Abstract
MicroRNA (miRNA) serves as a pivotal regulator of numerous cellular processes, and the identification of miRNA-disease associations (MDAs) is crucial for comprehending complex diseases. Recently, graph neural networks (GNN) have made significant advancements in MDA prediction. However, these methods tend to learn one type of node representation from a single heterogeneous network, ignoring the importance of multiple network topologies and node attributes. Here, we propose SMDAP (Sequence hierarchical modeling-based Mirna-Disease Association Prediction framework), a novel GNN-based framework that incorporates multiple network topologies and various node attributes including miRNA seed and full-length sequences to predict potential MDAs. Specifically, SMDAP consists of two types of MDA representation: following a heterogeneous pattern, we construct a transfer learning-like synchronous mutual learning network to learn the first MDA representation in conjunction with the miRNA seed sequence. Meanwhile, following a homogeneous pattern, we design a subgraph-inspired asynchronous multi-scale embedding network to obtain the second MDA representation based on the miRNA full-length sequence. Subsequently, an adaptive fusion approach is designed to combine the two branches such that we can score the MDAs by the downstream classifier and infer novel MDAs. Comprehensive experiments demonstrate that SMDAP integrates the advantages of multiple network topologies and node attributes into two branch representations. Moreover, the area under the receiver operating characteristic curve is 0.9622 on DB1, which is a 5.06% increase from the baselines. The area under the precision–recall curve is 0.9777, which is a 7.33% increase from the baselines. In addition, case studies on three human cancers validated the predictive performance of SMDAP. Overall, SMDAP represents a powerful tool for MDA prediction.
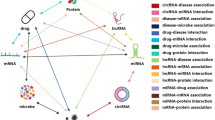
Graphical Abstract












Similar content being viewed by others
Data availability
Data will be made available on request.
References
Salmena L, Poliseno L, Tay Y et al (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146:353–358. https://doi.org/10.1016/j.cell.2011.07.014
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. https://doi.org/10.1038/nature02871
Reichenstein I, Eitan C, Diaz-Garcia S et al (2019) Human genetics and neuropathology suggest a link between miR-218 and amyotrophic lateral sclerosis pathophysiology. Sci Transl Med 11:eaav5264. https://doi.org/10.1126/scitranslmed.aav5264
Ucar A, Gupta SK, Fiedler J et al (2012) The miRNA-212/132 family regulates both cardiac hypertrophy and cardiomyocyte autophagy. Nat Commun 3:1–11. https://doi.org/10.1038/ncomms2090
Norsworthy PJ, Thompson AGB, Mok TH et al (2020) A blood miRNA signature associates with sporadic Creutzfeldt–Jakob disease diagnosis. Nat Commun 11:1–11. https://doi.org/10.1038/s41467-020-17655-x
Zheng K, You Z-H, Wang L et al (2020) DBMDA: a unified embedding for sequence-based miRNA similarity measure with applications to predict and validate miRNA-disease associations. Mol Ther Nucleic Acids 19:602–611. https://doi.org/10.1016/j.omtn.2019.12.010
Li H-Y, You Z-H, Wang L et al (2021) DF-MDA: an effective diffusion-based computational model for predicting miRNA-disease association. Mol Ther 29:1501–1511. https://doi.org/10.1016/j.ymthe.2021.01.003
Ji B-Y, You Z-H, Cheng L et al (2020) Predicting miRNA-disease association from heterogeneous information network with GraRep embedding model. Sci Rep 10:1–12. https://doi.org/10.1038/s41598-020-63735-9
Zhang Y, Chen M, Cheng X et al (2020) MSFSP: a novel miRNA–disease association prediction model by federating multiple-similarities fusion and space projection. Front Genet 11:389. https://doi.org/10.3389/fgene.2020.00389
Tang X, Luo J, Shen C et al (2021) Multi-view multichannel attention graph convolutional network for miRNA–disease association prediction. Brief Bioinform 22:bbab174. https://doi.org/10.1093/bib/bbab174
Li J, Chen X, Huang Q et al (2020) Seq-SymRF: a random forest model predicts potential miRNA-disease associations based on information of sequences and clinical symptoms. Sci Rep 10:1–11. https://doi.org/10.1038/s41598-020-75005-9
Li J, Zhang S, Liu T et al (2020) Neural inductive matrix completion with graph convolutional networks for miRNA-disease association prediction. Bioinformatics 36:2538–2546. https://doi.org/10.1093/bioinformatics/btz965
Liu D, Huang Y, Nie W et al (2021) SMALF: miRNA-disease associations prediction based on stacked autoencoder and XGBoost. BMC Bioinform 22:1–18. https://doi.org/10.1186/s12859-021-04135-2
Ding Y, Tian L-P, Lei X et al (2021) Variational graph auto-encoders for miRNA-disease association prediction. Methods 192:25–34. https://doi.org/10.1016/j.ymeth.2020.08.004
Chen X, Xie D, Wang L et al (2018) BNPMDA: bipartite network projection for MiRNA–disease association prediction. Bioinformatics 34:3178–3186. https://doi.org/10.1093/bioinformatics/bty333
Li Z, Li J, Nie R et al (2021) A graph auto-encoder model for miRNA-disease associations prediction. Brief Bioinform 22:bbaa240. https://doi.org/10.1093/bib/bbaa240
Jin C, Shi Z, Lin K et al (2022) Predicting miRNA-disease association based on neural inductive matrix completion with graph autoencoders and self-attention mechanism. Biomolecules 12:64. https://doi.org/10.3390/biom12010064
Zhu R, Ji C, Wang Y et al (2020) Heterogeneous graph convolutional networks and matrix completion for miRNA-disease association prediction. Front Bioeng Biotechnol 8:901. https://doi.org/10.3389/fbioe.2020.00901
Ding Y, Lei X, Liao B et al (2021) Predicting mirna-disease associations based on multi-view variational graph auto-encoder with matrix factorization. IEEE J Biomed Health Inform 26:446–457. https://doi.org/10.1109/jbhi.2021.3088342
Chen X, Wang L, Qu J et al (2018) Predicting miRNA–disease association based on inductive matrix completion. Bioinformatics 34:4256–4265. https://doi.org/10.1093/bioinformatics/bty503
Chen X, Sun L-G, Zhao Y (2021) NCMCMDA: miRNA–disease association prediction through neighborhood constraint matrix completion. Brief Bioinform 22:485–496. https://doi.org/10.1093/bib/bbz159
Chen X, Li S-X, Yin J et al (2020) Potential miRNA-disease association prediction based on kernelized Bayesian matrix factorization. Genomics 112:809–819. https://doi.org/10.1016/j.ygeno.2019.05.021
Öztürk Ş, Çukur T (2023) Focal modulation network for lung segmentation in chest X-ray images. Turk J Electr Eng Comput Sci 31:1006–1020. https://doi.org/10.55730/1300-0632.4031
Daşdemir Y, Özakar R (2022) Affective states classification performance of audio-visual stimuli from EEG signals with multiple-instance learning. Turk J Electr Eng Comput Sci 30:2707–2724. https://doi.org/10.55730/1300-0632.3964
Zheng K, You Z-H, Wang L et al (2019) MLMDA: a machine learning approach to predict and validate MicroRNA–disease associations by integrating of heterogenous information sources. J Transl Med 17:1–14. https://doi.org/10.1186/s12967-019-2009-x
Wang L, You Z-H, Chen X et al (2019) LMTRDA: Using logistic model tree to predict MiRNA-disease associations by fusing multi-source information of sequences and similarities. PLoS Comput Biol 15:e1006865. https://doi.org/10.1371/journal.pcbi.1006865
Chen X, Li T-H, Zhao Y et al (2021) Deep-belief network for predicting potential miRNA-disease associations. Brief Bioinform 22:bbaa186. https://doi.org/10.1093/bib/bbaa186
Giorgi JM, Bader GD (2018) Transfer learning for biomedical named entity recognition with neural networks. Bioinformatics 34:4087–4094. https://doi.org/10.1093/bioinformatics/bty449
Petegrosso R, Park S, Hwang TH et al (2017) Transfer learning across ontologies for phenome–genome association prediction. Bioinformatics 33:529–536. https://doi.org/10.1093/bioinformatics/btw649
Akalin F, Yumuşak N (2022) Detection and classification of white blood cells with an improved deep learning-based approach. Turk J Electr Eng Comput Sci 30:2725–2739. https://doi.org/10.55730/1300-0632.3965
Ünal Y, Öztürk Ş, Dudak MN et al (2022) Comparison of current convolutional neural network architectures for classification of damaged and undamaged cars. In: Advances in deep learning, artificial intelligence and robotics: proceedings of the 2nd international conference on deep learning, artificial intelligence and robotics (ICDLAIR) 2020. Springer, pp 141–149. https://doi.org/10.1007/978-3-030-85365-5_14
Ma M, Na S, Zhang X et al (2022) SFGAE: a self-feature-based graph autoencoder model for miRNA–disease associations prediction. Brief Bioinform 23:bbac340. https://doi.org/10.1093/bib/bbac340
Vaswani A, Shazeer N, Parmar N et al (2017) Attention is all you need. Adv Neural Inf Process Syst. https://doi.org/10.48550/arXiv.1706.03762
Berg R van den, Kipf TN, Welling M (2017) Graph convolutional matrix completion. arXiv preprint arXiv:170602263. https://doi.org/10.48550/arXiv.1706.02263
Wu F, Souza A, Zhang T et al (2019) Simplifying graph convolutional networks. In: International conference on machine learning, vol 97. PMLR, pp 6861–6871.
Wang X, Ji H, Shi C et al (2019) Heterogeneous graph attention network. In: The world wide web conference. pp 2022–2032. https://doi.org/10.1145/3308558.3313562
Wang X, He X, Wang M et al (2019) Neural graph collaborative filtering. In: Proceedings of the 42nd international ACM SIGIR conference on research and development in information retrieval. pp 165–174. https://doi.org/10.1145/3331184.3331267
Kipf TN, Welling M (2016) Semi-supervised classification with graph convolutional networks. arXiv preprint arXiv:160902907. https://doi.org/10.48550/arXiv.1609.02907
Xu K, Li C, Tian Y et al (2018) Representation learning on graphs with jumping knowledge networks. In: International conference on machine learning. PMLR, pp 5453–5462.
Chen M, Wei Z, Huang Z et al (2020) Simple and deep graph convolutional networks. In: International conference on machine learning. PMLR, pp 1725–1735
Sun F, Sun J, Zhao Q (2022) A deep learning method for predicting metabolite–disease associations via graph neural network. Brief Bioinform 23:bbac266. https://doi.org/10.1093/bib/bbac266
Wang W, Zhang L, Sun J et al (2022) Predicting the potential human lncRNA–miRNA interactions based on graph convolution network with conditional random field. Brief Bioinform 23:bbac463. https://doi.org/10.1093/bib/bbac463
Gao H, Sun J, Wang Y et al (2023) Predicting metabolite–disease associations based on auto-encoder and non-negative matrix factorization. Brief Bioinform 24:bbad259. https://doi.org/10.1093/bib/bbad259
Wang T, Sun J, Zhao Q (2023) Investigating cardiotoxicity related with hERG channel blockers using molecular fingerprints and graph attention mechanism. Comput Biol Med 153:106464. https://doi.org/10.1016/j.compbiomed.2022.106464
Zhang L, Yang P, Feng H et al (2021) Using network distance analysis to predict lncRNA–miRNA interactions. Interdiscip Sci 13:535–545. https://doi.org/10.1007/s12539-021-00458-z
Chen X, Yin J, Qu J et al (2018) MDHGI: matrix decomposition and heterogeneous graph inference for miRNA-disease association prediction. PLoS Comput Biol 14:e1006418. https://doi.org/10.1371/journal.pcbi.1006418
Wang Y-T, Wu Q-W, Gao Z et al (2021) MiRNA-disease association prediction via hypergraph learning based on high-dimensionality features. BMC Med Inform Decis Mak 21:1–13. https://doi.org/10.1186/s12911-020-01320-w
Ji C, Gao Z, Ma X et al (2021) AEMDA: inferring miRNA–disease associations based on deep autoencoder. Bioinformatics 37:66–72. https://doi.org/10.1093/bioinformatics/btaa670
Jeffrey HJ (1990) Chaos game representation of gene structure. Nucleic Acids Res 18:2163–2170. https://doi.org/10.1093/nar/18.8.2163
Wang Y, Yao H, Zhao S (2016) Auto-encoder based dimensionality reduction. Neurocomputing 184:232–242. https://doi.org/10.1016/j.neucom.2015.08.104
Wang D, Wang J, Lu M et al (2010) Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26:1644–1650. https://doi.org/10.1093/bioinformatics/btq241
Grover A, Leskovec J (2016) node2vec: Scalable feature learning for networks. In: Proceedings of the 22nd ACM SIGKDD international conference on knowledge discovery and data mining. pp 855–864. https://doi.org/10.1145/2939672.2939754
Shu H, Wang X, Zhu H (2020) D-CCA: A decomposition-based canonical correlation analysis for high-dimensional datasets. J Am Stat Assoc 115:292–306. https://doi.org/10.1080/01621459.2018.1543599
Yang Z, Wu L, Wang A et al (2017) dbDEMC 2.0: updated database of differentially expressed miRNAs in human cancers. Nucleic Acids Res 45:D812–D818. https://doi.org/10.1093/nar/gkw1079
Jiang Q, Wang Y, Hao Y et al (2009) miR2Disease: a manually curated database for microRNA deregulation in human disease. Nucleic Acids Res 37:D98–D104. https://doi.org/10.1093/nar/gkn714
Huang Z, Shi J, Gao Y et al (2019) HMDD v3. 0: a database for experimentally supported human microRNA–disease associations. Nucleic Acids Res 47:D1013–D1017. https://doi.org/10.1093/nar/gky1010
Kozomara A, Birgaoanu M, Griffiths-Jones S (2019) miRBase: from microRNA sequences to function. Nucleic Acids Res 47:D155–D162. https://doi.org/10.1093/nar/gky1141
Van Der Maaten L, Hinton G (2008) Visualizing data using t-SNE. J Mach Learn Res 9:2579–2625
Zhou S, Wang S, Wu Q et al (2020) Predicting potential miRNA-disease associations by combining gradient boosting decision tree with logistic regression. Comput Biol Chem 85:107200. https://doi.org/10.1016/j.compbiolchem.2020.107200
Dai Q, Wang Z, Liu Z et al (2022) Predicting miRNA-disease associations using an ensemble learning framework with resampling method. Brief Bioinform 23:bbab543. https://doi.org/10.1093/bib/bbab543
Cho WCS, Chow ASC, Au JSK (2011) MiR-145 inhibits cell proliferation of human lung adenocarcinoma by targeting EGFR and NUDT1. RNA Biol 8:125–131. https://doi.org/10.4161/rna.8.1.14259
Grose D, Morrison DS, Devereux G et al (2015) The impact of comorbidity upon determinants of outcome in patients with lung cancer. Lung Cancer 87:186–192. https://doi.org/10.1016/j.lungcan.2014.11.012
Iorio MV, Ferracin M, Liu C-G et al (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65:7065–7070. https://doi.org/10.1158/0008-5472.CAN-05-1783
Takahashi R, Miyazaki H, Ochiya T (2015) The roles of microRNAs in breast cancer. Cancers (Basel) 7:598–616. https://doi.org/10.3390/cancers7020598
Luo X, Burwinkel B, Tao S et al (2011) MicroRNA signatures: novel biomarker for colorectal cancer? microRNA and colorectal cancer. Cancer Epidemiol Biomark Prev 20:1272–1286. https://doi.org/10.1158/1055-9965.EPI-11-0035
Bandres E, Agirre X, Bitarte N et al (2009) Epigenetic regulation of microRNA expression in colorectal cancer. Int J Cancer 125:2737–2743. https://doi.org/10.1002/ijc.24638
Agarwal V, Bell GW, Nam J-W et al (2015) Predicting effective microRNA target sites in mammalian mRNAs. Elife 4:e05005. https://doi.org/10.7554/eLife.05005
Sakre N, Wildey G, Behtaj M et al (2017) RICTOR amplification identifies a subgroup in small cell lung cancer and predicts response to drugs targeting mTOR. Oncotarget 8:5992. https://doi.org/10.18632/oncotarget.13362
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31771430), and Hubei Hongshan Laboratory. Model training was performed at the Hefei Advanced Computing Center.
Author information
Authors and Affiliations
Contributions
WS: Methodology, Software, Writing—original draft. PZ: Methodology, Conceptualization, Writing—review and editing. WZ: Visualization, Investigation. JX: Formal analysis, Validation. YH: Data Curation. LL: Supervision, Writing—review and editing, Funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, W., Zhang, P., Zhang, W. et al. Synchronous Mutual Learning Network and Asynchronous Multi-Scale Embedding Network for miRNA-Disease Association Prediction. Interdiscip Sci Comput Life Sci (2024). https://doi.org/10.1007/s12539-023-00602-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12539-023-00602-x