Abstract
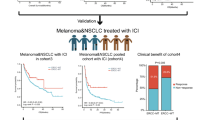
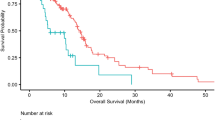
Glutamate-NMDAR receptors (GRINs) have been reported to influence cancer immunogenicity; however, the relationship between GRIN alterations and the response to immune checkpoint inhibitors (ICIs) has not been determined. This study combined clinical characteristics and mutational profiles from multiple cohorts to form a discovery cohort (n = 901). The aim of this study was to investigate the correlation between the mutation status of the GRIN gene and the response to ICI therapy. Additionally, an independent ICI-treated cohort from the Memorial Sloan Kettering Cancer Center (MSKCC, N = 1513) was used for validation. Furthermore, this study explored the associations between GRIN2A mutations and intrinsic and extrinsic immunity using multiomics analysis. In the discovery cohort, patients with GRIN2A-MUTs had improved clinical outcomes, as indicated by a higher objective response rate (ORR: 36.8% vs 25.8%, P = 0.020), durable clinical benefit (DCB: 55.2% vs 38.7%, P = 0.005), prolonged progression-free survival (PFS: HR = 0.65; 95% CI 0.49 to 0.87; P = 0.003), and increased overall survival (OS: HR = 0.67; 95% CI 0.50 to 0.89; P = 0.006). Similar results were observed in the validation cohort, in which GRIN2A-MUT patients exhibited a significant improvement in overall survival (HR = 0.66; 95% CI = 0.49 to 0.88; P = 0.005; adjusted P = 0.045). Moreover, patients with GRIN2A-MUTs exhibited an increase in tumor mutational burden, high expression of costimulatory molecules, increased activity of antigen-processing machinery, and infiltration of various immune cells. Additionally, gene sets associated with cell cycle regulation and the interferon response were enriched in GRIN2A-mutated tumors. In conclusion, GRIN2A mutation is a novel biomarker associated with a favorable response to ICIs in multiple cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The clinicopathological data of patients in the cohorts or patients treated with immune checkpoint agents are described in the Materials and Methods and Supporting Data. The resources, tools and codes used in our analyses are described in the “Methods” section. Other data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Havel JJ, Chowell D, Chan TA. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer. 2019;19:133–50.
Gandhi L, Rodriguez-Abreu D, Gadgeel S, Esteban E, Felip E, De Angelis F, et al. Pembrolizumab plus chemotherapy in metastatic non-small cell lung cancer. N Engl J Med. 2018;378:2078–92.
Sharma P, Hu-Lieskovan S, Wargo JA, Ribas A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell. 2017;168:707–23.
Grant MJ, Herbst RS, Goldberg SB. Selecting the optimal immunotherapy regimen in driver-negative metastatic NSCLC. Nat Rev Clin Oncol. 2021;18:625–44.
Davoli T, Uno H, Wooten EC, Elledge SJ. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science. 2017;355:eaaf8399.
Wu HX, Chen YX, Wang ZX, Zhao Q, He MM, Wang YN, et al. Alteration in TET1 as potential biomarker for immune checkpoint blockade in multiple cancers. J Immunother Cancer. 2019;7:264.
Ott PA, Bang YJ, Piha-Paul SA, Razak ARA, Bennouna J, Soria JC, et al. T-cell-inflamed gene-expression profile, programmed death ligand 1 expression, and tumor mutational burden predict efficacy in patients treated with pembrolizumab across 20 cancers: KEYNOTE-028. J Clin Oncol. 2019;37:318–27.
Hakimi AA, Voss MH, Kuo F, Sanchez A, Liu M, Nixon BG, et al. Transcriptomic profiling of the tumor microenvironment reveals distinct subgroups of clear cell renal cell cancer: data from a randomized phase III trial. Cancer Discov. 2019;9:510–25.
Zhou H, Liu J, Zhang Y, Huang Y, Shen J, Yang Y, et al. PBRM1 mutation and preliminary response to immune checkpoint blockade treatment in non-small cell lung cancer. NPJ Precision Oncol. 2020;4:6.
Gonzalez-Cao M, Viteri S, Karachaliou N, Aguilar A, Garcia-Mosquera JJ, Rosell R. Tumor mutational burden as predictive factor of response to immunotherapy. Trans Lung Cancer Res. 2018;7:S358–61.
Ready N, Hellmann MD, Awad MM, Otterson GA, Gutierrez M, Gainor JF, et al. First-line nivolumab plus ipilimumab in advanced non-small cell lung cancer (CheckMate 568): outcomes by programmed death ligand 1 and tumor mutational burden as biomarkers. J Clin Oncol. 2019;37:992–1000.
Prickett TD, Gartner JJ, Samuels Y. Genetic and functional analysis of GRIN2A in tumor samples. Methods Mol Biol. 2017;1677:93–116.
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev. 2010;62:405–96.
Lang UE, Puls I, Muller DJ, Strutz-Seebohm N, Gallinat J. Molecular mechanisms of schizophrenia. Cell Physiol Biochem. 2007;20:687–702.
Prickett TD, Zerlanko BJ, Hill VK, Gartner JJ, Qutob N, Jiang J, et al. Somatic mutation of GRIN2A in malignant melanoma results in loss of tumor suppressor activity via aberrant NMDAR complex formation. J Investig Dermatol. 2014;134:2390–8.
Wei X, Walia V, Lin JC, Teer JK, Prickett TD, Gartner J, et al. Exome sequencing identifies GRIN2A as frequently mutated in melanoma. Nat Genet. 2011;43:442–6.
Carvill GL, Regan BM, Yendle SC, O’Roak BJ, Lozovaya N, Bruneau N, et al. GRIN2A mutations cause epilepsy-aphasia spectrum disorders. Nat Genet. 2013;45:1073–6.
Lemke JR, Lal D, Reinthaler EM, Steiner I, Nothnagel M, Alber M, et al. Mutations in GRIN2A cause idiopathic focal epilepsy with rolandic spikes. Nat Genet. 2013;45:1067–72.
Wu WC, Sun HW, Chen J, OuYang HY, Yu XJ, Chen HT, et al. Immunosuppressive immature myeloid cell generation is controlled by glutamine metabolism in human cancer. Cancer Immunol Res. 2019;7:1605–18.
Gualandris A, Noghero A, Geuna M, Arese M, Valdembri D, Serini G, et al. Microenvironment drives the endothelial or neural fate of differentiating embryonic stem cells coexpressing neuropilin-1 and Flk-1. FASEB J. 2009;23:68–78.
Ferguson HJ, Wragg JW, Ward S, Heath VL, Ismail T, Bicknell R. Glutamate dependent NMDA receptor 2D is a novel angiogenic tumor endothelial marker in colorectal cancer. Oncotarget. 2016;7:20440–54.
Bagaev A, Kotlov N, Nomie K, Svekolkin V, Gafurov A, Isaeva O, et al. Conserved pancancer microenvironment subtypes predict response to immunotherapy. Cancer Cell. 2021;39:845–65.e7.
Taylor MH, Lee CH, Makker V, Rasco D, Dutcus CE, Wu J, et al. Phase IB/II trial of lenvatinib plus pembrolizumab in patients with advanced renal cell carcinoma, endometrial cancer, and other selected advanced solid tumors. J Clin Oncol. 2020;38:1154–63.
Finn RS, Ikeda M, Zhu AX, Sung MW, Baron AD, Kudo M, et al. Phase Ib study of lenvatinib plus pembrolizumab in patients with unresectable hepatocellular carcinoma. J Clin Oncol. 2020;38:2960–70.
Huang L, Luo S, Zhang X, Cai Y, Xue F, Hu H, et al. Distinct genomic landscape of colorectal mucinous carcinoma determined via comprehensive genomic profiling: steps to a new treatment strategy. Front Oncol. 2021;11:603564.
Qiu Y, Liu L, Yang H, Chen H, Deng Q, Xiao D, et al. Integrating histologic and genomic characteristics to predict tumor mutation burden of early-stage non-small cell lung cancer. Front Oncol. 2020;10:608989.
Braun DA, Hou Y, Bakouny Z, Ficial M, Sant’ Angelo M, Forman J, et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat Med. 2020;26:909–18.
Liu D, Schilling B, Liu D, Sucker A, Livingstone E, Jerby-Arnon L, et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat Med. 2019;25:1916–27.
Miao D, Margolis CA, Vokes NI, Liu D, Taylor-Weiner A, Wankowicz SM, et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat Genet. 2018;50:1271–81.
Hellmann MD, Nathanson T, Rizvi H, Creelan BC, Sanchez-Vega F, Ahuja A, et al. Genomic features of response to combination immunotherapy in patients with advanced non-small cell lung cancer. Cancer Cell. 2018;33:843–52.e4.
Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L, et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science. 2015;350:207–11.
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371:2189–99.
Long J, Wang D, Yang X, Wang A, Lin Y, Zheng M, et al. Identification of NOTCH4 mutation as a response biomarker for immune checkpoint inhibitor therapy. BMC Med. 2021;19:154.
Samstein RM, Lee CH, Shoushtari AN, Hellmann MD, Shen R, Janjigian YY, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51:202–6.
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, et al. The immune landscape of cancer. Immunity. 2019;51:411–2.
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell. 2016;165:35–44.
Zehir A, Benayed R, Shah RH, Syed A, Middha S, Kim HR, et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat Med. 2017;23:703–13.
Liu J, Lichtenberg T, Hoadley KA, Poisson LM, Lazar AJ, Cherniack AD, et al. An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell. 2018;173:400–16.e11.
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–7.
Reimand J, Isserlin R, Voisin V, Kucera M, Tannus-Lopes C, Rostamianfar A, et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat Protoc. 2019;14:482–517.
Gao J, Chang MT, Johnsen HC, Gao SP, Sylvester BE, Sumer SO, et al. 3D clusters of somatic mutations in cancer reveal numerous rare mutations as functional targets. Genome Med. 2017;9:4.
Garrido F, Aptsiauri N. Cancer immune escape: MHC expression in primary tumors versus metastases. Immunology. 2019;158:255–66.
Spranger S. Mechanisms of tumor escape in the context of the T-cell-inflamed and the non-T-cell-inflamed tumor microenvironment. Int Immunol. 2016;28:383–91.
Liu H, Yang Z, Lu W, Chen Z, Chen L, Han S, et al. Chemokines and chemokine receptors: a new strategy for breast cancer therapy. Cancer Med. 2020;9:3786–99.
Bergamaschi C, Pandit H, Nagy BA, Stellas D, Jensen SM, Bear J, et al. Heterodimeric IL-15 delays tumor growth and promotes intratumoral CTL and dendritic cell accumulation by a cytokine network involving XCL1, IFN-gamma, CXCL9 and CXCL10. J Immunotherapy Cancer. 2020;8:e000599.
Jia Q, Wu W, Wang Y, Alexander PB, Sun C, Gong Z, et al. Local mutational diversity drives intratumoral immune heterogeneity in non-small cell lung cancer. Nat Commun. 2018;9:5361.
Xu DH, Li Q, Hu H, Ni B, Liu X, Huang C, et al. Transmembrane protein GRINA modulates aerobic glycolysis and promotes tumor progression in gastric cancer. J Exp Clin Cancer Res. 2018;37:308.
D’Mello SA, Joseph WR, Green TN, Leung EY, During MJ, Finlay GJ, et al. Selected GRIN2A mutations in melanoma cause oncogenic effects that can be modulated by extracellular glutamate. Cell Calcium. 2016;60:384–95.
Huinen ZR, Huijbers EJM, van Beijnum JR, Nowak-Sliwinska P, Griffioen AW. Anti-angiogenic agents - overcoming tumor endothelial cell anergy and improving immunotherapy outcomes. Nature Rev Clin Oncol. 2021;18:527–40.
Long J, Lin J, Wang A, Wu L, Zheng Y, Yang X, et al. PD-1/PD-L blockade in gastrointestinal cancers: lessons learned and the road toward precision immunotherapy. J Hematol Oncol. 2017;10:146.
Wood MA, Nellore A, Thompson RF. Tumor mutation burden-from doubts to concerns. JAMA Oncol. 2019;5:1808–9.
Tsao MS, Kerr KM, Kockx M, Beasley MB, Borczuk AC, Botling J, et al. PD-L1 immunohistochemistry comparability study in real-life clinical samples: results of blueprint phase 2 project. J Thoracic Oncol. 2018;13:1302–11.
Acknowledgements
The authors would like to thank Drs. Xundong Zhang, and Xing-jie Hao (School of Public Health, Tongji Medical College, HUST) for their support in the bioinformatics analyses of this study.
Funding
This work was supported by the National Nature Science Foundation of China (No. 82273441 and 81874065), the National Basic Research Program of China (2020YFA0710700), the Knowledge Innovation Program of Wuhan-Shuguang Project (No. 2022020801020456), the Tongji Hospital (HUST) Foundation for Excellent Young Scientist (No. 2020YQ05), and the first level of the Public Health Youth Top Talent Project of Hubei Province (No. 2022SCZ051).
Author information
Authors and Affiliations
Contributions
Study concept and design: ZD, WZ, and BZ. Acquisition, analysis, interpretation of the data, critical revision of the manuscript: all the authors. Drafting of the manuscript: GL, TL, GJ and ZD. Study supervision: BZ and WZ.
Corresponding authors
Ethics declarations
Competing interests
ZD served as a speaker and consultant for Bayer, Eisai, Roche, MSD, Astra-Zeneca, Innovent, Hengrui, and BeiGene. The remaining authors have nothing to disclose.
Ethics approval and consent to participate
All the data used in this study are deidentified and publicly available. Therefore, the Institutional Review Board (IRB) of Tongji Hospital was waived.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Gx., Chang, Rz., Liu, Tt. et al. GRIN2A mutation is a novel indicator of stratifying beneficiaries of immune checkpoint inhibitors in multiple cancers. Cancer Gene Ther 31, 586–598 (2024). https://doi.org/10.1038/s41417-024-00730-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-024-00730-6