Abstract
Continuously progressive hepatic fibrosis might cause chronic liver diseases, resulting in hepatic failure. The activation of hepatic stellate cells (HSCs) residing in the liver might induce and influence hepatic fibrosis. In the present study, microRNA 3074 (miR-3074) was found increased within transforming growth factorβ (TGF-β)-activated HSCs and enriched within the TGF-β signaling. In activated HSCs by TGF-β, miR-3074 overexpression aggravated TGF-β-induced fibrotic changes, whereas miR-3074 inhibition exerted opposite effects. miR-3074 directly targeted bone morphogenetic protein 7 (BMP7) and inhibited BMP7 expression. Under TGF-β induction, overexpressed BMP7 notably attenuated the promotive roles of miR-3074 overexpression in TGF-β-activated HSCs. Within carbon tetrachloride (CCl4)-caused liver fibrosis murine model, miR-3074 agomir administration promoted, while LV-BMP7 administration alleviated CCl4-induced fibrotic changes; LV-BMP7 significantly attenuated the effects of miR-3074 agomir. Lastly, mmu-miR-3074 also targeted mouse BMP7 and inhibited mouse BMP7 expression. In conclusion, the miR-3074/BMP7 axis regulates TGF-β-caused activation of HSCs in vitro and CCl4-caused murine liver fibrosis in vivo. BMP7-mediated Smad1/5/8 activation might be involved.
Similar content being viewed by others
Introduction
Liver fibrosis, a chronic liver disease caused by the imbalance between generation and deposition of extracellular matrix (ECM) and primarily collagen [1], occurs in most types of chronic liver diseases, resulting in portal hypertension, cirrhosis, and hepatic failure [2], eventually raising the incidence of hepatocellular carcinoma [3].
The dynamic crosstalk between hepatocytes and non-parenchymal cells might induce and influence hepatic fibrosis, where myoblasts that produce ECM exert a critical effect [4]. It has been proposed that hepatic fibrosis cells mainly come from hepatic stellate cells (HSC) residing in liver [5]. HSCs maintain a non-proliferative and quiescent phenotype in normal liver. Under hepatic damage or in vitro inflammation, quiescent HSCs transdifferentiate into proliferative, migratory, contractile, and ECM-producing myofibroblasts [6]. Thus, in experimental liver parenchymal injury and cholestatic liver injury, the activated HSCs may make great contributions to ECM production [5]. Reportedly, multiple signaling pathways could trigger or promote the activation of HSCs [7]; among which transforming growth factorβ (TGF-β) plays a crucial role in liver fibrogenesis.
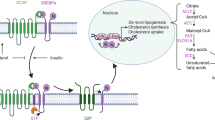
The TGF-β superfamily contains many structurally and functionally similar proteins, including bone morphogenetic proteins (BMPs) [8]. It has shown that TGF-β and BMP-7 share similar downstream SMAD family member (SMAD) pathways, and they balance their biological activities by mutual regulation. Previous studies have pointed out that TGF-β can transmit intracellular signals via SMAD 2/3 phosphorylation in mammals; subsequently, in activated HSCs, activated SMAD3 enhanced type I and type III collagen transcription [9]. On the contrary, SMAD7, an inhibitory SMAD, exerts a negative regulatory effect on the fibrosis induced by SMAD3 [10]. Moreover, SMAD1/5/8 are activated by BMP-7. Phosphorylated SMADs form heterotrimeric complexes via interaction with a distinct co-partner SMAD, SMAD4. The obtained complexes (SMAD1/5/8-SMAD4 and SMAD2/3-SMAD4) translocate from the membrane to the nucleus, and there they can modulate SMAD responsive gene transcription. Within hepatic fibrosis patients, elevated TGF-β1 concentrations are linked with liver fibrosis grade, indicating that TGF-β expression is correlated with progressive hepatic damage [11,12,13]. Interestingly, the abundance of p-SMAD2/3 is increased during rat liver fibrosis [14]. Thus, searching for inhibitory agent of the TGF-β/Smad pathway might provide new directions for treating liver fibrosis.
MicroRNAs (miRNAs), a class of short non-coding RNAs with 18–24 nucleotides in length, modulate gene expression via interaction with target mRNA 3'-UTRs through mRNA destabilization OR translation repression [15, 16]. There is mounting evidence that miRNAs contribute to hepatic fibrogenesis and HSC activation, and miRNAs bind to SMAD proteins in the liver to exert their effects. miR-199a plays an essential role in EMT and enhances hepatic fibrogenesis through promoting the expression of fibrotic genes [17]. miR-146a overexpression inhibits the TGF-β1-induced proliferation of HSCs and promotes HSC apoptosis and alpha-smooth muscle actin (α-SMA) expression, at least partly, by downregulating SMAD4 [18]. TGF-β1 triggers miR-33a expression, which then targets SMAD7 to stimulate HSC activation [19]. Furthermore, miR-21 binds to SMAD7, which further indicates that SMAD7 is targeted in the process of miRNA-related fibrosis [20]. In our previous study, we demonstrated that the miR-503/SMAD7 axis contributes to promoting TGF-β-induced activation of HSCs with the involvement of TGF-β/SMAD signaling [21]. Thus, miRNAs are potential agents regulating the TGF-β/SMAD signaling and affecting TGF-β-caused HSC activation.
Herein, miRNAs that are deregulated in TGF-β-stimulated HSCs and enriched in the TGF-β signaling were analyzed and cross-checked; 7 miRNAs were obtained and miR-3074 was selected because its role in TGF-β-caused activation of HSCs was unclear. TGF-β-caused HSC activation was conducted and the effects of miR-3074 overexpression or inhibition were investigated. We employed three online tools to analyze possible downstream targets of miR-3074, and found BMP7. We validated the putative binding between miR-3074 and BMP7, and investigated the impacts of miR-3074/BMP7 axis upon TGF-β-caused activation of HSCs. Lastly, we constructed a CCl4-caused liver fibrosis murine model, and investigated the miR-3074/BMP7 axis influence in CCl4-caused fibrotic changes.
Materials and methods
Cell lineage and treatment
The human HSCs line LX-2 (SCC064; Sigma-Aldrich, St. Louis, MO, USA) was maintained in 10% fetal bovine serum (FBS, Invitrogen, Carlsbad, CA, USA)-contained RPMI-1640 medium (Invitrogen). The mouse HSCs (mHSCs, BNCC359737) was acquired from BeNa Culture Collection (Beijing, China) and cultured in high glucose DMEM medium (Invitrogen) with 10% FBS. TGF-β (10 ng/ml) was for human HSCs stimulation for 24/48 h [22]. To assess how TGF-β affects HSCs stimulation, cells were treated with 10 µM of SB-431542 (a TGF-β inhibitor, HY-10431, MedChem Express, Monmouth Junction, NJ, USA) for 12 h to specifically inhibit TGF-β [23].
qRT-PCR
As described previously, total RNA extraction, and determination of miRNA and mRNA expression were carried out [24, 25]. RNU6B (for miRNA) or β-actin (for mRNA) expression served as the endogenous control. Relative fold changes were analyzed using the 2−ΔΔCT method. The primer sequences are listed in Table 1.
Immunoblotting
Total protein of cells or tissues was extracted. RIPA buffer was utilized for cell lysis. Protein concentrations were determined using a Bicinchoninic Acid Protein Assay kit (Beyotime, Shanghai, China). Protein samples were subjected to SDS-PAGE for separation. After protein transferring to PVDF membranes (Sigma–Aldrich), primary antibodies anti-α-SMA (55135-1-AP; Proteintech, Wuhan, China), anti-Collagen I (COL1A2, 66761-1-Ig, Proteintech), BMP7 (CSB-PA05805A0Rb, Cusabio, Wuhan, China), p-Smad1/5/8 (AB3848-I, Sigma-Aldrich), Smad1/5/8 (SAB2702532, Sigma-Aldrich), p-Smad2 (CSB-PA783890, Cusabio), Smad2 (12570-1-AP, Proteintech), p-Smad3 (ab52903, Abcam, Cambridge, MA, USA), Smad3 (66516-1-Ig, Proteintech), PKA (55346-1-AP, Proteintech), CREB (12208-1-AP, Proteintech) or β-actin (ab8226; Abcam) were added for incubation, followed by incubation with HRP-conjugated secondary antibody (1:5000). Image J software (NIH, Bethesda, MD, USA) was applied for signal visualization. β-actin acted as the internal control.
Immunofluorescence (IF) staining
IF staining was carried out to measure α-SMA protein content and distribution, as previously described [26]. Namely, the coverslips underwent an incubation with primary antibody against α-SMA (55135-1-AP, Proteintech), and then with FITC-conjugated secondary antibody (SA00003-2, Proteintech) and DAPI (Beyotime). Photographing was done using a fluorescence microscopy (BX53, Olympus, Tokyo, Japan).
Cell transfection and lentivirus infection
The BMP7 overexpression lentivirus (Lv-BMP7) and miR-3074-5p overexpression lentivirus (Lv-miR-3074-5p) were purchased from GenePharma (Shanghai, China). The lentivirus constructions were as previous description [27]. Briefly, the CDS region of BMP7 (human or mouse) and pri-miR-3074 with flanking sequences were cloned by PCR and inserted into pLVX-puro vector, respectively. The primers for PCR are listed in Table 2. Then, the pLVX-puro vector and packaging plasmids were co-transfected into 293T cells using the help of Lipofectamine 3000 (Invitrogen). Forty-eight hours after transfection, the culture medium was refreshed and cells were further culture for 48 h. The culture supernatants were collected for lentivirus purification (Lv-BMP7 and Lv-miR-3074-5p). The empty pLVX-puro vector served as the control vector (Lv-NC). The mouse miR-2074-5p sponge was designed and synthesized by Genepharma and inserted into pLVX-puro vector. The lentivirus packaging was as above mentioned. The obtained lentivirus named Lv-anti-miR-3074-5p. The viral particles were purified and concentrated using ultracentrifugation and the supernatant titer was determined to be 1 × 108 TU/ml. The purified lentivirus of different treatments was infected to HSCs for 24 h.
miR-3074-5p agomir/antagomir (GenePharma) transfection was done for miR-3074-5p inhibition/overexpression in cells. Two specific short hairpin RNA (shRNA) of BMP7 (sh-BMP7#1 and sh-BMP7#2) and scramble oligonucleotides shRNA (sh-NC) were designed and synthesized by Genepharma and then inserted into pLVX-shRNA1 vecotr for silencing BMP7 expression. The vector contained scramble ShRNA sequence was set as negative control (sh-NC). Lipofectamine 3000 (Invitrogen) was utilized for transfection of vectors and agomir/antagomir in HSCs as per the protocols. The sequences of miR-3074-5p agomir/antagomir and sh-BMP7#1/sh-BMP7#2 are listed in Table 2.
MTT assay
MTT assay was used for cell viability detection as previously described [28]. The OD490 nm values were measured and cell viability was detected. Cells treated with only DMEM and MTT were used as the control.
Luciferase reporter assay
The wild-type and mutant-type of human and mouse BMP7 3'UTR luciferase reporter vectors were constructed to verify BMP7 3'UTR bindings with human miR-3074-5p or mouse mmu-miR-3074-5p. The predicted human and mouse miR-3074-5p binding sites in human and mouse BMP7 3'UTR were mutated in mutant-type reporter vectors for the removal of complementarity. Next, the above vectors were co-transfected with miR-3074-5p mimics/inhibitor into 293T cells, HSCs or mHSCs, followed by the determination of luciferase activity utilizing the Dual-Luciferase Reporter Assay System (Promega, WI, USA).
Construction of hepatic fibrosis mouse model
Total 64 C57BL/6 mice (Hunan SJA Laboratory Animal Co., Ltd., Changsha, China) were housed (20–22 °C with a light/dark cycle of 12 h/12 h, with free access to water and diet. Among then, 56 mice were intraperitoneally injected with carbon tetrachloride (CCl4) (diluted in corn oil; 0.5 mL/kg body weight) twice a week for 6 weeks [29], for the establishment of hepatic fibrosis mouse model. Eight mice injected with corn oil served as negative control. After CCl4 initial injection for 2 weeks, mice were allocated into the CCl4 + (Lv-NC), CCl4 + (Lv-miR-3074-5p), CCl4 + (Lv-anti-NC), CCl4 + (Lv-anti-miR-3074-5p), – CCl4 + (Lv-miR-3074-5p + Lv-BMP7), and control groups. Each mouse in the experimental groups was injected with a single dose of 108 TU/ml of Lv-miR-3074-5p, Lv-anti-miR-3074-5p, and Lv-miR-3074-5p along with Lv-BMP7 or its corresponding negative control (100 μl) in tail vein. Mice in the control group were only injected with the same volume of saline. After CCl4 last injection for 2 weeks, mice were euthanized to harvest their livers for subsequent experiments. All experimental protocols got approval from the animal care and use committee of the First Affiliated Hospital, Hengyang Medical School, University of South China.
Hematoxylin and eosin (H&E) staining and Masson’s staining
The collected mouse liver tissues were fixed in 10% buffered formalin for 24 h, paraffin-embedded, and sectioned (5 μm). After that, the sections were stained with H&E or Masson’s staining. Representative images were photographed.
Immunohistochemical (IHC) staining
Following 10-min fixing in 10% formaldehyde, the liver sections were rinsed 3 times with PBS, and then subjected to an incubation (37 °C, 1 h) with antibody against BMP7 (CSB-PA05805A0Rb, Cusabio). The next day, after PBS washing, the sections underwent 30-min incubation with poly-HRP antibody, followed by DAB staining using the DAB staining kit (Boster, Wuhan, China). Finally, a microscope (Olympus) was applied for section observation.
Statistical analysis
The cell experiments were performed for three biological replicates. The animal experiments were performed for eight biological replicates. Experimental data were represented as means ± standard deviation (SD). Differences among groups were analyzed using a one-way analysis of variance (ANOVA) followed by Tukey’s test. GraphPad Prism software 7.0 (GraphPad, La Jolla, CA, USA) was utilized for statistical analysis. p < 0.05 is indicative of statistically significant.
Results
Indentification of miRNAs that might affect TGF-β-induced HSC activation
To identify miRNAs participating in TGF-β-caused HSC activation, we analyzed online dataset GSE151251. In HSCs stimulated with TGF-β, RNAseq analysis detected a total of 1968 deregulated miRNAs. After standardization and homogenization using DEseq2 package in R language, a total of 563 deregulated miRNAs were obtained, 17 of which were downregulated and 20 upregulated with the∣ logFc∣ > 0.2, p < 0.05 (Table S1 and Fig. S1A). The heatmap showed the top 15 up- and downregulated miRNA (Fig. S1B).
Next, based on mirPath V3 (http://snf-515788.vm.okeanos.grnet.gr/index.php? R = miPath/reverse) (contained the data from TargetScan, Tarbase V7.0, and MicroT-CDS5.0) were used to screen for the miRNAs enriched in the TGF-β pathway simultaneously. Among them, 217 miRNAs were associated with the TGF-β signaling in all three sets of data (Fig. S1C). Compared with deregulated miRNAs reported in GSE151251, seven miRNA host genes (MIR22, MIR3074, MIR199a, MIR145, MIR143, MIR21, and MIR214) were found to be enriched in the TGF-β pathway; meanwhile, they were among the deregulated miRNAs reported in GSE151251 (Fig. S1D). Reportedly, of these seven miRNAs, miR-214 [30], miR-21 [31], miR-145 [32], miR-143 [33], miR-199a [34], and miR-22 [35, 36] exert an effect on hepatic fibrogenesis and/or HSC activation (Fig. S1D); thus, miR-3074 has been selected for further experiments.
TGF-β induces fibrotic alterations in HSCs
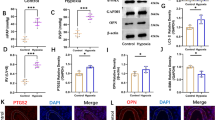
To investigate the specific role of miR-3074, TGF-β-caused HSCs activation was first conducted. We treated HSCs with TGF-β (10 ng/ml) for 0/24/48 h, followed by the determination of COL1A2 and α-SMA expression, which was found increased time-dependently in TGF-β-induced HSCs (Fig. 1A, B). IF staining also showed an increase in α-SMA, a marker of HSCs activation, in TGF-β-induced HSCs (Fig. 1C). TGF-β stimulation notably promoted HSCs viability with time-dependently (Fig. 1D). These results indicated that TGF-β stimulation results in HSCs activation. Then, the expression of miR-3074 by qRT-PCR; Fig. 1E shows that miR-3074 was dramatically upregulated time-dependently in TGF-β stimulated HSCs. Moreover, SB-431542 (a TGF-β inhibitor) markedly reversed upregulated of miR-3074 in TGF-β-induced HSCs (Fig. 1F). According to GSE151251, the expression of MIR3074 in HSCs stimulated with TGF-β showed to be dramatically increased (Fig. 1G). From all the above, miR-3074 might participate in TGF-β-caused activation of HSCs.
TGF-β induces fibrotic changes in hepatic stellate cells (HSCs). HSCs were stimulated with 10 ng/ml TGF-β for 0, 24, or 48 h and examined for the mRNA expression and protein levels of COL1A2 and α-SMA by qRT-PCR (A) and Immunoblotting (B); the levels and distribution of α-SMA by Immunofluorescent (IF) staining (C); the cell viability by MTT (D). F HSCs were stimulated with 10 ng/ml TGF-β for 24 h and 10 µM of SB-431542 (a TGF-β inhibitor) for 12 h and the expression of miR-3074 in HSCs were detected by qRT-PCR. G The expression of MIR3074 in HSCs stimulated or non-stimulated with TGF-β, according to GSE151251. N = 3 (biological replicates). *p < 0.05; **p < 0.01 compared with 0 h (A, B, C, E) or or control group (D, F) or normal group (G). ##p < 0.01, compared with 24 h group (A, B, E) or TGF-β group (F)
miR-3074 overexpression aggravates TGF-β-caused fibrotic changes in HSCs
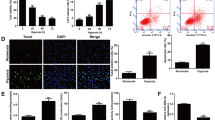
To investigate the specific functions of miR-3074 upon TGF-β-caused HSC activation, we transfected miR-3074 agomir/antagomir to achieve miR-3074 overexpression/inhibition in HSCs, as confirmed by qRT-PCR (Fig. 2A). For cell viability determination, HSCs were transfected with miR-3074 agomir/antagomir, treated with TGF-β (10 ng/ml) for 24 h. MTT assay was then carried out. In response to TGF-β treatment, miR-3074 overexpression facilitated TGF-β-induced HSC cell viability (Fig. 2B), while miR-3074 inhibition showed the opposite trends. Regarding the activation of HSCs and fibrotic markers, TGF-β-caused elevated COL1A2 and α-SMA expressions within HSCs showed to be further increased via miR-3074 overexpression while partially reduced via miR-3074 inhibition (Fig. 2C, D). Consistently, IF staining also showed that TGF-β-induced increase in fluorescent intensity that represented α-SMA level was promoted via miR-3074 overexpression while partially reversed via miR-3074 inhibition (Fig. 2E). Moreover, miR-3074-5p has been identified as a regulatory factor of cAMP responsive element binding protein (CREB) [37], hence, the effect of miR-3074-5p on cAMP signaling in HSCs was detected (Fig. S2). TGFβ treatment dramatically elevated PKA and CREB contents, whereas miR-3074 overexpression partially decreased TGFβ-induced increases in these proteins and miR-3074 inhibition further promoted PKA and CREB levels (Fig. S2). Thus, miR-3074 overexpression enhances TGF-β-induced HSCs activation.
miR-3074 overexpression aggravates TGF-β-induced fibrotic changes in HSCs. A miR-3074 overexpression or inhibition was achieved in HSCs by transfecting miR-3074 agomir or antagomir; miR-3074 expression was confirmed using qRT-PCR. B HSCs were transfected with miR-3074 agomir or antagomir, stimulated with 10 ng/ml TGF-β for 0, 24, or 48 h, and examined for cell viability by MTT assay. Then, HSCs were transfected with miR-3074 agomir or antagomir, stimulated with 10 ng/ml TGF-β for 48 h, and examined for the mRNA expression of COL1A2 and α-SMA by qRT-PCR (C); the protein levels of COL1A2 and α-SMA by Immunoblotting (D); the levels and distribution of α-SMA by Immunofluorescent (IF) staining (E). N = 3 (biological replicates). **p < 0.01 compared with agomir NC group. ##p < 0.01, compared with antagomir NC group
In vivo effects of miR-3074 overexpression and inhibition in liver fibrosis mouse model
The CCl4-caused liver fibrosis model in mouse was established, and mice were divided into the CCl4 + (lv-NC), CCl4 + (lv-miR-3074-5p), CCl4 + (lv-anti-NC), and CCl4 + (lv-anti-miR-3074-5p) groups; mice in each group received corresponding treatment and injection. H&E staining revealed that miR-3074 overexpression aggravated CCl4-induced structural disorder in mice liver; while miR-3074 inhibition improved liver damage of CCl4-induced mice (Fig. 3A). Masson’s staining showed miR-3074 overexpression aggravated sinusoidal fibrosis within the liver of CCl4-stimulated mice, whereas miR-3074 inhibition partially ameliorated sinusoidal fibrosis (Fig. 3B). Fibrotic area was calculated and Fig. 3C shows that the fibrotic area in CCl4-stimulated mice could be dramatically increased by miR-3074 overexpression, while reduced by miR-3074 inhibition. Regarding the miR-3074 expression in mice liver tissues, miR-3074 expression level showed to be significantly promoted by miR-3074 overexpression, whereas inhibited by miR-3074 inhibition (Fig. 3D).
In vivo effects of miR-3074 on CCl4-induced hepatic fibrosis in mouse. Mice were assigned into four groups: CCl4 + (lv-NC), CCl4 + (lv-miR-3074-5p), CCl4 + (lv-anti-NC), and CCl4 + (lv-anti-miR-3074-5p); mice in each group received corresponding treatment and injection. A Histopathological changes in mouse liver were assessed by H&E staining. B, C ECM deposition in mouse liver was examined by Masson’s staining. D The expression of miR-3074 in mouse liver from each group was determined by qRT-PCR. N = 8 (biological replicates). **p < 0.01 compared with CCl4 + (lv-NC) group. ##p < 0.01, compared with CCl4 + (lv-anti-NC) group
miR-3074 targets BMP7
miRNAs post-transcriptionally modulate gene expression via interaction with the 3'-UTR of target mRNAs and suppression of protein production through mRNA destabilization and translation repression [38]. Regarding the underlying mechanism, we employed miRDB, DIANA TOOLS, and TargetScan to analyze possible downstream targets of miR-3074; predicted targets with a Target Score ≥ 85 (by miRDB), miTG score ≥ 0.9 (by DIANA TOOLS), or Total context + + score ≤ -0.25 (by TargetScan) were selected and a total of 7 genes (CIT, PHACTR3, R3HDM1, FBXO3, JPH4, BMP7, and TRIB2) were enrolled (Fig. 4A, Table S2). As Fig. S3 revealed, overexpression of miR-3074 notably inhibited PHACTR3, R3HDM1, JPH4 and BMP7 mRNA expression, among which BMP7 had the lowest expression. Reportedly, BMP7 repressed liver fibrogenesis [39,40,41,42]; thus, BMP7 was selected for further experiments.
BMP7 is a direct downstream target of miR-3074. A A schematic diagram showing the process of selecting miR-3074 downstream targets; BMP7 was selected. B HSCs were transfected with miR-3074 agomir or antagomir, stimulated with 10 ng/ml TGF-β for 48 h, and examined for BMP7 protein levels by Immunoblotting. C Human wild- and mutant-type BMP7 luciferase reporter plasmids were constructed as described and co-transfected in 293 T cells and human HSCs with miR-3074 agomir/antagomir; the luciferase activity was determined. N = 3 (biological replicates). **p < 0.01 compared with agomir NC group. ##p < 0.01, compared with antagomir NC group
We transfected HSCs with miR-3074 agomir/antagomir, which was then treated with TGF-β (10 ng/ml) for 48 h; BMP7 protein contents were determined by immunoblotting; Fig. 4B shows that, in response to TGF-β induction, miR-3074 overexpression notably decreased BMP7 protein levels, while miR-3074 inhibition exhibited the opposite trends. Then, human wild-type and mutant-type BMP7 luciferase reporter plasmids were constructed, and then co-transfected with miR-3074 agomir/antagomir into 293 T cells and human HSCs; the luciferase activity was detected. As shown in Fig. 4C, when co-transfecting 293 T cells and human HSCs with human wt-BMP7 and agomir/antagomir, miR-3074 overexpression inhibited, whereas miR-3074 inhibition promoted the luciferase activity; when co-transfecting 293 T cells and human HSCs with human mut-BMP7 and agomir/antagomir, miR-3074 overexpression or inhibition did not change the luciferase activity. In summary, miR-3074 directly targets BMP7 and inhibits BMP7 expression.
miR-3074 modulates TGF-β-induced fibrotic changes through BMP7 in HSCs
After confirming the direct binding between miR-3074 and BMP7, next, we investigated miR-3074/BMP7 effects on TGF-β-caused HSC activation. We transfected Lv-BMP7 to achieve BMP7 overexpression within HSCs, as confirmed by Immunoblotting and qRT-PCR (Fig. 5A, B). Then, HSCs were co-transfected with miR-3074 agomir and Lv-BMP7, treated with TGF-β (10 ng/ml) for 24 h; HSC activation was measured. In TGF-β-treated HSCs, miR-3074 overexpression further promoted cell viability, whereas BMP7 overexpression partially inhibited cell viability, and BMP7 overexpression could remarkably reversed overexpressed miR-3074 effects on HSCs (Fig. 5C). Upon TGF-β treatment, miR-3074 overexpression decreased BMP7 mRNA and protein expression but increased COL1A2 and α-SMA, whereas BMP7 overexpression increased BMP7 but decreased COL1A2 and α-SMA; BMP7 overexpression significantly reversed the effects of miR-3074 overexpression on these factors (Fig. 5D, E). Moreover, after TGF-β treatment, miR-3074 overexpression inhibited the phosphorylation of Smad1/5/8 but promoted the phosphorylation of Smad2 and Smad3, whereas BMP7 overexpression exerted opposite effects on Smad1/5/8, Smad2 and Smad3 phosphorylation (Fig. 5E); the effects of miR-3074 overexpression on Smads phosphorylation could be partially attenuated by BMP7 overexpression (Fig. 5E). As further evidence of TGF-β-induced fibrotic changes, miR-3074 overexpression elevated the levels of α-SMA under TGF-β stimulation, whereas BMP7 overexpression exerted opposite effects on α-SMA levels (Fig. 5F); similarly, BMP7 overexpression significantly attenuated the impacts of overexpressed miR-3074 upon α-SMA levels (Fig. 5F).
Overexpression of miR-3074 modulates TGF-β-induced fibrotic changes through BMP7 in HSCs. A, B BMP7 overexpression was achieved in HSCs by transfecting Lv-BMP7; BMP7 overexpression was confirmed using Immunoblotting and qRT-PCR. Then, HSCs were co-transfected with miR-3074 agomir and Lv-BMP7, stimulated with 10 ng/ml TGF-β for 24 h, and examined for C cell viability by MTT assay; D mRNA expression of BMP7, COL1A2, and α-SMA by qRT-PCR; E the protein levels of BMP7, p-Smad1/5/8, Smad1/5/8, p-Smad2, Smad2, p-Smad3, Smad3, COL1A2, and α-SMA by Immunoblotting; F the levels and distribution of α-SMA by IF staining. N = 3 (biological replicates). *p < 0.05, **p < 0.01 compared with Lv-NC group (A, B) or agomir NC + Lv-NC group (C–E). #p < 0.05, ##p < 0.01, compared with miR-3074-5p agomir + Lv-BMP7 group
Moreover, two shRNA of BMP7 vectors (sh-BMP7#1 and sh-BMP7#2) were transfected into HSCs to achieve BMP7 silencing, as confirmed by Immunoblotting and qRT-PCR (Fig. 6A, B). Then, HSCs were co-transfected with miR-3074 antagomir and sh-BMP7#2, treated with TGF-β (10 ng/ml) for 24 h; HSC activation was measured. In TGF-β-treated HSCs, miR-3074 silencing inhibited cell viability, whereas BMP7 silencing promoted cell viability (Fig. 6C). Upon TGF-β treatment, miR-3074 knockdown increased BMP7 mRNA and protein expression but decreased COL1A2 and α-SMA, whereas BMP7 overexpression increased BMP7 but decreased COL1A2 and α-SMA (Fig. 6D, E). Moreover, after TGF-β treatment, miR-3074 knockdown promoted the phosphorylation of Smad1/5/8 but inhibited the phosphorylation of Smad2 and Smad3, whereas BMP7 knockdown exerted opposite effects on Smad1/5/8, Smad2 and Smad3 phosphorylation (Fig. 6E). As further evidence of TGF-β-induced fibrotic changes, miR-3074 silencing inhibited the levels of α-SMA under TGF-β stimulation, whereas BMP7 silencing exerted opposite effects on α-SMA levels (Fig. 6F); BMP7 silencing significantly attenuated the impacts of overexpressed miR-3074 upon α-SMA levels (Fig. 6F).
Silencing of miR-3074 modulates TGF-β-induced fibrotic changes through BMP7 in HSCs. A, B BMP7 knockdown was achieved in HSCs by transfecting with two shRNA of BMP7 vectors (sh-BMP7#1 and sh-BMP7#2); BMP7 silencing was confirmed using Immunoblotting and qRT-PCR. Then, HSCs were co-transfected with miR-3074 antagomir and sh-BMP7#2, stimulated with 10 ng/ml TGF-β for 24 h, and examined for C cell viability by MTT assay; D mRNA expression of BMP7, COL1A2, and α-SMA by qRT-PCR; E the protein levels of BMP7, p-Smad1/5/8, Smad1/5/8, p-Smad2, Smad2, p-Smad3, Smad3, COL1A2, and α-SMA by Immunoblotting; F the levels and distribution of α-SMA by IF staining. N = 3 (biological replicates). **p < 0.01 compared with sh-NC group (A, B) or antagomir NC + sh-NC group (C–E). #p < 0.05, ##p < 0.01, compared with miR-3074-5p antagomir + sh-BMP7 group
In vivo influence of miR-3074/BMP7 in liver fibrosis mouse model
Finally, we established CCl4-caused liver fibrosis model in mouse, and investigated the dynamic effects of the miR-3074/BMP7 axis. Mice were divided into the control, CCl4 + (Lv-NC), CCl4 + (Lv-miR-3074-5p), and CCl4 + (miR-3074-5p + Lv-BMP7) groups; mice in each group received corresponding treatment and injection. H&E and Masson’s staining demonstrated inflammatory cell recruitment with related collagen fibers all over hepatic parenchyma of mice in the CCl4 + (Lv-NC) group, whereas LV-miR-3074-5p injection further enhanced hepatic injuries (Fig. 7A, B). On the contrary, LV-BMP7 injection improved inflammation and fibrosis as compared to mice in the CCl4 + (Lv-NC) and CCl4 + (Lv-miR-3074-5p) groups (Fig. 7A, B). CCl4 stimulation reduced BMP7 levels, and Lv-miR-3074-5p injection further decreased BMP7 levels; in the CCl4 + (Lv-miR-3074-5p + Lv-BMP7) group, LV-BMP7 injection partially increased BMP7 levels (Fig. 7C). Contrariwise, CCl4 stimulation induced miR-3074 upregulation, and Lv-miR-3074-5p injection further upregulated miR-3074 expression; Lv-BMP7 injection caused no changes in miR-3074 expression (Fig. 7D). Lastly, wild-type and mutant-type mouse BMP7 luciferase reporter plasmids were constructed as described, which was then co-transfected with mmu-miR-3074 agomir/antagomir into 293T cells, followed by the detection of luciferase activity. Consistent with earlier experiments, when co-transfecting 293T cells with wt-BMP7 (mouse) and mmu-miR-3074 agomir/antagomir, mmu-miR-3074 overexpression inhibited, whereas mmu-miR-3074 inhibition promoted the luciferase activity; when co-transfecting 293T cells with mut-BMP7 (mouse) and mmu-miR-3074 agomir/antagomir, mmu-miR-3074 overexpression or inhibition caused no changes in the luciferase activity (Fig. 7E). Moreover, mouse wild-type and mutant-type BMP7 3’UTR luciferase reporter plasmids were constructed, and then co-transfected with human miR-3074 agomir/antagomir into mHSCs and the luciferase activity was detected. As indicated in Fig. S4A, when co-transfected with mouse wt-BMP7, the luciferase activity was inhibited by miR-3074 overexpression, while promoted by miR-3074 inhibition; in cells co-transfected with mouse mut-BMP7, miR-3074 overexpression or inhibition did not alter the luciferase activity (Fig. S4A). In mHSCs, human source miR-3074 overexpression markedly inhibited BMP7 expression, while miR-3074 inhibition notably promoted BMP7 expression (Fig. S4B). In summary, mmu-miR-3074 directly targets mouse BMP7.
In vivo effects of the miR-3074/BMP7 axis on CCl4-induced hepatic fibrosis mouse model Mice were divided into four groups: control, CCl4 + (Lv-NC), CCl4 + (Lv-miR-3074-5p), and CCl4 + (Lv-miR-3074-5p + LV-BMP7); mice in each group received corresponding treatment and injection. A Histopathological changes in mouse liver were assessed by H&E staining. B ECM deposition in mouse liver was examined by Masson’s staining. C The levels and distribution of BMP7 were determined by Immunohistochemical (IHC) staining. D The expression of miR-3074 in mouse liver from each group was determined by qRT-PCR. N = 8 (biological replicates). E Wild- and mutant-type mouse BMP7 luciferase reporter plasmids were constructed as described and co-transfected in 293T cells with mmu-miR-3074 agomir/antagomir; the luciferase activity was determined. N = 3 (biological replicates). **p < 0.01 compared with control or agomir NC group. ##p < 0.01, compared with CCl4 + (Lv-NC) or antagomir NC group. &&p < 0.01, compared to CCl4 + (Lv-miR-3074-5p) group
Discussion
In the present study, miR-3074 was found increased within TGF-β-activated HSCs and enriched within the TGF-β signaling. In activated HSCs by TGF-β, miR-3074 overexpression aggravated TGF-β-induced fibrotic changes, whereas miR-3074 inhibition exerted opposite effects. miR-3074 directly targeted BMP7 and inhibited BMP7 expression. Under TGF-β induction, overexpressed BMP7 notably attenuated the promotive roles of miR-3074 overexpression in TGF-β-activated HSCs. Within CCl4-caused liver fibrosis murine model, miR-3074 overexpression administration promoted, while LV-BMP7 administration alleviated CCl4-induced fibrotic changes; LV-BMP7 significantly attenuated the effects of miR-3074. Lastly, mmu-miR-3074 also targeted mouse BMP7 and inhibited mouse BMP7 expression.
miRNAs modulate biological behaviors and affect gene expression in multiple organs, including liver. Within liver fibrogenesis, miR-146a [18], miR-454 [43] and miR-200a [44] exhibit negative correlation with SMAD4 and SMAD3, separately. Moreover, miR-199a [17], miR-33a [19], and miR-21 [20] exhibit positive correlation with SMAD3 and SMAD7, separately. Through targeting different Smads, miRNAs play crucial roles in hepatic fibrosis. Previously, we have demonstrated the binding between miR-503 and Smad7 and miR-503's pro-fibrotic roles in TGF-β-activated HSCs [21]. Herein, comprehensive bioinformatics analyses indicated miR-3074 upregulation within TGF-β-induced HSCs. Moreover miR-3074 was enriched in the TGF-β signaling pathway according to DIANA-mirPath analysis. miR-3074-5p has been recognized as a regulatory factor of carcinogenic cAMP responsive element binding protein [45]. In a previous study, miR-3074-5p is linked to a satisfactory prognosis of papillary renal cell carcinoma patients, which might act mainly to suppress oligodendrocyte precursor cell proliferative ability and neuronal differentiation [46]; the effects of miR-3074 on hepatic fibrosis have never been examined. Herein, TGF-β-induced upregulation of miR-3074 suggests that miR-3074 might participate in TGF-β-induced HSC activation.
In the normal liver, HSCs retain the quiescent non-proliferative state as monitored by star-shaped morphology. Vitamin A is stored as retinyl esters in the lipid droplets in HSC cytoplasm [47]. In the process of acute and chronic hepatic damage, TGF-β shows to be activated from ECM deposits and expressed and produced from a variety of cellular types, inducing HSC activation and trans-differentiation to myofibroblasts, consisting of intracellular vitamin A droplet loss, fibroblast shape adaption, and development of phenotype with the characteristics of contractility, proliferation, and migration [47]. Consistently, herein, TGF-β markedly enhanced the viability of HSC cells and upregulated α-SMA and COL1A2 proteins, which indicated that TGF-β treatment caused HSC activation. More importantly, overexpression of miR-3074 can further enhance TGF-β-caused HSC activation, whereas miR-3074 inhibition exhibited opposite effects, as evidenced by the inhibited cell viability and decreased α-SMA and COL1A2 levels. In summary, miR-3074 overexpression promotes activation of TGF-β-treated HSCs.
As aforementioned, miRNAs modulate gene expression via interacting with target mRNAs through mRNA destabilization or translational repression [15, 16]. In the present study, BMP7, belonging to the TGF-β superfamily and signal via SMAD1/5/8 [48, 49], is a direct downstream target of miR-3074. Interestingly, exogenous BMP7 inhibited HSC activation, migratory ability, and proliferative ability and exerted anti-hepatic fibrosis effects through selectively regulating TGF-β/Smad signaling-related factors [39]. In the process of hepatic fibrogenesis, BMP7 inhibited fibrosis via activating Smad1/5/8 complexes [10, 50]. Herein, considering that miR-3074 is triggered via TGF-β treatment and miR-3074 inhibits BMP7 expression, the miR-3074/BMP7 axis might serve a dynamic part in the activation of TGF-β-stimulated HSCs. As expected, within TGF-β-stimulated HSCs, BMP7 overexpression inhibited HSC viability and decreased α-SMA and COL1A2 levels; in the meantime, BMP7 overexpression promoted SMAD1/5/8 phosphorylation but hindered Smad2 phosphorylation. More importantly, in TGF-β-stimulated HSCs, BMP7 overexpression notably reversed miR-3074 overexpression impacts upon activation of TGF-β-treated HSCs, as well as the Smads phosphorylation. Thus, within TGF-β-treated HSCs, the miR-3074/BMP7 axis modulates TGF-β-caused activation of HSCs, possibly via BMP7 activating Smad1/5/8.
BMP7 helps to suppress the effects of fibrogenesis induced by TGF-β. Within a CCl4-caused hepatic fibrosis murine model, BMP7 overexpression in the liver via tail injection suppressed hepatic fibrogenesis [51]. Within a CCl4-caused hepatic fibrosis murine model, hepatic fibrosis could also be ameliorated by the oral administration of recombinant BMP7, which could be related to influencing TGF-β/Smad signaling [42]. It has shown that BMP7 modulates liver fibrosis through the TGF-β/Smad signaling in hepatic fibrosis model. Herein, we established CCl4-caused hepatic fibrosis murine model; fibrotic changes and liver injuries were observed in mouse liver. Consistent with the in vitro results, miR-3074 overexpression aggravated, while BMP7 overexpression alleviated the fibrotic changes and liver injuries; BMP7 overexpression partially reversed the effects of miR-3074 overexpression. In mouse, mmu-miR-3074 also targets mouse BMP7 and inhibits BMP7 expression.
In conclusion, the miR-3074/BMP7 axis regulates TGF-β-caused activation of HSCs in vitro and CCl4-caused murine liver fibrosis in vivo. BMP7-mediated Smad1/5/8 activation might be involved.
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper.
References
Jiao J, Friedman SL, Aloman C. Hepatic fibrosis. Curr Opin Gastroenterol. 2009;25(3):223–9.
Bataller R, Brenner DA. Liver fibrosis. J Clin Investig. 2005;115(2):209–18.
Ellis EL, Mann DA. Clinical evidence for the regression of liver fibrosis. J Hepatol. 2012;56(5):1171–80.
Seki E, Schwabe RF. Hepatic inflammation and fibrosis: functional links and key pathways. Hepatology. 2015;61(3):1066–79.
Mederacke I, et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat Commun. 2013;4:2823.
Puche JE, Saiman Y, Friedman SL. Hepatic stellate cells and liver fibrosis. Compr Physiol. 2013;3(4):1473–92.
Lee YA, Wallace MC, Friedman SL. Pathobiology of liver fibrosis: a translational success story. Gut. 2015;64(5):830–41.
Massague J, Wotton D. Transcriptional control by the TGF-beta/Smad signaling system. EMBO J. 2000;19(8):1745–54.
Friedman SL. Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev. 2008;88(1):125–72.
Xu F, et al. TGF-beta/SMAD pathway and its regulation in hepatic fibrosis. J Histochem Cytochem. 2016;64(3):157–67.
Friedman SL. Mechanisms of hepatic fibrogenesis. Gastroenterology. 2008;134(6):1655–69.
Lee UE, Friedman SL. Mechanisms of hepatic fibrogenesis. Best Pract Res Clin Gastroenterol. 2011;25(2):195–206.
Gressner AM, Weiskirchen R. Modern pathogenetic concepts of liver fibrosis suggest stellate cells and TGF-beta as major players and therapeutic targets. J Cell Mol Med. 2006;10(1):76–99.
Zhang L, et al. Smad2 protects against TGF-beta1/Smad3-mediated collagen synthesis in human hepatic stellate cells during hepatic fibrosis. Mol Cell Biochem. 2015;400(1–2):17–28.
Szabo G, Bala S. MicroRNAs in liver disease. Nat Rev Gastroenterol Hepatol. 2013;10(9):542–52.
Eulalio A, Huntzinger E, Izaurralde E. Getting to the root of miRNA-mediated gene silencing. Cell. 2008;132(1):9–14.
Murakami Y, et al. The progression of liver fibrosis is related with overexpression of the miR-199 and 200 families. PLoS ONE. 2011;6(1):e16081.
He Y, et al. MicroRNA-146a modulates TGF-beta1-induced hepatic stellate cell proliferation by targeting SMAD4. Cell Signal. 2012;24(10):1923–30.
Huang CF, et al. miR-33a levels in hepatic and serum after chronic HBV-induced fibrosis. J Gastroenterol. 2015;50(4):480–90.
Marquez RT, et al. Correlation between microRNA expression levels and clinical parameters associated with chronic hepatitis C viral infection in humans. Lab Investig. 2010;90(12):1727–36.
Xie X, et al. MicroRNA-503 targets mothers against decapentaplegic homolog 7 enhancing hepatic stellate cell activation and hepatic fibrosis. Dig Dis Sci. 2020;66:1928–39.
Kang H, et al. TGF-beta activates NLRP3 inflammasome by an autocrine production of TGF-beta in LX-2 human hepatic stellate cells. Mol Cell Biochem. 2022;477(5):1329–38.
Wang Q, et al. MicroRNA-101 inhibits renal tubular epithelial-to-mesenchymal transition by targeting TGF-β1 type I receptor. Int J Mol Med. 2021. https://doi.org/10.3892/ijmm.2021.4952.
Ge S, et al. Role of growth factor receptor-bound 2 in CCl4-induced hepatic fibrosis. Biomed Pharmacother. 2017;92:942–51.
Ge S, et al. MicroRNA-146b regulates hepatic stellate cell activation via targeting of KLF4. Ann Hepatol. 2016;15(6):918–28.
He H, et al. FBXO31 modulates activation of hepatic stellate cells and liver fibrogenesis by promoting ubiquitination of Smad7. J Cell Biochem. 2019;121:3711–9.
Hao Y, et al. TRIM27-mediated ubiquitination of PPARγ promotes glutamate-induced cell apoptosis and inflammation. Exp Cell Res. 2021;400(1):112437.
Yang JJ, et al. MeCP2 silencing of LncRNA H19 controls hepatic stellate cell proliferation by targeting IGF1R. Toxicology. 2016;359–360:39–46.
Jung HJ, et al. Ethanol extract of Pharbitis nil ameliorates liver fibrosis through regulation of the TGFbeta1-SMAD2/3 pathway. J Ethnopharmacol. 2022;294:115370.
Ma L, et al. MicroRNA-214 promotes hepatic stellate cell activation and liver fibrosis by suppressing Sufu expression. Cell Death Dis. 2018;9(7):718.
Caviglia JM, et al. MicroRNA-21 and Dicer are dispensable for hepatic stellate cell activation and the development of liver fibrosis. Hepatology. 2018;67(6):2414–29.
Men R, et al. MircoRNA-145 promotes activation of hepatic stellate cells via targeting kruppel-like factor 4. Sci Rep. 2017;7:40468.
Tu H, et al. microRNA-143-3p attenuated development of hepatic fibrosis in autoimmune hepatitis through regulation of TAK1 phosphorylation. J Cell Mol Med. 2020;24(2):1256–67.
Yang X, et al. Twist1-induced miR-199a-3p promotes liver fibrosis by suppressing caveolin-2 and activating TGF-beta pathway. Signal Transduct Target Ther. 2020;5(1):75.
Ji D, et al. MiR-22 suppresses BMP7 in the development of cirrhosis. Cell Physiol Biochem. 2015;36(3):1026–36.
Huang W, et al. LncRNA Neat1 expedites the progression of liver fibrosis in mice through targeting miR-148a-3p and miR-22-3p to upregulate Cyth3. Cell Cycle. 2021;20(5–6):490–507.
Wang J, et al. lncRNA Eif4g2 improves palmitate-induced dysfunction of mouse β-cells via modulation of Nrf2 activation. Exp Cell Res. 2020;396(2):112291.
Cannell IG, Kong YW, Bushell M. How do microRNAs regulate gene expression? Biochem Soc Trans. 2008;36(Pt 6):1224–31.
Zou GL, et al. Bone morphogenetic protein-7 represses hepatic stellate cell activation and liver fibrosis via regulation of TGF-beta/Smad signaling pathway. World J Gastroenterol. 2019;25(30):4222–34.
Chen L, et al. rSjp40 inhibits activated hepatic stellate cells by promoting nuclear translocation of YB1 and inducing BMP-7/Smad1/5/8 pathway. Parasit Vectors. 2019;12(1):279.
Wang SL, et al. Inhibitory effect of bone morphogenetic protein-7 on hepatic fibrosis in rats. Int J Clin Exp Pathol. 2013;6(5):897–903.
Hao ZM, et al. Oral administration of recombinant adeno-associated virus-mediated bone morphogenetic protein-7 suppresses CCl(4)-induced hepatic fibrosis in mice. Mol Ther. 2012;20(11):2043–51.
Zhu D, et al. Expression of microRNA-454 in TGF-beta1-stimulated hepatic stellate cells and in mouse livers infected with Schistosoma japonicum. Parasit Vectors. 2014;7:148.
Wang B, et al. miR-200a prevents renal fibrogenesis through repression of TGF-beta2 expression. Diabetes. 2011;60(1):280–7.
Wang J, et al. lncRNA Eif4g2 improves palmitate-induced dysfunction of mouse beta-cells via modulation of Nrf2 activation. Exp Cell Res. 2020;396(2):112291.
Yang J, et al. Oligodendrocyte precursor cell transplantation promotes functional recovery following contusive spinal cord injury in rats and is associated with altered microRNA expression. Mol Med Rep. 2018;17(1):771–82.
Tsuchida T, Friedman SL. Mechanisms of hepatic stellate cell activation. Nat Rev Gastroenterol Hepatol. 2017;14(7):397–411.
Derynck R, Budi EH. Specificity, versatility, and control of TGF-beta family signaling. Sci Signal. 2019. https://doi.org/10.1126/scisignal.aav5183.
Herrera B, Addante A, Sanchez A. BMP signalling at the crossroad of liver fibrosis and regeneration. Int J Mol Sci. 2017;19(1):39.
Guo J, et al. BMP7 suppresses excessive scar formation by activating the BMP7/Smad1/5/8 signaling pathway. Mol Med Rep. 2017;16(2):1957–63.
Kinoshita K, et al. Adenovirus-mediated expression of BMP-7 suppresses the development of liver fibrosis in rats. Gut. 2007;56(5):706–14.
Acknowledgements
Not applicable.
Funding
This study was supported by the fund of Hunan Provincial Health Commission (202103080306 and 202203082548).
Author information
Authors and Affiliations
Contributions
LJ, L-BJ and XX designed the research and wrote the manuscript. L-BJ, XX and YX performed the majority of experiments and assisted with the animal experiments. D-CY and T-HB performed the figures and the statistical analyses. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Consent for publication
All authors agree to publication.
Ethics approval and consent to participate
All animal experiments were performed in accordance with the guidelines of Health Guide for the Care and Use of Laboratory Animals, and the experimental protocols were approved by the Institutional Animal Care and Use Committee of First Affiliated Hospital of University of South China.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2023_1017_MOESM1_ESM.tif
The selection of miRNAs involved in TGF-β-caused HSC activation. A Based on the online dataset GSE151251, RNAseq analysis detected a total of 1968 deregulated miRNAs in HSCs stimulated with TGF-β. After standardization and homogenization using DEseq2 package in R language, a total of 563 deregulated miRNAs were obtained, 17 of which were downregulated and 20 upregulated with the∣ logFc∣ > 0.2, P < 0.05. B The heatmap showed the top 15 up- and downregulated miRNAs. C Based on mirPath V3 (containing the data from TargetScan, Tarbase V7.0, and MicroT-CDS5.0) were used to screen for the miRNAs enriched in the TGF-β pathway. 217 miRNAs were associated with the TGF-β signaling in all three sets of data. D Compared with deregulated miRNAs reported in GSE151251, seven miRNA host genes (MIR22, MIR3074, MIR199a, MIR145, MIR143, MIR21, and MIR214) were found to be enriched in the TGF-β pathway. Among the 7 miRNAs, miR-3074 has not been reported that associated with liver fibrosis (TIF 1207 KB)
13577_2023_1017_MOESM2_ESM.tif
The effect of miR-3074-5p on cAMP signaling in HSCs. Human HSCs were transfected with miR-3074 agomir or antagomir, stimulated with 10 ng/ml TGF-β for 48 h, and examined for PKA and CREB protein levels by Immunoblotting. N = 3 (biological replicates). *p < 0.05, **p < 0.01 (TIF 764 KB)
13577_2023_1017_MOESM3_ESM.tif
The effect of miR-3074 agomir on the expression levels of 7 potential target genes of miR-3074 HSCs were transfected with miR-3074 agomir, stimulated with 10 ng/ml TGF-β for 48 h, and examined for CIT, PHACTR3, R3HDM1, FBXO3, JPH4, BMP7, and TRIB2 mRNA levels by qRT-PCR. N = 3 (biological replicates). **p < 0.01 compared with agomir NC group (TIF 207 KB)
13577_2023_1017_MOESM4_ESM.tif
Mmu-BMP7 is a direct downstream target of miR-3074. A Mouse wild- and mutant-type BMP7 luciferase reporter plasmids were constructed and co-transfected into mHSCs cells transfected with human source miR-3074 agomir/antagomir; the luciferase activity was determined. B mHSCs were transfected with human source miR-3074 agomir or antagomir, and examined for BMP7 mRNA levels by qRT-PCR. N = 3 (biological replicates). **p < 0.01 compared with agomir NC group. ##p < 0.01, compared with antagomir NC group (TIF 641 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, B., Xie, X., Yang, X. et al. The miR-3074/BMP7 axis regulates TGF-β-caused activation of hepatic stellate cells in vitro and CCl4-caused murine liver fibrosis in vivo. Human Cell 37, 435–450 (2024). https://doi.org/10.1007/s13577-023-01017-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-023-01017-y