (2010) Tob1 is a constitutively expressed repressor of liver regeneration. J Exp Med 207(6):1197–1208. https://doi.org/10.1084/jem.20092434
Hoang TV, Kumar PK, Sutharzan S, Tsonis PA, Liang C, Robinson ML (2014) Comparative transcriptome analysis of epithelial and fiber cells in newborn mouse lenses with RNA sequencing. Mol Vis 20:1491–1517
Hosoda N, Funakoshi Y, Hirasawa M, Yamagishi R, Asano Y, Miyagawa R, Ogami K, Tsujimoto M, Hoshino S (2011) Anti-proliferative protein Tob negatively regulates CPEB3 target by recruiting Caf1 deadenylase. EMBO J 30(7):1311–1323. https://doi.org/10.1038/emboj.2011.37
Huang KL, Chadee AB, Chen CY, Zhang Y, Shyu AB (2013) Phosphorylation at intrinsically disordered regions of PAM2 motif-containing proteins modulates their interactions with PABPC1 and influences mRNA fate. RNA (New York NY) 19(3):295–305. https://doi.org/10.1261/rna.037317.112
Jain S, Parker R (2013) The discovery and analysis of P bodies. Adv Exp Med Biol 768:23–43. https://doi.org/10.1007/978-1-4614-5107-5_3
Jeon P, Ham HJ, Park S, Lee JA (2022) Regulation of Cellular Ribonucleoprotein granules: from Assembly to Degradation via post-translational modification. Cells 11(13). https://doi.org/10.3390/cells11132063
Kakrana A, Yang A, Anand D, Djordjevic D, Ramachandruni D, Singh A, Huang H, Ho JWK, Lachke SA (2018) iSyTE 2.0: a database for expression-based gene discovery in the eye. Nucleic Acids Res 46(D1):D875–d885. https://doi.org/10.1093/nar/gkx837
Kallifatidis G, Boros J, Shin EH, McAvoy JW, Lovicu FJ (2011) The fate of dividing cells during lens morphogenesis, differentiation and growth. Exp Eye Res 92(6):502–511. https://doi.org/10.1016/j.exer.2011.03.012
Kawamura-Tsuzuku J, Suzuki T, Yoshida Y, Yamamoto T (2004) Nuclear localization of Tob is important for regulation of its antiproliferative activity. Oncogene 23(39):6630–6638. https://doi.org/10.1038/sj.onc.1207890
Kerr CL, Huang J, Williams T, West-Mays JA (2012) Activation of the hedgehog signaling pathway in the developing lens stimulates ectopic FoxE3 expression and disruption in fiber cell differentiation. Investig Ophthalmol Vis Sci 53(7):3316–3330. https://doi.org/10.1167/iovs.12-9595
Kundu J, Wahab SM, Kundu JK, Choi YL, Erkin OC, Lee HS, Park SG, Shin YK (2012) Tob1 induces apoptosis and inhibits proliferation, migration and invasion of gastric cancer cells by activating Smad4 and inhibiting betacatenin signaling. Int J Oncol 41(3):839–848. https://doi.org/10.3892/ijo.2012.1517
Lachke SA (2022) RNA-binding proteins and post-transcriptional regulation in lens biology and cataract: mediating spatiotemporal expression of key factors that control the cell cycle, transcription, cytoskeleton and transparency. Exp Eye Res 214:108889. https://doi.org/10.1016/j.exer.2021.108889
Lachke SA, Maas RL (2011) RNA granules and cataract. Expert Rev Ophthalmol 6(5):497–500. https://doi.org/10.1586/eop.11.53
Lachke SA, Alkuraya FS, Kneeland SC, Ohn T, Aboukhalil A, Howell GR, Saadi I, Cavallesco R, Yue Y, Tsai AC, Nair KS, Cosma MI, Smith RS, Hodges E, Alfadhli SM, Al-Hajeri A, Shamseldin HE, Behbehani A, Hannon GJ, Bulyk ML, Drack AV, Anderson PJ, John SW, Maas RL (2011) Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Sci (New York NY) 331(6024):1571–1576. https://doi.org/10.1126/science.1195970
Lachke SA, Ho JW, Kryukov GV, O’Connell DJ, Aboukhalil A, Bulyk ML, Park PJ, Maas RL (2012) iSyTE: integrated systems Tool for Eye gene discovery. Investig Ophthalmol Vis Sci 53(3):1617–1627. https://doi.org/10.1167/iovs.11-8839
Li XA, Beebe DC (1991) Messenger RNA stabilization in chicken lens development: a reexamination. Dev Biol 146(1):239–241
Lovicu FJ, McAvoy JW, de Iongh RU (2011) Understanding the role of growth factors in embryonic development: insights from the lens. Philosophical Trans Royal Soc Lond Ser B Biol Sci 366(1568):1204–1218. https://doi.org/10.1098/rstb.2010.0339
Maekawa M, Nishida E, Tanoue T (2002) Identification of the anti-proliferative protein Tob as a MAPK substrate. J Biol Chem 277(40):37783–37787. https://doi.org/10.1074/jbc.M204506200
Maekawa M, Yamamoto T, Nishida E (2004) Regulation of subcellular localization of the antiproliferative protein Tob by its nuclear export signal and bipartite nuclear localization signal sequences. Exp Cell Res 295(1):59–65. https://doi.org/10.1016/j.yexcr.2003.12.016
Martinez G, de Iongh RU (2010) The lens epithelium in ocular health and Disease. Int J Biochem Cell Biol 42(12):1945–1963
Martinez G, Wijesinghe M, Turner K, Abud HE, Taketo MM, Noda T, Robinson ML, de Iongh RU (2009) Conditional mutations of beta-catenin and APC reveal roles for canonical wnt signaling in lens differentiation. Investig Ophthalmol Vis Sci 50(10):4794–4806 doi:iovs.09-3567 [pii]. https://doi.org/10.1167/iovs.09-3567
McAvoy JW (1978) Cell division, cell elongation and distribution of alpha-, beta- and gamma-crystallins in the rat lens. J Embryol Exp Morphol 44:149–165
McAvoy JW, Dawes LJ, Sugiyama Y, Lovicu FJ (2017) Intrinsic and extrinsic regulatory mechanisms are required to form and maintain a lens of the correct size and shape. Exp Eye Res 156:34–40. https://doi.org/10.1016/j.exer.2016.04.009
Mochizuki T, Masai I (2014) The lens equator: a platform for molecular machinery that regulates the switch from cell proliferation to differentiation in the vertebrate lens. Dev Growth Differ 56(5):387–401. https://doi.org/10.1111/dgd.12128
Moser J, Fritzler M (2013) Relationship of Other Cytoplasmic Ribonucleoprotein Bodies (cRNPB) to GW/P Bodies. In: Chan EKL, Fritzler MJ (eds) Ten Years of Progress in GW/P Body Research, vol 768. Advances in Experimental Medicine and Biology. Springer New York, pp 213–242. https://doi.org/10.1007/978-1-4614-5107-5_13
Ogami K, Hosoda N, Funakoshi Y, Hoshino S (2014) Antiproliferative protein Tob directly regulates c-myc proto-oncogene expression through cytoplasmic polyadenylation element-binding protein CPEB. Oncogene 33(1):55–64. https://doi.org/10.1038/onc.2012.548
Olszewska M, Bujarski JJ, Kurpisz M (2012) P-bodies and their functions during mRNA cell cycle: mini-review. Cell Biochem Funct 30(3):177–182. https://doi.org/10.1002/cbf.2804
Pathania M, Wang Y, Simirskii VN, Duncan MK (2016) β1-integrin controls cell fate specification in early lens development. Differ Res Biol Divers 92(4):133–147. https://doi.org/10.1016/j.diff.2016.08.002
Peek R, McAvoy JW, Lubsen NH, Schoenmakers JG (1992) Rise and fall of crystallin gene messenger levels during fibroblast growth factor induced terminal differentiation of lens cells. Dev Biol 152(1):152–160
Putnam A, Thomas L, Seydoux G (2023) RNA granules: functional compartments or incidental condensates? Genes Dev 37(9–10):354–376. https://doi.org/10.1101/gad.350518.123
Rajagopal R, Huang J, Dattilo LK, Kaartinen V, Mishina Y, Deng CX, Umans L, Zwijsen A, Roberts AB, Beebe DC (2009) The type I BMP receptors, Bmpr1a and Acvr1, activate multiple signaling pathways to regulate lens formation. Dev Biol 335(2):305–316. https://doi.org/10.1016/j.ydbio.2009.08.027
Ramaswami M, Taylor JP, Parker R (2013) Altered ribostasis: RNA-protein granules in degenerative disorders. Cell 154(4):727–736. https://doi.org/10.1016/j.cell.2013.07.038
Ripin N, Parker R (2022) Are stress granules the RNA analogs of misfolded protein aggregates? RNA (New York NY) 28(1):67–75. https://doi.org/10.1261/rna.079000.121
Salerno F, van Lier RA, Wolkers MC (2014) Better safe than sorry: TOB1 employs multiple parallel regulatory pathways to keep Th17 cells quiet. Eur J Immunol 44(3):646–649. https://doi.org/10.1002/eji.201444465
Shapouri F, Saeidi S, de Iongh RU, Casagranda F, Western PS, McLaughlin EA, Sutherland JM, Hime GR, Familari M (2015) Tob1 is expressed in developing and adult gonads and is associated with the P-body marker, Dcp2. Cell and tissue research. https://doi.org/10.1007/s00441-015-2328-z
Suzuki T, Ajima JKT, Nakamura R, Yoshida T, Yamamoto Y T (2002) Phosphorylation of three regulatory serines of Tob by Erk1 and Erk2 is required for ras-mediated cell proliferation and transformation. Genes Dev 16(11):1356–1370. https://doi.org/10.1101/gad.962802
Teo ZL, McQueen-Miscamble L, Turner K, Martinez G, Madakashira B, Dedhar S, Robinson ML, de Iongh RU (2014) Integrin linked kinase (ILK) is required for lens epithelial cell survival, proliferation and differentiation. Exp Eye Res 121:130–142. https://doi.org/10.1016/j.exer.2014.01.013
Terrell AM, Anand D, Smith SF, Dang CA, Waters SM, Pathania M, Beebe DC, Lachke SA (2015) Molecular characterization of mouse lens epithelial cell lines and their suitability to study RNA granules and cataract associated genes. Exp Eye Res 131:42–55. https://doi.org/10.1016/j.exer.2014.12.011
Tzachanis D, Freeman GJ, Hirano N, van Puijenbroek AA, Delfs MW, Berezovskaya A, Nadler LM, Boussiotis VA (2001) Tob is a negative regulator of activation that is expressed in anergic and quiescent T cells. Nat Immunol 2(12):1174–1182. https://doi.org/10.1038/ni730
West-Mays JA, Pino G, Lovicu FJ (2010) Development and use of the lens epithelial explant system to study lens differentiation and cataractogenesis. Prog Retin Eye Res 29(2):135–143
Winkler GS (2010) The mammalian anti-proliferative BTG/Tob protein family. J Cell Physiol 222(1):66–72. https://doi.org/10.1002/jcp.21919
Wolf L, Gao CS, Gueta K, Xie Q, Chevallier T, Podduturi NR, Sun J, Conte I, Zelenka PS, Ashery-Padan R, Zavadil J, Cvekl A, Bethesda (2013) Md) 3 (12):2239–2255. doi:https://doi.org/10.1534/g3.113.008698
Wolozin B (2014) Physiological protein aggregation run amuck: stress granules and the genesis of neurodegenerative Disease. Discov Med 17(91):47–52
Xie Q, McGreal R, Harris R, Gao CY, Liu W, Reneker LW, Musil LS, Cvekl A (2015) Regulation of c-Maf and alphaa-crystallin in ocular Lens by FGF signaling. J Biol Chem. https://doi.org/10.1074/jbc.M115.705103
Xiong B, Rui Y, Zhang M, Shi K, Jia S, Tian T, Yin K, Huang H, Lin S, Zhao X, Chen Y, Chen YG, Lin SC, Meng A (2006) Tob1 controls dorsal development of zebrafish embryos by antagonizing maternal beta-catenin transcriptional activity. Dev Cell 11(2):225–238. https://doi.org/10.1016/j.devcel.2006.06.012
Yoshida Y, Tanaka S, Umemori H, Minowa O, Usui M, Ikematsu N, Hosoda E, Imamura T, Kuno J, Yamashita T, Miyazono K, Noda M, Noda T, Yamamoto T (2000) Negative regulation of BMP/Smad signaling by Tob in osteoblasts. Cell 103(7):1085–1097
Yoshida Y, Nakamura T, Komoda M, Satoh H, Suzuki T, Tsuzuku JK, Miyasaka T, Yoshida EH, Umemori H, Kunisaki RK, Tani K, Ishii S, Mori S, Suganuma M, Noda T, Yamamoto T (2003a) Mice lacking a transcriptional corepressor Tob are predisposed to cancer. Genes Dev 17(10):1201–1206. https://doi.org/10.1101/gad.1088003
Yoshida Y, von Bubnoff A, Ikematsu N, Blitz IL, Tsuzuku JK, Yoshida EH, Umemori H, Miyazono K, Yamamoto T, Cho KW (2003b) Tob proteins enhance inhibitory smad-receptor interactions to repress BMP signaling. Mech Dev 120(5):629–637
Acknowledgements
The authors gratefully acknowledge Professor Tadashi Yamamoto and Dr Toru Suzuki, from Okinawa Institute of Science and Technology Graduate School, for supplying tissues from the Tob1-/- mice. Confocal microscopy was performed at the Biological Optical Microscopy Platform (BOMP), The University of Melbourne (www.microscopy.unimelb.edu.au).
Funding
This work was funded by internal funds (R.deI.) at the University of Melbourne. R.C.P. gratefully acknowledges the support of an internship from the “Programa CiÊncias Sem Fronteiras” funded by the Brazilian government.
Author information
Authors and Affiliations
Contributions
Conceptualization, R.deI., M.F. and G.R.H.; formal analysis, R.C.P., X.Y., and R.deI.; ex-perimental investigation, R.C.P., X.Y., G.M. and F.J.L.; resources, R.deI. and F.J.L.; data cu-ration, R.deI.; writing—original draft preparation, R.C.P., X.Y. and R.deI.; writing—review and editing, R.deI., M.F. F.J.L. and G.R.H.; visualization, X.X.; supervision, R.deI., G.M. and M.F.; project administration, R.deI. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional Review Board
All animal procedures were approved by the Animal Ethics Committees of the University of Melbourne (AEC# 1413343) and University of Sydney (AEC# 2015/844; 714) and were carried out in accordance with the Association for Research in Vision and Ophthalmology (ARVO) statement for the Use of Animals in Ophthalmic and Vision Research.
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Perez, R.C., Yang, X., Familari, M. et al. TOB1 and TOB2 mark distinct RNA processing granules in differentiating lens fiber cells. J Mol Histol 55, 121–138 (2024). https://doi.org/10.1007/s10735-023-10177-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10735-023-10177-y