Abstract
The protein S-nitrosylation (SNO) is a significant post-translational modification that affects the stability, activity, cellular localization, and function of proteins. Therefore, highly accurate prediction of SNO sites aids in grasping biological function mechanisms. In this document, we have constructed a predictor, named PPSNO, forecasting protein SNO sites using stacked integrated learning. PPSNO integrates multiple machine learning techniques into an ensemble model, enhancing its predictive accuracy. First, we established benchmark datasets by collecting SNO sites from various sources, including literature, databases, and other predictors. Second, various techniques for feature extraction are applied to derive characteristics from protein sequences, which are subsequently amalgamated into the PPSNO predictor for training. Five-fold cross-validation experiments show that PPSNO outperformed existing predictors, such as PSNO, PreSNO, pCysMod, DeepNitro, RecSNO, and Mul-SNO. The PPSNO predictor achieved an impressive accuracy of 92.8%, an area under the curve (AUC) of 96.1%, a Matthews correlation coefficient (MCC) of 81.3%, an F1-score of 85.6%, an SN of 79.3%, an SP of 97.7%, and an average precision (AP) of 92.2%. We also employed ROC curves, PR curves, and radar plots to show the superior performance of PPSNO. Our study shows that fused protein sequence features and two-layer stacked ensemble models can improve the accuracy of predicting SNO sites, which can aid in comprehending cellular processes and disease mechanisms. The codes and data are available at https://github.com/serendipity-wly/PPSNO.
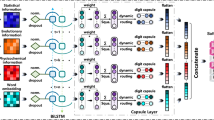
Graphical Abstract









Similar content being viewed by others
Data Availability
Publicly available datasets were analyzed in this study. Codes and data are available at https://github.com/serendipity-wly/PPSNO.
References
Jia J, Arif A, Terenzi F et al (2014) Target-selective protein S-nitrosylation by sequence motif recognition. Cell 159:623–634. https://doi.org/10.1016/j.cell.2014.09.032
Fernando V, Zheng X, Walia Y et al (2019) S-nitrosylation: an emerging paradigm of redox signaling. Antioxidants 8:404. https://doi.org/10.3390/antiox8090404
Hayashi H, Hess DT, Zhang R et al (2018) S-nitrosylation of β-arrestins biases receptor signaling and confers ligand independence. Mol Cell 70:473-487.e6. https://doi.org/10.1016/j.molcel.2018.03.034
Rizza S, Cardaci S, Montagna C et al (2018) S-nitrosylation drives cell senescence and aging in mammals by controlling mitochondrial dynamics and mitophagy. Proc Natl Acad Sci 115:E3388–E3397. https://doi.org/10.1073/pnas.1722452115
Li F, Sonveaux P, Rabbani ZN et al (2007) Regulation of HIF-1α stability through S-nitrosylation. Mol Cell 26:63–74. https://doi.org/10.1016/j.molcel.2007.02.024
Wang Z (2012) Protein S-nitrosylation and cancer. Cancer Lett 2:123–129. https://doi.org/10.1016/j.canlet.2012.03.009
Wijasa TS, Sylvester M, Brocke-Ahmadinejad N et al (2020) Quantitative proteomics of synaptosome S-nitrosylation in Alzheimer’s disease. J Neurochem 152:710–726. https://doi.org/10.1111/jnc.14870
Piroddi M, Palmese A, Pilolli F et al (2011) Plasma nitroproteome of kidney disease patients. Amino Acids 40:653–667. https://doi.org/10.1007/s00726-010-0693-1
Hao G, Derakhshan B, Shi L et al (2006) SNOSID, a proteomic method for identification of cysteine S-nitrosylation sites in complex protein mixtures. Proc Natl Acad Sci 103:1012–1017. https://doi.org/10.1073/pnas.0508412103
Xue Y, Liu Z, Gao X et al (2010) GPS-SNO: computational prediction of protein S-nitrosylation sites with a modified GPS algorithm. PLoS ONE 5:e11290. https://doi.org/10.1371/journal.pone.0011290
Lee T-Y, Chen Y-J, Lu T-C et al (2011) SNOSite: exploiting maximal dependence decomposition to identify cysteine S-nitrosylation with substrate site specificity. PLoS ONE 6:e21849. https://doi.org/10.1371/journal.pone.0021849
Li B-Q, Hu L-L, Niu S et al (2012) Predict and analyze S-nitrosylation modification sites with the mRMR and IFS approaches. J Proteom 75:1654–1665. https://doi.org/10.1016/j.jprot.2011.12.003
Xu Y, Ding J, Wu L-Y (2013) iSNO-PseAAC: predict cysteine S-nitrosylation sites in proteins by incorporating position specific amino acid propensity into pseudo amino acid composition. PLoS ONE 8:e55844. https://doi.org/10.1371/journal.pone.0055844
Xu Y, Shao X-J, Wu L-Y et al (2013) iSNO-AAPair: incorporating amino acid pairwise coupling into PseAAC for predicting cysteine S-nitrosylation sites in proteins. PeerJ 1:e171. https://doi.org/10.7717/peerj.171
Zhang J, Zhao X, Sun P (2014) PSNO: predicting cysteine S-nitrosylation sites by incorporating various sequence-derived features into the general form of Chou’s PseAAC. Int J Mol Sci 15:11204–11219. https://doi.org/10.3390/ijms150711204
Hasan MdM, Manavalan B, MstS K (2019) Prediction of S-nitrosylation sites by integrating support vector machines and random forest. Mol Omics 15:451–458. https://doi.org/10.1039/C9MO00098D
Qiu W-R, Wang Q-K, Guan M-Y et al (2021) Predicting S-nitrosylation proteins and sites by fusing multiple features. Math Biosci Eng 18:9132–9147. https://doi.org/10.3934/mbe.2021450
Xie Y, Luo X, Li Y et al (2018) DeepNitro: prediction of protein nitration and nitrosylation sites by deep learning. Genom Proteom Bioinform 16:294–306. https://doi.org/10.1016/j.gpb.2018.04.007
Siraj A, Chantsalnyam T, Tayara H (2021) RecSNO: prediction of protein S-nitrosylation sites using a recurrent neural network. IEEE Access 9:6674–6682. https://doi.org/10.1109/ACCESS.2021.3049142
Fu H, Yang Y, Wang X et al (2019) DeepUbi: a deep learning framework for prediction of ubiquitination sites in proteins. BMC Bioinformatics 20:1–10. https://doi.org/10.1186/s12859-019-2677-9
Zhao Q, Ma J, Wang Y et al (2022) Mul-SNO: a novel prediction tool for S-nitrosylation sites based on deep learning methods. IEEE J Biomed Health Inform 26:2379–2387. https://doi.org/10.1109/JBHI.2021.3123503
Li S, Yu K, Wu G et al (2021) pCysMod: prediction of multiple cysteine modifications based on deep learning framework. Front Cell Dev Biol 9:617366. https://doi.org/10.3389/fcell.2021.617366
Fu L, Niu B, Zhu Z et al (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Chen Y-J, Ku W-C, Lin P-Y et al (2010) S-alkylating labeling strategy for site-specific identification of the s-nitrosoproteome. J Proteome Res 9:6417–6439. https://doi.org/10.1021/pr100680a
Stephen FA (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Gao W, Xu D, Li H et al (2023) Identification of adaptor proteins by incorporating deep learning and PSSM profiles. Methods 209:10–17. https://doi.org/10.1016/j.ymeth.2022.11.001
Lee T-Y, Chen S-A, Hung H-Y (2011) Incorporating distant sequence features and radial basis function networks to identify ubiquitin conjugation sites. PLoS ONE 6:e17331. https://doi.org/10.1371/journal.pone.0017331
Chen Z, Zhao P, Li F et al (2018) iFeature: a python package and web server for features extraction and selection from protein and peptide sequences. Bioinformatics 34:2499–2502. https://doi.org/10.1093/bioinformatics/bty140
Li Z-R, Lin HH, Han LY et al (2006) PROFEAT: a web server for computing structural and physicochemical features of proteins and peptides from amino acid sequence. Nucleic Acids Res 34:W32–W37
Ju Z, Wang S-Y (2020) Prediction of lysine formylation sites using the composition of k-spaced amino acid pairs via Chou’s 5-steps rule and general pseudo components. Genomics 112:859–866. https://doi.org/10.1016/j.ygeno.2019.05.027
Zhao Y, He N, Chen Z (2020) Identification of protein lysine crotonylation sites by a deep learning framework with convolutional neural networks. IEEE Access 8:14244–14252. https://doi.org/10.1109/ACCESS.2020.2966592
Chou K-C (2005) Using amphiphilic pseudo amino acid composition to predict enzyme subfamily classes. Bioinformatics 21:10–19. https://doi.org/10.1093/bioinformatics/bth466
Zhang J, Liu B (2019) A review on the recent developments of sequence-based protein feature extraction methods. Curr Bioinforma 14:190–199. https://doi.org/10.2174/1574893614666181212102749
Saravanan V, Gautham N (2015) Harnessing computational biology for exact linear B-cell epitope prediction: a novel amino acid composition-based feature descriptor. Omics J Integr Biol 19:648–658. https://doi.org/10.1089/omi.2015.0095
Dubchak I, Muchnik I, Holbrook SR (1995) Prediction of protein folding class using global description of amino acid sequence. Proc Natl Acad Sci 92:8700–8704. https://doi.org/10.1073/pnas.92.19.8700
Uzma MU, Halim Z (2023) Protein encoder: an autoencoder-based ensemble feature selection scheme to predict protein secondary structure. Expert Syst Appl 213:119081. https://doi.org/10.1016/j.eswa.2022.119081
Kalyan KS, Rajasekharan A, Sangeetha S (2022) AMMU: a survey of transformer-based biomedical pretrained language models. J Biomed Inform 126:103982. https://doi.org/10.1016/j.jbi.2021.103982
Deng L, Pan J, Xu X et al (2018) PDRLGB: precise DNA-binding residue prediction using a light gradient boosting machine. BMC Bioinformatics 19:522. https://doi.org/10.1186/s12859-018-2527-1
Song X, Zhu J, Tan X et al (2022) XGBoost-based feature learning method for mining COVID-19 novel diagnostic markers. Front Public Health 10:926069. https://doi.org/10.3389/fpubh.2022.926069
Hancock JT, Khoshgoftaar TM (2020) CatBoost for big data: an interdisciplinary review. J Big Data 7:94. https://doi.org/10.1186/s40537-020-00369-8
Rigatti SJ (2017) Random forest. J Insur Med 47:31–39. https://doi.org/10.17849/insm-47-01-31-39.1
Talebi H, Peeters LJM, Otto A (2022) A truly spatial random forests algorithm for geoscience data analysis and modelling. Math Geosci 54:1–22. https://doi.org/10.1007/s11004-021-09946-w
Qiu Z, Liu Q (2021) Protein–protein interaction site prediction using random forest proximity distance. J Bioinform Comput Biol 19:2050042. https://doi.org/10.1142/S0219720020500420
Cabras S, Castellanos ME, Staffetti E (2016) A random forest application to contact-state classification for robot programming by human demonstration. Appl Stoch Models Bus Ind 32:209–227. https://doi.org/10.1002/asmb.2145
Hua S, Sun Z (2001) A novel method of protein secondary structure prediction with high segment overlap measure: support vector machine approach. J Mol Biol 308:397–407. https://doi.org/10.1006/jmbi.2001.4580
Pan X, Shen H-B (2018) Predicting RNA–protein binding sites and motifs through combining local and global deep convolutional neural networks. Bioinformatics 34:3427–3436. https://doi.org/10.1093/bioinformatics/bty364
Fukushima K (1980) Neocognitron: a self-organizing neural network model for a mechanism of pattern recognition unaffected by shift in position. Biol Cybern 36:193–202. https://doi.org/10.1007/BF00344251
Hochreiter S, Schmidhuber J (1997) Long short-term memory. Neural Comput 9:1735–1780. https://doi.org/10.1162/neco.1997.9.8.1735
Bühlmann P, Yu B (2002) Analyzing bagging. Ann Stat 30:927–961. https://doi.org/10.1214/aos/1031689014
Freund Y, Schapire RE (1997) A decision-theoretic generalization of on-line learning and an application to boosting. J Comput Syst Sci 55:119–139. https://doi.org/10.1006/jcss.1997.1504
Le NQK, Huynh T-T (2019) Identifying SNAREs by incorporating deep learning architecture and amino acid embedding representation. Front Physiol 10:1501. https://doi.org/10.3389/fphys.2019.01501
Funding
This research was funded by the Jiangsu Students' platform for innovation and entrepreneurship training program (Grant No. 202310292023Z) and the Natural Science Foundation of Jiangsu Province of China (Grant No. BK20230626).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Ethical Approval
This article does not contain any studies with animals performed by any of the authors.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhu, L., Wang, L., Yang, Z. et al. PPSNO: A Feature-Rich SNO Sites Predictor by Stacking Ensemble Strategy from Protein Sequence-Derived Information. Interdiscip Sci Comput Life Sci 16, 192–217 (2024). https://doi.org/10.1007/s12539-023-00595-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-023-00595-7