Abstract
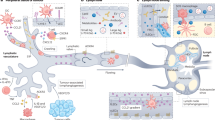
Glioblastoma is the most lethal primary brain tumor with glioblastoma stem cells (GSCs) atop a cellular hierarchy. GSCs often reside in a perivascular niche, where they receive maintenance cues from endothelial cells, but the role of heterogeneous endothelial cell populations remains unresolved. Here, we show that lymphatic endothelial-like cells (LECs), while previously unrecognized in brain parenchyma, are present in glioblastomas and promote growth of CCR7-positive GSCs through CCL21 secretion. Disruption of CCL21–CCR7 paracrine communication between LECs and GSCs inhibited GSC proliferation and growth. LEC-derived CCL21 induced KAT5-mediated acetylation of HMGCS1 on K273 in GSCs to enhance HMGCS1 protein stability. HMGCS1 promoted cholesterol synthesis in GSCs, favorable for tumor growth. Expression of the CCL21–CCR7 axis correlated with KAT5 expression and HMGCS1K273 acetylation in glioblastoma specimens, informing patient outcome. Collectively, glioblastomas contain previously unrecognized LECs that promote the molecular crosstalk between endothelial and tumor cells, offering potentially alternative therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The RNA-seq data related to this study are available at the NCBI Gene Expression Omnibus under accession no. GSE218043, the SMART-seq data in this study are available at the Genome Sequence Archive for Human under accession no. HRA003200 and the MS data related to this study are available at OMIX (China National Center for Bioinformation) under accession code PRJCA014088. All other supporting data of this study are available from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
The analyses presented in this study were conducted using standard workflows.
References
Ostrom, Q. T. et al. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2013–2017. Neuro. Oncol. 22, iv1–iv96 (2020).
Stupp, R. et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 10, 459–466 (2009).
Quail, D. F. & Joyce, J. A. The microenvironmental landscape of brain tumors. Cancer Cell 31, 326–341 (2017).
Singh, S. K. et al. Identification of human brain tumour initiating cells. Nature 432, 396–401 (2004).
Seano, G. & Jain, R. K. Vessel co-option in glioblastoma: emerging insights and opportunities. Angiogenesis 23, 9–16 (2020).
Gilbert, M. R. et al. A randomized trial of bevacizumab for newly diagnosed glioblastoma. N. Engl. J. Med. 370, 699–708 (2014).
Chinot, O. L. et al. Bevacizumab plus radiotherapy-temozolomide for newly diagnosed glioblastoma. N. Engl. J. Med. 370, 709–722 (2014).
Obernier, K. & Alvarez-Buylla, A. Neural stem cells: origin, heterogeneity and regulation in the adult mammalian brain. Development 146, 156059 (2019).
Zhou, W. et al. Periostin secreted by glioblastoma stem cells recruits M2 tumour-associated macrophages and promotes malignant growth. Nat. Cell Biol. 17, 170–182 (2015).
Osswald, M. et al. Brain tumour cells interconnect to a functional and resistant network. Nature 528, 93–98 (2015).
Plate, K. H., Breier, G., Weich, H. A. & Risau, W. Vascular endothelial growth factor is a potential tumour angiogenesis factor in human gliomas in vivo. Nature 359, 845–848 (1992).
Ricci-Vitiani, L. et al. Tumour vascularization via endothelial differentiation of glioblastoma stem-like cells. Nature 468, 824–828 (2010).
Wang, R. et al. Glioblastoma stem-like cells give rise to tumour endothelium. Nature 468, 829–833 (2010).
Cheng, L. et al. Glioblastoma stem cells generate vascular pericytes to support vessel function and tumor growth. Cell 153, 139–152 (2013).
Winkler, E. A. et al. A single-cell atlas of the normal and malformed human brain vasculature. Science 375, eabi7377 (2022).
Louveau, A. et al. Structural and functional features of central nervous system lymphatic vessels. Nature 523, 337–341 (2015).
Da Mesquita, S. et al. Functional aspects of meningeal lymphatics in ageing and Alzheimer’s disease. Nature 560, 185–191 (2018).
Da Mesquita, S. et al. Meningeal lymphatics affect microglia responses and anti-Aβ immunotherapy. Nature 593, 255–260 (2021).
Louveau, A. et al. CNS lymphatic drainage and neuroinflammation are regulated by meningeal lymphatic vasculature. Nat. Neurosci. 21, 1380–1391 (2018).
Song, E. et al. VEGF-C-driven lymphatic drainage enables immunosurveillance of brain tumours. Nature 577, 689–694 (2020).
Ahn, J. H. et al. Meningeal lymphatic vessels at the skull base drain cerebrospinal fluid. Nature 572, 62–66 (2019).
Hu, X. et al. Meningeal lymphatic vessels regulate brain tumor drainage and immunity. Cell Res. 30, 229–243 (2020).
Lee, E. et al. Breast cancer cells condition lymphatic endothelial cells within pre-metastatic niches to promote metastasis. Nat. Commun. 5, 4715 (2014).
Wei, J. C. et al. Tumor-associated Lymphatic endothelial cells promote lymphatic metastasis by highly expressing and secreting SEMA4C. Clin. Cancer Res. 23, 214–224 (2017).
Wei, Y. et al. IL-17A secreted from lymphatic endothelial cells promotes tumorigenesis by upregulation of PD-L1 in hepatoma stem cells. J Hepatol. 71, 1206–1215 (2019).
Xie, Y. et al. Key molecular alterations in endothelial cells in human glioblastoma uncovered through single-cell RNA sequencing. JCI Insight 6, 150861 (2021).
LeBlanc, V. G. et al. Single-cell landscapes of primary glioblastomas and matched explants and cell lines show variable retention of inter- and intratumor heterogeneity. Cancer Cell 40, 379–392 (2022).
Mabbott, N. A., Baillie, J. K., Brown, H., Freeman, T. C. & Hume, D. A. An expression atlas of human primary cells: inference of gene function from coexpression networks. BMC Genomics 14, 632 (2013).
Hansen, A. et al. KSHV-encoded miRNAs target MAF to induce endothelial cell reprogramming. Genes Dev. 24, 195–205 (2010).
Wong, H. L. et al. MT1-MMP sheds LYVE-1 on lymphatic endothelial cells and suppresses VEGF-C production to inhibit lymphangiogenesis. Nat. Commun. 7, 10824 (2016).
Pan, Y. et al. Podoplanin requires sialylated O-glycans for stable expression on lymphatic endothelial cells and for interaction with platelets. Blood 124, 3656–3665 (2014).
Chakarov, S. et al. Two distinct interstitial macrophage populations coexist across tissues in specific subtissular niches. Science 363, aau0964 (2019).
Lim, H. Y. et al. Hyaluronan receptor LYVE-1-expressing macrophages maintain arterial tone through hyaluronan-mediated regulation of smooth muscle cell collagen. Immunity 49, 326–341 (2018).
Drieu, A. et al. Parenchymal border macrophages regulate the flow dynamics of the cerebrospinal fluid. Nature 611, 585–593 (2022).
Gravendeel, L. A. et al. Intrinsic gene expression profiles of gliomas are a better predictor of survival than histology. Cancer Res. 69, 9065–9072 (2009).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Mack, S. C. et al. Chromatin landscapes reveal developmentally encoded transcriptional states that define human glioblastoma. J. Exp. Med. 216, 1071–1090 (2019).
Buonamici, S. et al. CCR7 signalling as an essential regulator of CNS infiltration in T-cell leukaemia. Nature 459, 1000–1004 (2009).
Vaahtomeri, K. et al. Locally triggered release of the chemokine CCL21 promotes dendritic cell transmigration across lymphatic endothelia. Cell Rep. 19, 902–909 (2017).
Maire, C. L. et al. Glioma escape signature and clonal development under immune pressure. J. Clin. Invest. 130, 5257–5271 (2020).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
Brasaemle, D. L. et al. Perilipin A increases triacylglycerol storage by decreasing the rate of triacylglycerol hydrolysis. J. Biol. Chem. 275, 38486–38493 (2000).
Jaeger, K. et al. Structural basis for allosteric ligand recognition in the human CC chemokine receptor 7. Cell 178, 1222–1230 (2019).
Alegaon, S. G. et al. Synthesis, molecular docking and ADME studies of thiazole-thiazolidinedione hybrids as antimicrobial agents. J. Biomol. Struct. Dyn. https://doi.org/10.1080/07391102.2021.1880479 (2022).
Fu, C. et al. Oral bioavailability comparison of artemisinin, deoxyartemisinin, and 10-deoxoartemisinin based on computer simulations and pharmacokinetics in rats. ACS Omega 6, 889–899 (2021).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Kuczynski, E. A., Vermeulen, P. B., Pezzella, F., Kerbel, R. S. & Reynolds, A. R. Vessel co-option in cancer. Nat. Rev. Clin. Oncol. 16, 469–493 (2019).
Tavora, B. et al. Tumoural activation of TLR3-SLIT2 axis in endothelium drives metastasis. Nature 586, 299–304 (2020).
Calabrese, C. et al. A perivascular niche for brain tumor stem cells. Cancer Cell 11, 69–82 (2007).
Chen, J. et al. Cerebrovascular injuries induce lymphatic invasion into brain parenchyma to guide vascular regeneration in zebrafish. Dev. Cell 49, 697–710 (2019).
Jin, X. et al. Targeting glioma stem cells through combined BMI1 and EZH2 inhibition. Nat. Med. 23, 1352–1361 (2017).
Han, M. et al. Therapeutic implications of altered cholesterol homeostasis mediated by loss of CYP46A1 in human glioblastoma. EMBO Mol. Med. 12, e10924 (2020).
Wang, X. et al. MYC-regulated mevalonate metabolism maintains brain tumor-initiating cells. Cancer Res. 77, 4947–4960 (2017).
Villa, G. R. et al. An LXR–cholesterol axis creates a metabolic co-dependency for brain cancers. Cancer Cell 30, 683–693 (2016).
Cheng, C. et al. Glucose-mediated N-glycosylation of SCAP is essential for SREBP-1 activation and tumor growth. Cancer Cell 28, 569–581 (2015).
Yuan, H. et al. Lysine catabolism reprograms tumour immunity through histone crotonylation. Nature 617, 818–826 (2023).
Ye, Z. et al. Targeting microglial metabolic rewiring synergizes with immune-checkpoint blockade therapy for glioblastoma. Cancer Discov. 13, 974–1001 (2023).
Yu, H., Xing, S., Nierves, L., Lange, P. F. & Huan, T. Fold-change compression: an unexplored but correctable quantitative bias caused by nonlinear electrospray ionization responses in untargeted metabolomics. Anal. Chem. 92, 7011–7019 (2020).
Yu, H. & Huan, T. Patterned signal ratio biases in mass spectrometry-based quantitative metabolomics. Anal. Chem. 93, 2254–2262 (2021).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kong, S. et al. Global analysis of lysine acetylome reveals the potential role of CCL18 in non-small cell lung cancer. Proteomics 21, e2000144 (2021).
Gong, Y. et al. Age-associated proteomic signatures and potential clinically actionable targets of colorectal cancer. Mol. Cell. Proteom. 20, 100115 (2021).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Chou, M. F. & Schwartz, D. Biological sequence motif discovery using motif-x. Curr. Protoc. Bioinform. 13, 13 15–13 24 (2011).
Acknowledgements
The authors thank S. Ma from UCSD for the plasmid. This work was supported by grants from the National Institutes of Health (NIH) (T32 CA094186 to K.Y.; F30 CA217065 to R.C.G.; F31 CA243296 to D. Lee; R35 CA197718, R01 CA238662, R01 CA268634 and R01 NS103434 to J.N.R.; and R01NS106155, R01CA159859 and R01CA255369 to M.D.T.) and from the National Natural Science Foundation of China (82273255, 81822034 and 81821002 to S.Z.; 82273473 to Z.Q.; and 82072779 to Xiuxing W.), The National Key Research and Development Program of China (2022YFA1106600, 2017YFA0106800 and 2018YFA0109200 to S.Z.), The Sichuan Science-Technology International Cooperation Project (22ZYZYTS0070 and 2019YFH0144 to S.Z.) and The Shanghai Municipal Health Commission (2022YQ062 to Z.Q.). All graphics were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Conceptualization was the responsibility of L.Z., S.Z. and J.N.R. Methodology was the responsibility of L.Z., Z.Q., Z.Y., L.X., Q.W., H.Y., V.V., J.L. and V.F. Software was the responsibility of Z.Q., O.S., L.D.H., V.F., J.Z. and W.W. Investigation was the responsibility of L.Z., Z.Q., Z.Y., F.L., L.O., T.M.P., W.W., Xiang W. and T.H. Resources were the responsibility of F.F., A.E.S., H.W. and Xiuxing W. Data curation was carried out by L.Z., Z.Q. and Z.Y. Writing of the original draft was the responsibility of L.Z. and Z.Q. Review and editing of the manuscript was carried out by J.N.R., S.Z., K.Y., S.A.G., R.C.G., D. Lv, D. Lee, B.M.L., C.J. and D.D. Supervision was carried out by J.M.B., M.D.T., S.Z. and J.N.R.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LEC-like cells are present in glioblastoma parenchyma.
a, UMAP plot showing cell clusters in scRNA-seq analysis from 4 integrated glioblastoma patients colored by individuals (GSE162631). b, Bubble plot showing expression levels of marker genes in cell clusters in scRNA-seq analysis from 4 integrated glioblastoma patients (GSE162631). c, UMAP plot showing cell clusters in scRNA-seq analysis from 4 integrated glioblastoma patients could express endothelial cell makers ESAM. d-g, Prediction of percentage of predicted LECs in endothelial cell clusters in scRNA-seq analysis from a panel of glioblastoma patients (GSE162631). h, Expression of key LEC-specific and BEC-specific markers, respectively in the GSE162631 dataset. i, Bubble plot showing expression levels of marker genes in cell clusters in scRNA-seq analysis from 12 integrated glioblastoma patients (GSE173278). j–k, UMAP showing expression levels of endothelial cell specific genes ESAM and PECAM in two endothelial cell populations in scRNA-seq analysis of 12 glioblastoma patients. l, LEC signature scores between predicted LECs and BECs from 12 integrated glioblastoma patients (GSE173278). For box-and-whisker plot, the box indicates interquartile range (IQR), the line in the box indicates the median, the whiskers indicate points within Q3 + 1.5 × IQR and Q1 – 1.5 × IQR, and the points beyond whiskers indicate outliers. Q1 and Q3 denote the first and third quartiles, respectively. P values were calculated using unpaired two-tailed t-test. m, Bubble plot showing expression levels of LEC marker genes Podoplanin, FOXC2, PROX1, and GATA2 are highly enriched in LEC clusters compared with BEC clusters in scRNA-seq analysis from 12 integrated glioblastoma patients (GSE173278). n, The large-scale chromosomal copy number variations (CNVs) in single cells (GSE173278) were inferred using inferCNV packages. The myeloid cells, oligodendrocytes and pericytes were used as normal references. CNVs in endothelial cell cluster and malignant cluster were shown in the heat map.
Extended Data Fig. 2 Flow cytometry gating strategies for the LEC-like cell in glioblastoma patients.
Gating strategies for FACS analysis shown in Fig. 1n and the percentage of LYVE1+podoplanin+ LECs in CD45− CD31+ endothelial cells of a panel of GBM patient fresh samples.
Extended Data Fig. 3 LEC-like cell abundance is correlated with patient survival.
a-b, Gating strategies for FACS analysis shown in Fig. 2a,b. c, Representative images of immunohistochemical staining of pan-endothelial cell maker CD31 and LEC marker Podoplanin in non-neoplastic brain tissues and glioblastoma tissues. Glioblastoma tissue fractions exhibit enriched LECs. Scale bars, 50 μm. Images are representative of three independent experiments. d, Two-tailed Spearman correlation between HBEC cells and GBM patient 25-derived BECs using proteomics data. e, Two-tailed Spearman correlation between HBEC cells and glioblastoma patient 26-derived BECs using proteomics data. f, Spearman correlation between HBE-5i cells and glioblastoma patient-derived BECs using SMART-seq data. g, Spearman correlation between commercial LECs and glioblastoma patient-derived LECs using SMART-seq data. h, FACS plots demonstrating isolation of LEC-like cells from GL261 glioma cells. Cells from mouse tumor tissue were stained with CD45-APC-Cy7, CD31-PE, and Podoplanin-FITC.
Extended Data Fig. 4 CCR7 mediates the interplay between LECs and GSCs.
a, Differential overall survival of glioblastoma patients stratified by different CCR6 expressions in the TCGA glioblastoma dataset. P values were calculated using log-rank test. b, Differential overall survival of glioblastoma patients stratified by CXCR4 expression in TCGA glioblastoma data. P values were calculated using log-rank test. c, mRNA levels of top 5 upregulated chemokine receptors in GSCs and NSCs analyzed by qPCR assay. Data are presented as mean ± SD from 3 independent experiments. P values were calculated using two-tailed unpaired Student’s t-test. d, Differential expression of CCR7 between normal and glioblastoma patients in Gravendeel data. GBM (n = 159) vs. normal brain (n = 8). P values were calculated using two-tailed unpaired Student’s t-test. e, Differential expression of CCR7 between normal and glioblastoma patients in TCGA glioblastoma dataset. GBM (n = 528) vs. normal brain (n = 10). P values were calculated using two-tailed unpaired Student’s t-test. f, CCR7+SOX2+/SOX2+ cells in both GSCs and NSCs were detected (n = 5). P values were calculated using two-tailed unpaired Student’s t-test. g, FACS dot plots showing the CD133− CCR7+ and CD133− CCR7− DGC2907. 96.3% of CD133− DGC2907 were CCR7 negative, 95% CI. h, FACS dot plots showing CD133-CCR7+ and CD133−CCR7− DGC20. 96.3% of CD133− DGC2907 were CCR7 negative, 95% CI. i, Spheroid formation assay was performed for GSCs transduced with either shCONT, shCCR7.291, or shCCR7.183 and co-cultured with LECs. n = 6 each group. j, Statistics for spheroid formation assay were performed for GSCs expressing shCONT, shCCR7.291, or shCCR7.183 co-cultured with LECs. Data are presented as mean ± SD from six independent experiments. P values were calculated using two-tailed unpaired Student’s t-test. k and l, Representative FACS dot plots showing CCR7high and CCR7low GSC2907(k) and GSC20(l). For box-and-whisker plot, the box indicates interquartile range (IQR), the line in the box indicates the median, the top of whiskers indicates the maxima and the bottom of the whiskers indicate the minima, Q1 and Q3 denote the first and third quartiles, respectively.
Extended Data Fig. 5 LEC-secreted CCL21, but not CCL19, stimulates proliferation of CCR7+ GSCs in vitro.
a, Cell counting was performed for GSC1552 0, 2, 4, and 6 days after adding either IgG (10 ng ml−1), CCL19 recombinant protein (10 ng ml−1), or CCL21 recombinant protein (10 ng ml−1). b, Quantification of spheres formation of GSC2907 (left), GSC20 (middle), or GSC1552 (right) when treated with either IgG, CCL19 recombinant protein, or CCL21 recombinant protein. Data are presented as mean ± SD from 6 independent experiments. P-values were calculated using two-tailed unpaired Student’s t-test. c, Representative images of spheres formed by GSCs when treated with either IgG, CCL19 recombinant protein, or CCL21 recombinant protein. Scale bar, 50 μm. Images are representative of six independent experiments. d, Cell counting was performed 0, 2, 4, 6 days after adding either IgG or CCL21 monoclonal antibody. P-values were calculated using two-tailed unpaired Student’s t-test. e, Statistics for spheroid formation of GSCs treated with CCL21 monoclonal antibody when co-cultured with LECs. Data are presented as mean ± SD from 6 independent experiments. P-values were calculated using two-tailed unpaired Student’s t-test. f, Cell counting was performed for GSC1552 cells transfected with either shCONT or shCCR7 and treated with CCL21(n = 6). g and h, Representative FACS dot plots showing CCR7+ and CD133+ CT2A cells (g) or GL261 cells (h). i, Schematic model that illustrates cell sorting strategy of CD133+CCR7+ murine glioma cell lines (GL261 and CT2A) and implantation into CCL21−/− mice. j, qPCR analysis showing mRNA expression of CCL21 following shRNA-mediated knockdown. Data are presented as mean ± SD from three independent experiments. P-values were calculated using two-tailed unpaired Student’s t-test. k, Cell counting for GSCs co-cultured with either LECs-shCONT, LECs-shCCL21.242, or LECs-shCCL21.214 was performed on Day 0, 2, 4, and 6. (a-b, d–f, k) Data are presented as mean ± SD from six independent experiments. P-values were calculated using one-way ANOVA test.
Extended Data Fig. 6 Acetylproteome of GSCs stimulated with CCL21.
a, Distribution of subcellular localization of proteins with quantified acetylation sites in GSC20. b, Protein-protein interaction modules for metabolic processes were demonstrated. Proteins with differential acetylation sites obtained from GSC20 cells when treated with either IgG or CCL21 recombinant protein. c, Protein-protein interaction modules for proteins with differential acetylation sites obtained from GSC20 cells when treated with either IgG or CCL21 recombinant protein.
Extended Data Fig. 7 HMGCS1 is acetylated at K273 in GSCs stimulated with CCL21.
a, Specificity of the anti-acetylated lysine 273 antibody was confirmed by its ability to recognize the acetylated, but not unacetylated, peptide in GSC1552 cells. Images are representative of three independent experiments. b, Flag-tagged wild-type HMGCS1-expressing GSC20 cells were lysed, which were subjected to immunoprecipitation with a Flag antibody. The immunoprecipitated HMGCS1 was de-acetylated by CobB. c, Enzymatic activity of HMGCS1WT or HMGCS1K273Q was tested using an in vitro assay using purified His-tagged HMGCS1 or HMGCS1K273Q protein with acetyl-CoA. The reactions were performed at 30 °C for 30 minutes. Data are presented as mean ± SD from three independent experiments. P-values were calculated using two-tailed unpaired Student’s t-test. d, HMGCS1 was stably knocked down with shRNA in GSC20 and GSC23 cells. shRNA-resistant wild-type and K273Q mutant of HMGCS1 were stably re-expressed in the knockdown cells to a level compatible with endogenous HMGCS1. HMGCS1 knockdown efficiency and re-expression level were determined by western blot. Images are representative of three independent experiments.
Extended Data Fig. 8 The BBB-penetrating properties of CMP2105.
a, ADME properties of conformers of Cmp2105 using Qikprop. b, HRMS Extracted ion chromatogram (XIC) of CMP2105, m/z 476.1962 (−0.01ppm mass error from theoretical) at retention time 4.6min in homogenized brain extract. c, HRMS HCD MS2 fragmentation spectrum of CMP2105, m/z 476.1962 parent ion, showing diagnostic fragment of 115.1122.
Supplementary information
Supplementary Table 1
Supplementary Tables 1 and 2.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 7
Unprocessed western blots.
Source Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, L., Qiu, Z., Yang, Z. et al. Lymphatic endothelial-like cells promote glioblastoma stem cell growth through cytokine-driven cholesterol metabolism. Nat Cancer 5, 147–166 (2024). https://doi.org/10.1038/s43018-023-00658-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43018-023-00658-0
This article is cited by
-
Cholesterol-fuelled glioblastoma
Nature Reviews Cancer (2024)