Abstract
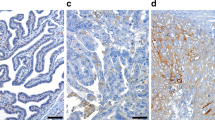
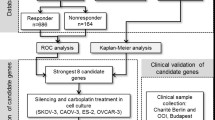
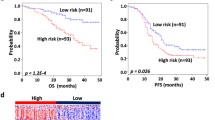
Chemotherapy resistance is the main reason for the poor prognosis of ovarian cancer (OC). FHL1 is an important tumour regulator, but its relationship with the prognosis, drug resistance, and tumour microenvironment of OC is unknown. Immunohistochemistry was used to determine FHL1 expression in OC. Kaplan‒Meier plotter was used for survival analysis. The value of gene expression in predicting drug resistance was estimated using the area under the curve (AUC). Bivariate correlation was used to determine the coexpression of two genes. Functional cluster and pathway enrichment were used to uncover hidden signalling pathways. The relationship between gene levels and the tumour microenvironment was visualised through the ggstatsplot and pheatmap packages. The mRNA and protein levels of FHL1 were downregulated in 426 and 100 OC tissues, respectively. Low FHL1 expression was correlated with good progression-free survival (PFS), postprogression survival, and overall survival (OS) in 1815 OC patients, and was further confirmed to be associated with good OS by immunohistochemistry in 152 OC tissues. Furthermore, FHL1 was downregulated in drug-sensitive tissues, while its high expression predicted drug resistance (AUC > 0.65). Mechanistically, FHL1 was coexpressed with FLNC, CAV1, PPP1R12B, and FLNA at the mRNA and protein levels in 558 and 174 OC tissues, respectively, and their expression was downregulated in OC. Additionally, very strong coexpression of FHL1 with the four genes was identified in at least 23 different tumours. Low expression of the four genes was associated with good PFS, and the combination of FHL1 with the four genes provided better prognostic power. Meanwhile, the expression of all five genes was strongly and positively associated with the abundance of macrophages. Low FHL1 expression acts as a favourable factor in OC, probably via positive coexpression with FLNC, CAV1, PPP1R12B, and FLNA.









Similar content being viewed by others
Data availability
The data required for immunohistochemical analysis are available from the corresponding author, and other data can be found in the relevant open online databases.
References
Asada K, Ando T, Niwa T et al (2013) FHL1 on chromosome X is a single-hit gastrointestinal tumor-suppressor gene and contributes to the formation of an epigenetic field defect. Oncogene 32:2140–2149. https://doi.org/10.1038/onc.2012.228
Bingle L, Brown NJ, Lewis CE (2002) The role of tumour-associated macrophages in tumour progression: implications for new anticancer therapies. J Pathol 196:254–265. https://doi.org/10.1002/path.1027
Bjorge L, Hakulinen J, Vintermyr OK et al (2005) Ascitic complement system in ovarian cancer. Br J Cancer 92:895–905. https://doi.org/10.1038/sj.bjc.6602334
Bu D, Luo H, Huo P et al (2021) KOBAS-i: intelligent prioritization and exploratory visualization of biological functions for gene enrichment analysis. Nucleic Acids Res 49:W317–W325. https://doi.org/10.1093/nar/gkab447
Celus W, Di Conza G, Oliveira AI et al (2017) Loss of caveolin-1 in metastasis-associated macrophages drives lung metastatic growth through increased angiogenesis. Cell Rep 21:2842–2854. https://doi.org/10.1016/j.celrep.2017.11.034
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Chandrashekar DS, Karthikeyan SK, Korla PK et al (2022) UALCAN: an update to the integrated cancer data analysis platform. Neoplasia 25:18–27. https://doi.org/10.1016/j.neo.2022.01.001
Chatterjee M, Ben-Josef E, Thomas DG et al (2015) Caveolin-1 is associated with tumor progression and confers a multi-modality resistance phenotype in pancreatic cancer. Sci Rep 5:10867. https://doi.org/10.1038/srep10867
Chen L, Zhang D, Zheng S et al (2022a) Stemness analysis in hepatocellular carcinoma identifies an extracellular matrix gene-related signature associated with prognosis and therapy response. Front Genet 13:959834. https://doi.org/10.3389/fgene.2022.959834
Chen Y, Pei M, Li J et al (2022b) Disruption of the CCDC43-FHL1 interaction triggers apoptosis in gastric cancer cells. Exp Cell Res 415:113107. https://doi.org/10.1016/j.yexcr.2022.113107
de Haas T, Hasselt N, Troost D et al (2008) Molecular risk stratification of medulloblastoma patients based on immunohistochemical analysis of MYC, LDHB, and CCNB1 expression. Clin Cancer Res 14:4154–4160. https://doi.org/10.1158/1078-0432.Ccr-07-4159
Deng L, Wang D, Chen S et al (2021) Epiphycan predicts poor outcomes and promotes metastasis in ovarian cancer. Front Oncol 11:653782. https://doi.org/10.3389/fonc.2021.653782
Diao C, Xi Y, Xiao T (2017) Identification and analysis of key genes in osteosarcoma using bioinformatics. Oncol Lett. https://doi.org/10.3892/ol.2017.7649
Diaz-Pena R, Boekstegers F, Silva RS et al (2020) Amerindian ancestry influences genetic susceptibility to chronic obstructive pulmonary disease. J Pers Med 10. https://doi.org/10.3390/jpm10030093
Dong C, Yin F, Zhu D et al (2020) NCALD affects drug resistance and prognosis by acting as a ceRNA of CX3CL1 in ovarian cancer. J Cell Biochem 121:4470–4483. https://doi.org/10.1002/jcb.29670
Drexler R, Fahy R, Kuchler M et al (2021) Association of subcellular localization of TEAD transcription factors with outcome and progression in pancreatic ductal adenocarcinoma. Pancreatology 21:170–179. https://doi.org/10.1016/j.pan.2020.12.003
Edwards NJ, Oberti M, Thangudu RR et al (2015) The CPTAC data portal: a resource for cancer proteomics research. J Proteome Res 14:2707–2713. https://doi.org/10.1021/pr501254j
Fan S, Liang Z, Gao Z et al (2018) Identification of the key genes and pathways in prostate cancer. Oncol Lett 16:6663–6669. https://doi.org/10.3892/ol.2018.9491
Fekete JT, Gyorffy B (2019) ROCplot.org: validating predictive biomarkers of chemotherapy/hormonal therapy/anti-HER2 therapy using transcriptomic data of 3,104 breast cancer patients. Int J Cancer 145:3140–3151. https://doi.org/10.1002/ijc.32369
Fu Y, Xu M, Cui Z et al (2020) Genome-wide identification of FHL1 as a powerful prognostic candidate and potential therapeutic target in acute myeloid leukaemia. EBioMedicine 52:102664. https://doi.org/10.1016/j.ebiom.2020.102664
Gou R, Zhu L, Zheng M et al (2019) Annexin A8 can serve as potential prognostic biomarker and therapeutic target for ovarian cancer: based on the comprehensive analysis of Annexins. J Transl Med 17:275. https://doi.org/10.1186/s12967-019-2023-z
Gyorffy B, Lanczky A, Eklund AC et al (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123:725–731. https://doi.org/10.1007/s10549-009-0674-9
Gyorffy B, Lanczky A, Szallasi Z (2012) Implementing an online tool for genome-wide validation of survival-associated biomarkers in ovarian-cancer using microarray data from 1287 patients. Endocr Relat Cancer 19:197–208. https://doi.org/10.1530/ERC-11-0329
He S, Yang L, Xiao Z et al (2021) Identification of key carcinogenic genes in Wilms’ tumor. Genes Genet Syst 96:141–149. https://doi.org/10.1266/ggs.21-00015
Junnikkala S, Hakulinen J, Jarva H et al (2002) Secretion of soluble complement inhibitors factor H and factor H-like protein (FHL-1) by ovarian tumour cells. Br J Cancer 87:1119–1127. https://doi.org/10.1038/sj.bjc.6600614
Kamil M, Shinsato Y, Higa N et al (2019) High filamin-C expression predicts enhanced invasiveness and poor outcome in glioblastoma multiforme. Br J Cancer 120:819–826. https://doi.org/10.1038/s41416-019-0413-x
Li JH, Liu S, Zhou H et al (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92-97. https://doi.org/10.1093/nar/gkt1248
Li B, Severson E, Pignon JC et al (2016) Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol 17:174. https://doi.org/10.1186/s13059-016-1028-7
Liao C, Zhao YX, Han WD et al (2022) RIPK4 is an immune regulating-associated biomarker for ovarian cancer and possesses generalization value in pan-cancer. J Immunol Res 2022:7599098. https://doi.org/10.1155/2022/7599098
Lisowska KM, Olbryt M, Dudaladava V et al (2014) Gene expression analysis in ovarian cancer - faults and hints from DNA microarray study. Front Oncol 4:6. https://doi.org/10.3389/fonc.2014.00006
Liu M, Liu X, Liu S et al (2020) Big data-based identification of multi-gene prognostic signatures in liver cancer. Front Oncol 10:847. https://doi.org/10.3389/fonc.2020.00847
Liu Y, Wang C, Cheng P et al (2021) FHL1 inhibits the progression of colorectal cancer by regulating the Wnt/beta-catenin signaling pathway. J Cancer 12:5345–5354. https://doi.org/10.7150/jca.60543
Liu Z, Liu J, Liu X et al (2022) CTR-DB, an omnibus for patient-derived gene expression signatures correlated with cancer drug response. Nucleic Acids Res 50:D1184–D1199. https://doi.org/10.1093/nar/gkab860
Liu X, Li X, Kuang Q et al (2023) Screening of immunotherapy-related genes in bladder cancer based on GEO datasets. Front Oncol 13:1176637. https://doi.org/10.3389/fonc.2023.1176637
Lu W, Mengxuan Z, Ming R et al (2022) TRIP13/FLNA complex promotes tumor progression and is associated with unfavorable outcomes in melanoma. J Oncol 2022:1419179. https://doi.org/10.1155/2022/1419179
Matsumoto M, Kawakami K, Enokida H et al (2010) CpG hypermethylation of human four-and-a-half LIM domains 1 contributes to migration and invasion activity of human bladder cancer. Int J Mol Med 26:241–247. https://doi.org/10.3892/ijmm_00000458
Niu C, Liang C, Guo J et al (2012) Downregulation and growth inhibitory role of FHL1 in lung cancer. Int J Cancer 130:2549–2556. https://doi.org/10.1002/ijc.26259
Patil I (2021) Visualizations with statistical details: the ‘ggstatsplot’ approach. Journal of Open Source Software 6. https://doi.org/10.21105/joss.03167
Popek S, Kapka-Skrzypczak L, Sawicki K et al (2016) IL-6 and IL-8 enhance factor H binding to the cell membranes. Mol Med Rep 13:3886–3894. https://doi.org/10.3892/mmr.2016.5012
Ren W, Lian P, Cheng L et al (2015) FHL1 inhibits the growth of tongue squamous cell carcinoma cells via G1/S cell cycle arrest. Mol Med Rep 12:3958–3964. https://doi.org/10.3892/mmr.2015.3844
Shathasivam T, Kislinger T, Gramolini AO (2010) Genes, proteins and complexes: the multifaceted nature of FHL family proteins in diverse tissues. J Cell Mol Med 14:2702–2720. https://doi.org/10.1111/j.1582-4934.2010.01176.x
Siegel RL, Miller KD, Wagle NS et al (2023) Cancer statistics, 2023. CA Cancer J Clin 73:17–48. https://doi.org/10.3322/caac.21763
Sung H, Ferlay J, Siegel RL et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71:209–249. https://doi.org/10.3322/caac.21660
Tang Z, Li C, Kang B et al (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45:W98–W102. https://doi.org/10.1093/nar/gkx247
Thorsson V, Gibbs DL, Brown SD et al (2018) The immune landscape of cancer. Immunity 48(812–830):e814. https://doi.org/10.1016/j.immuni.2018.03.023
Tian W, Lei N, Zhou J et al (2022) Extracellular vesicles in ovarian cancer chemoresistance, metastasis, and immune evasion. Cell Death Dis 13:64. https://doi.org/10.1038/s41419-022-04510-8
Wang X, Wei X, Yuan Y et al (2018) Src-mediated phosphorylation converts FHL1 from tumor suppressor to tumor promoter. J Cell Biol 217:1335–1351. https://doi.org/10.1083/jcb.201708064
Wei X, Zhang H (2020) Four and a half LIM domains protein 1 can be as a double-edged sword in cancer progression. Cancer Biol Med 17:270–281. https://doi.org/10.20892/j.issn.2095-3941.2019.0420
Wick MR (2019) The hematoxylin and eosin stain in anatomic pathology-an often-neglected focus of quality assurance in the laboratory. Semin Diagn Pathol 36:303–311. https://doi.org/10.1053/j.semdp.2019.06.003
Wu P, Heins ZJ, Muller JT et al (2019) Integration and analysis of CPTAC proteomics data in the context of cancer genomics in the cBioPortal. Mol Cell Proteomics 18:1893–1898. https://doi.org/10.1074/mcp.TIR119.001673
Wu S, Liu M, Xiao S et al (2023) Identification and verification of novel ferroptosis biomarkers predicts the prognosis of hepatocellular carcinoma. Genomics 115. https://doi.org/10.1016/j.ygeno.2023.110733
Xu X, Fan Z, Liang C et al (2017) A signature motif in LIM proteins mediates binding to checkpoint proteins and increases tumour radiosensitivity. Nat Commun 8:14059. https://doi.org/10.1038/ncomms14059
Yang H, Yu M, Zhong S et al (2022) Neoantigens and the tumor microenvironment play important roles in the prognosis of high-grade serous ovarian cancer. J Ovarian Res 15:18. https://doi.org/10.1186/s13048-022-00955-9
Ye G, Feng S, Yang Y et al (2022) Establishment and comprehensive analysis of underlying microRNA-mRNA interactive networks in ovarian cancer. J Oncol 2022:5120342. https://doi.org/10.1155/2022/5120342
Zeller C, Dai W, Steele NL et al (2012) Candidate DNA methylation drivers of acquired cisplatin resistance in ovarian cancer identified by methylome and expression profiling. Oncogene 31:4567–4576. https://doi.org/10.1038/onc.2011.611
Zhai L, Bell A, Ladomersky E et al (2021) Tumor cell IDO enhances immune suppression and decreases survival independent of tryptophan metabolism in glioblastoma. Clin Cancer Res 27:6514–6528. https://doi.org/10.1158/1078-0432.CCR-21-1392
Zhang K, Kong X, Feng G et al (2018) Investigation of hypoxia networks in ovarian cancer via bioinformatics analysis. J Ovarian Res 11:16. https://doi.org/10.1186/s13048-018-0388-x
Zhang S, Cao M, Yan S et al (2022) TRIM44 promotes BRCA1 functions in HR repair to induce cisplatin chemoresistance in lung adenocarcinoma by deubiquitinating FLNA. Int J Biol Sci 18:2962–2979. https://doi.org/10.7150/ijbs.71283
Zhang J, Li H, Guo M et al (2023) FHL1 as a novel prognostic biomarker and correlation with immune infiltration levels in lung adenocarcinoma. Immunotherapy. https://doi.org/10.2217/imt-2022-0195
Zhou L, Ding L, Liu J et al (2018) Four-and-a-half LIM protein 1 promotes paclitaxel resistance in hepatic carcinoma cells through the regulation of caspase-3 activation. J Cancer Res Ther 14:S767–S773. https://doi.org/10.4103/0973-1482.187304
Zou W, Ma X, Hua W et al (2015) Caveolin-1 mediates chemoresistance in cisplatin-resistant ovarian cancer cells by targeting apoptosis through the Notch-1/Akt/NF-kappaB pathway. Oncol Rep 34:3256–3263. https://doi.org/10.3892/or.2015.4320
Funding
This work was supported by Guangxi Natural Science Foundation (2021GXNSFAA075002), the National Natural Science Foundation of China (82260721, 81903644), the Advanced Innovation Teams and Xinghu Scholars Program of Guangxi Medical University (24/02304001018X), and the Program Foundation of Key Laboratory of High-Incidence Tumor Prevention and Treatment, Ministry of Education (GKE2019-20, GKE-ZZ202148).
Author information
Authors and Affiliations
Contributions
FY and XL designed and reviewed the study. XC drafted the manuscript and FQ revised the manuscript. XC and YY carried out the majority of the experiments and bioinformatics analysis. XC, YY, YS, LS, SX and YH were participated in material preparation and data collection. The final manuscript was read and approved by all authors.
Corresponding authors
Ethics declarations
Ethical approval
The authors take responsibility for this research to ensure that any questions in regard to its accuracy or integrity are sufficiently addressed. The work was performed following the Declaration of Helsinki guidelines (2013 revision).
Consent for publication
Not applicable.
Competing interests
All authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The original article contains an error. First, affiliation 1 should be corrected from: “Key Laboratory of Longevity and Ageing‑Related Disease of Chinese Ministry of Education, Centre for Translational Medicine and School of Preclinical Medicine, Guangxi Medical University, Nanning 530021, Guangxi, China” to “Key Laboratory of Longevity and Ageing-Related Disease of Chinese Ministry of Education, Centre for Translational Medicine and School of Basic Medical Sciences, Guangxi Medical University, Nanning 530021, Guangxi, China”. Secondly, the funding statement should be corrected from “This work was supported by Guangxi Natural Science Foundation (2021GXNSFAA075002), the National Natural Science Foundation of China (81903644), the Advanced Innovation Teams and Xinghu Scholars Program of Guangxi Medical University (24/02304001018X), and the Program Foundation of Key Laboratory of High-Incidence Tumor Prevention and Treatment, Ministry of Education (GKE2019-20, GKE-ZZ202148).” to “This work was supported by Guangxi Natural Science Foundation (2021GXNSFAA075002), the National Natural Science Foundation of China (82260721, 81903644), the Advanced Innovation Teams and Xinghu Scholars Program of Guangxi Medical University (24/02304001018X), and the Program Foundation of Key Laboratory of High-Incidence Tumor Prevention and Treatment, Ministry of Education (GKE2019-20, GKE-ZZ202148).
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, X., Yu, Y., Su, Y. et al. Low FHL1 expression indicates a good prognosis and drug sensitivity in ovarian cancer. Funct Integr Genomics 24, 25 (2024). https://doi.org/10.1007/s10142-024-01294-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-024-01294-2