Abstract
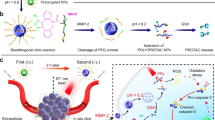
In some cancers mutant p53 promotes the occurrence, development, metastasis and drug resistance of tumours, with targeted protein degradation seen as an effective therapeutic strategy. However, a lack of specific autophagy receptors limits this. Here, we propose the synthesis of biomimetic nanoreceptors (NRs) that mimic selective autophagy receptors. The NRs have both a component for targeting the desired protein, mutant-p53-binding peptide, and a component for enhancing degradation, cationic lipid. The peptide can bind to mutant p53 while the cationic lipid simultaneously targets autophagosomes and elevates the levels of autophagosome formation, increasing mutant p53 degradation. The NRs are demonstrated in vitro and in a patient-derived xenograft ovarian cancer model in vivo. The work highlights a possible direction for treating diseases by protein degradation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The main data that support the findings of this study are available within the paper and its supplementary information. All the relevant data acquired during the study are available for research purposes from the corresponding author on reasonable request. Source data are provided with this paper.
References
Levine, A. J. & Oren, M. The first 30 years of p53: growing ever more complex. Nat. Rev. Cancer 9, 749–758 (2009).
Shaw, P. H. The role of p53 in cell cycle regulation. Pathol. Res. Pract. 192, 669–675 (1996).
Vousden, K. H. & Lane, D. P. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 8, 275–283 (2007).
Vousden, K. H. & Ryan, K. M. p53 and metabolism. Nat. Rev. Cancer 9, 691–700 (2009).
The Cancer Genome Atlas Research Network Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615 (2011).
Olivier, M., Hollstein, M. & Hainaut, P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb. Perspect. Biol. 2, a001008 (2010).
Bertheau, P. et al. p53 in breast cancer subtypes and new insights into response to chemotherapy. Breast 22, S27–S29 (2013).
Freed-Pastor, W. A. & Prives, C. Mutant p53: one name, many proteins. Genes Dev. 26, 1268–1286 (2012).
Muller, P. A. & Vousden, K. H. p53 mutations in cancer. Nat. Cell Biol. 15, 2–8 (2013).
Yue, X. et al. Mutant p53 in cancer: accumulation, gain-of-function, and therapy. J. Mol. Biol. 429, 1595–1606 (2017).
Lukashchuk, N. & Vousden, K. H. Ubiquitination and degradation of mutant p53. Mol. Cell Biol. 27, 8284–8295 (2007).
Schulz-Heddergott, R. et al. Therapeutic ablation of gain-of-function mutant p53 in colorectal cancer inhibits Stat3-mediated tumor growth and invasion. Cancer Cell 34, 298–314 (2018).
Zhang, C. et al. Gain-of-function mutant p53 in cancer progression and therapy. J. Mol. Cell Biol. 12, 674–687 (2020).
Bykov, V. J. N., Eriksson, S. E., Bianchi, J. & Wiman, K. G. Targeting mutant p53 for efficient cancer therapy. Nat. Rev. Cancer 18, 89–102 (2018).
Parrales, A. & Iwakuma, T. Targeting oncogenic mutant p53 for cancer therapy. Front. Oncol. 5, 288 (2015).
Zhang, Y. J. et al. Glutathionylation-dependent proteasomal degradation of wide-spectrum mutant p53 proteins by engineered zeolitic imidazolate framework-8. Biomaterials 271, 120720 (2021).
Qian, J. et al. Enhancing chemotherapy of p53‐mutated cancer through ubiquitination‐dependent proteasomal degradation of mutant p53 proteins by engineered ZnFe‐4 nanoparticles. Adv. Funct. Mater. 30, 2001994 (2020).
Kocaturk, N. M. & Gozuacik, D. Crosstalk between mammalian autophagy and the ubiquitin–proteasome system. Front. Cell Dev. Biol. 6, 128 (2018).
Jing, M. et al. Photoresponsive PAMAM-assembled nanocarrier loaded with autophagy inhibitor for synergistic cancer therapy. Small 17, e2102295 (2021).
Lee, C. W. et al. Selective autophagy degrades nuclear pore complexes. Nat. Cell Biol. 22, 159–166 (2020).
Zhang, Y. et al. Harnessing copper–palladium alloy tetrapod nanoparticle-induced pro-survival autophagy for optimized photothermal therapy of drug-resistant cancer. Nat. Commun. 9, 4236 (2018).
Khaminets, A., Behl, C. & Dikic, I. Ubiquitin-dependent and independent signals in selective autophagy. Trends Cell Biol. 26, 6–16 (2016).
Kirkin, V. & Rogov, V. V. A diversity of selective autophagy receptors determines the specificity of the autophagy pathway. Mol. Cell 76, 268–285 (2019).
Shaid, S., Brandts, C. H., Serve, H. & Dikic, I. Ubiquitination and selective autophagy. Cell Death Differ. 20, 21–30 (2013).
Sarraf, S. A. et al. Loss of TAX1BP1-directed autophagy results in protein aggregate accumulation in the brain. Mol. Cell 80, 779–795 (2020).
Jo, C. et al. Nrf2 reduces levels of phosphorylated tau protein by inducing autophagy adaptor protein NDP52. Nat. Commun. 5, 3496 (2014).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Guida, E. et al. Peptide aptamers targeting mutant p53 induce apoptosis in tumor cells. Cancer Res. 68, 6550–6558 (2008).
Man, N., Chen, Y., Zheng, F., Zhou, W. & Wen, L. P. Induction of genuine autophagy by cationic lipids in mammalian cells. Autophagy 6, 449–454 (2010).
Roberts, R. et al. Autophagy and formation of tubulovesicular autophagosomes provide a barrier against nonviral gene delivery. Autophagy 9, 667–682 (2013).
Li, M. et al. Mono- versus polyubiquitination: differential control of p53 fate by Mdm2. Science 302, 1972–1975 (2003).
Li, M., Luo, J., Brooks, C. L. & Gu, W. Acetylation of p53 inhibits its ubiquitination by Mdm2. J. Biol. Chem. 277, 50607–50611 (2002).
Maeda, H., Nakamura, H. & Fang, J. The EPR effect for macromolecular drug delivery to solid tumors: improvement of tumor uptake, lowering of systemic toxicity, and distinct tumor imaging in vivo. Adv. Drug Deliv. Rev. 65, 71–79 (2013).
Peer, D. et al. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotechnol. 2, 751–760 (2007).
Alexandrova, E. M. et al. Improving survival by exploiting tumour dependence on stabilized mutant p53 for treatment. Nature 523, 352–356 (2015).
Ghosh, M. et al. Mutant p53 suppresses innate immune signaling to promote tumorigenesis. Cancer Cell 39, 494–508 (2021).
Baslan, T. et al. Ordered and deterministic cancer genome evolution after p53 loss. Nature 608, 795–802 (2022).
Parrales, A. et al. DNAJA1 controls the fate of misfolded mutant p53 through the mevalonate pathway. Nat. Cell Biol. 18, 1233–1243 (2016).
Proia, D. A. & Bates, R. C. Ganetespib and HSP90: translating preclinical hypotheses into clinical promise. Cancer Res. 74, 1294–1300 (2014).
Padmanabhan, A. et al. USP15-dependent lysosomal pathway controls p53-R175H turnover in ovarian cancer cells. Nat. Commun. 9, 1270 (2018).
Garufi, A. et al. Degradation of mutant p53H175 protein by Zn(II) through autophagy. Cell Death Dis. 5, e1271 (2014).
Li, Z. et al. Allele-selective lowering of mutant HTT protein by HTT–LC3 linker compounds. Nature 575, 203–209 (2019).
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Li, H. J. et al. Stimuli-responsive clustered nanoparticles for improved tumor penetration and therapeutic efficacy. Proc. Natl Acad. Sci. USA 113, 4164–4169 (2016).
Acknowledgements
This work was supported by the National Key R&D Program of China (2022YFA1205700 to X.Y.), the National Natural Science Foundation of China (T2222014 and 32071398 to Y.Z., 82150118 and 32171375 to L.W., U22A20156 and 51822302 to X.Y., 82202305 to J.Q., 52203163 to Z.C.), the Key R&D Program of Guangdong Province (2020B1515120096 and 2022B0202010002 to Y.Z., 2020B0101030006 to L.W.), the China Postdoctoral Science Foundation (2023M741211 to X.H. and 2023T160230 to J.Q.), Basic and Applied Basic Research of Guangzhou (2023A04J1821 to J.Q.) and the Major Science and Technology projects in the Xinjiang Uygur Autonomous Region (2022A02004 to Y.Z.).
Author information
Authors and Affiliations
Contributions
X.H., Z.C. and J.Q. designed and performed the majority of experiments, analysed the data and wrote the manuscript. T.D. helped detect the SPR of proteins. Y.W. and H.Z. constructed the related plasmids. S.Z., Xiaoli Wang, X.R., W.Z. and Y.X. helped construct the animal models. G.Y provided samples for establishing PDX models. Xingwu Wang provided important support to carry out the in vitro ubiquitination experiment. Y.Z., L.W. and X.Y. conceived the idea, designed the study, interpreted data and revised the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
Y.Z., X.H., Z.C., X.Y. and L.W. have submitted two patent applications (202211119762.4, PCT/CN2023/113588) related to this study. The other authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Peter Tsevtkov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, Figs. 1–45 and Tables 1–3.
Supplementary Video 1
Video of live EGFP–LC3/MDA-MB-231 cells co-incubated with Cy5-labelled mNRs for 2 h. Red, mNRs; green, LC3.
Supplementary Video 2
Video of live EGFP–LC3/MDA-MB-231 cells co-incubated with Cy5-labelled NRs for 2 h. Red, NRs; green, LC3.
Supplementary Data 1
Unprocessed western blots and/or gels for Supplementary Figs. 4, 15, 17, 20–24, 28, 29, 31, 34a and 41.
Source data
Source Data Figs. 2–6
Unprocessed western blots and/or gels for Figs. 2a,c,e,g–i, 3a,b,f,g,i–k, 4b, 5h and 6e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, X., Cao, Z., Qian, J. et al. Nanoreceptors promote mutant p53 protein degradation by mimicking selective autophagy receptors. Nat. Nanotechnol. 19, 545–553 (2024). https://doi.org/10.1038/s41565-023-01562-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-023-01562-5
This article is cited by
-
Nanoreceptors take down mutant p53
Nature Reviews Cancer (2024)