Abstract
Chromatin accessibility is a hallmark of active regulatory regions and is functionally linked to transcriptional networks and cell identity. However, the molecular mechanisms and networks that govern chromatin accessibility have not been thoroughly studied. Here we conducted a genome-wide CRISPR screening combined with an optimized ATAC-see protocol to identify genes that modulate global chromatin accessibility. In addition to known chromatin regulators like CREBBP and EP400, we discovered a number of previously unrecognized proteins that modulate chromatin accessibility, including TFDP1, HNRNPU, EIF3D and THAP11 belonging to diverse biological pathways. ATAC-seq analysis upon their knockouts revealed their distinct and specific effects on chromatin accessibility. Remarkably, we found that TFDP1, a transcription factor, modulates global chromatin accessibility through transcriptional regulation of canonical histones. In addition, our findings highlight the manipulation of chromatin accessibility as an approach to enhance various cell engineering applications, including genome editing and induced pluripotent stem cell reprogramming.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The RNA-seq, ChIP–seq, ATAC-seq, MNase-seq and CRISPR-screening datasets in this research are available from the NCBI GEO repository: GSE144454. Data used in, but not generated in, this study include ChIP–seq datasets: GSE72800 (POLR2A), GSE94992 (SMC1A and CTCF), GSE108390 (H3K36me3, H3K4me1, H3K9me3 and H2AK119ub), PRJEB8671 (TFDP1 in LoVo cells), GSE105217 (TFDP1 in K562 cells), GSE80661 (TFDP1 in U266 cells) and GSE80661 (TFDP1 in MM1.S cells). Source data are provided with this paper.
Code availability
The publicly available softwares used are indicated in the Reporting Summary. The custom codes to analyze ATAC-seq data in Fig. 3 and Extended Data Fig. 7 are available in GitHub (https://github.com/Park-Sung-Joon/ATACprofWS, https://doi.org/10.5281/zenodo.10417228)76.
References
Boskovic, A., Bing, X. Y., Kaymak, E. & Rando, O. J. Control of noncoding RNA production and histone levels by a 5′ tRNA fragment. Genes Dev. 34, 118–131 (2020).
Celona, B. et al. Substantial histone reduction modulates genomewide nucleosomal occupancy and global transcriptional output. PLoS Biol. 9, e1001086 (2011).
Pal, S. & Tyler, J. K. Epigenetics and aging. Sci. Adv. 2, e1600584 (2016).
Klein, H.-U. et al. Epigenome-wide study uncovers large-scale changes in histone acetylation driven by tau pathology in aging and Alzheimer’s human brains. Nat. Neurosci. 22, 37–46 (2019).
Li, D. et al. Chromatin accessibility dynamics during iPSC reprogramming. Cell Stem Cell 21, 819–833.e6 (2017).
Liu, L. et al. An integrated chromatin accessibility and transcriptome landscape of human pre-implantation embryos. Nat. Commun. 10, 364 (2019).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Chen, X. et al. ATAC-see reveals the accessible genome by transposase-mediated imaging and sequencing. Nat. Methods 13, 1013–1020 (2016).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Zhou, M., Bhasin, A. & Reznikoff, W. S. Molecular genetic analysis of transposase-end DNA sequence recognition: cooperativity of three adjacent base-pairs in specific interaction with a mutant Tn5 transposase. J. Mol. Biol. 276, 913–925 (1998).
Essletzbichler, P. et al. Megabase-scale deletion using CRISPR/Cas9 to generate a fully haploid human cell line. Genome Res. 24, 2059–2065 (2014).
de Dieuleveult, M. et al. Genome-wide nucleosome specificity and function of chromatin remodellers in ES cells. Nature 530, 113–116 (2016).
Schick, S. et al. Systematic characterization of BAF mutations provides insights into intracomplex synthetic lethalities in human cancers. Nat. Genet. 51, 1399–1410 (2019).
Doench, J. G. et al. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR–Cas9. Nat. Biotechnol. 34, 184–191 (2016).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Görisch, S. M., Wachsmuth, M., Tóth, K. F., Lichter, P. & Rippe, K. Histone acetylation increases chromatin accessibility. J. Cell Sci. 118, 5825–5834 (2005).
Ryu, K., Kim, D.-S. & Kraus, L. W. New facets in the regulation of gene expression by ADP-ribosylation and poly(ADP-ribose) polymerases. Chem. Rev. 115, 2453–2481 (2015).
Azad, G. et al. PARP1-dependent eviction of the linker histone H1 mediates immediate early gene expression during neuronal activation. J. Cell Biol. 217, 473–481 (2018).
Kim, M. Y., Mauro, S., Gévry, N., Lis, J. T. & Kraus, W. NAD+-dependent modulation of chromatin structure and transcription by nucleosome binding properties of PARP-1. Cell 119, 803–814 (2004).
Krishnakumar, R. et al. Reciprocal binding of PARP-1 and histone H1 at promoters specifies transcriptional outcomes. Science 319, 819–821 (2008).
Schick, S. et al. Acute BAF perturbation causes immediate changes in chromatin accessibility. Nat. Genet. 53, 269–278 (2021).
Fischer, V., Schumacher, K., Tora, L. & Devys, D. Global role for coactivator complexes in RNA polymerase II transcription. Transcription 10, 1–8 (2018).
Stewart-Morgan, K. R., Nazaret, R.-G. & Groth, A. Transcription restart establishes chromatin accessibility after DNA replication. Mol. Cell 75, 284–297.e6 (2019).
Hauri, S. et al. A high-density map for navigating the human polycomb complexome. Cell Rep. 17, 583–595 (2016).
Wang, Q. et al. WDR68 is essential for the transcriptional activation of the PRC1-AUTS2 complex and neuronal differentiation of mouse embryonic stem cells. Stem Cell Res. 33, 206–214 (2018).
Bracken, A. P. et al. E2F target genes: unraveling the biology. Trends Biochem. Sci. 29, 409–417 (2004).
Wu, C. L., Classon, M., Dyson, N. & Harlow, E. Expression of dominant-negative mutant DP-1 blocks cell cycle progression in G1. Mol. Cell. Biol. 16, 3698–3706 (1996).
Lan, W. et al. E2F signature is predictive for the pancreatic adenocarcinoma clinical outcome and sensitivity to E2F inhibitors, but not for the response to cytotoxic-based treatments. Sci. Rep. 8, 8330 (2018).
Ma, Y. et al. A small-molecule E2F inhibitor blocks growth in a melanoma culture model. Cancer Res. 68, 6292–6299 (2008).
Yesbolatova, A. et al. The auxin-inducible degron 2 technology provides sharp degradation control in yeast, mammalian cells, and mice. Nat. Commun. 11, 5701 (2020).
Gokhman, D., Livyatan, I., Sailaja, B. S., Melcer, S. & Meshorer, E. Multilayered chromatin analysis reveals E2f, Smad and Zfx as transcriptional regulators of histones. Nat. Struct. Mol. Biol. 20, 119–126 (2013).
Oswald, F., Dobner, T. & Lipp, M. The E2F transcription factor activates a replication-dependent human H2A gene in early S phase of the cell cycle. Mol. Cell. Biol. 16, 1889–1895 (1996).
Rabinovich, A., Jin, V. X., Rabinovich, R., Xu, X. & Farnham, P. J. E2F in vivo binding specificity: comparison of consensus versus nonconsensus binding sites. Genome Res. 18, 1763–1777 (2008).
Singh, R., Kuscu, C., Quinlan, A., Qi, Y. & Adli, M. Cas9-chromatin binding information enables more accurate CRISPR off-target prediction. Nucleic Acids Res. 43, e118 (2015).
Wu, X. et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat. Biotechnol. 32, 670–676 (2014).
Knaupp, A. S. et al. Transient and permanent reconfiguration of chromatin and transcription factor occupancy drive reprogramming. Cell Stem Cell 21, 834–845.e6 (2017).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Nishimura, K. et al. Simple and effective generation of transgene-free induced pluripotent stem cells using an auto-erasable Sendai virus vector responding to microRNA-302. Stem Cell Res. 23, 13–19 (2017).
Chan, E. M. et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nat. Biotechnol. 27, 1033–1037 (2009).
Yang, C.-S., Chang, K.-Y. & Rana, T. M. Genome-wide functional analysis reveals factors needed at the transition steps of induced reprogramming. Cell Rep. 8, 327–337 (2014).
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nat. Genet. 45, 34–42 (2013).
Schmidt, R. & Plath, K. The roles of the reprogramming factors Oct4, Sox2 and Klf4 in resetting the somatic cell epigenome during induced pluripotent stem cell generation. Genome Biol. 13, 251 (2012).
Xu, Y. et al. Transcriptional control of somatic cell reprogramming. Trends Cell Biol. 26, 272–288 (2016).
Huangfu, D. et al. Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat. Biotechnol. 26, 795–797 (2008).
Mikkelsen, T. S. et al. Dissecting direct reprogramming through integrative genomic analysis. Nature 454, 49–55 (2008).
Shi, Y. et al. A combined chemical and genetic approach for the generation of induced pluripotent stem cells. Cell Stem Cell 2, 525–528 (2008).
Liu, B. et al. Inhibition of histone deacetylase 1 (HDAC1) and HDAC2 enhances CRISPR/Cas9 genome editing. Nucleic Acids Res. 48, 517–532 (2020).
Doege, C. A. et al. Early-stage epigenetic modification during somatic cell reprogramming by Parp1 and Tet2. Nature 488, 652–655 (2012).
Baranello, L., Kouzine, F., Sanford, S. & Levens, D. ChIP bias as a function of cross-linking time. Chromosome Res. 24, 175–181 (2016).
Birk, U. J. Super-resolution microscopy of chromatin. Genes 10, 493 (2019).
Nagaich, A. K., Walker, D. A., Wolford, R. & Hager, G. L. Rapid periodic binding and displacement of the glucocorticoid receptor during chromatin remodeling. Mol. Cell 14, 163–174 (2004).
Schmiedeberg, L., Skene, P., Deaton, A. & Bird, A. A temporal threshold for formaldehyde crosslinking and fixation. PLoS ONE 4, e4636 (2009).
Teytelman, L. et al. Impact of chromatin structures on DNA processing for genomic analyses. PLoS ONE 4, e6700 (2009).
Whelan, D. R. & Bell, T. D. M. Image artifacts in single molecule localization microscopy: why optimization of sample preparation protocols matters. Sci. Rep. 5, 1–10 (2015).
Oh, K.-S., Ha, J., Baek, S. & Sung, M.-H. XL-DNase-seq: improved footprinting of dynamic transcription factors. Epigenetics Chromatin 12, 30 (2019).
Zhang, H. et al. Extensive evaluation of ATAC-seq protocols for native or formaldehyde-fixed nuclei. BMC Genomics 23, 214 (2022).
Mulqueen, R. et al. Improved single-cell ATAC-seq reveals chromatin dynamics of in vitro corticogenesis. Preprint at bioRxiv https://doi.org/10.1101/637256 (2019).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Meers, M. P., Llagas, G., Janssens, D. H., Codomo, C. A. & Henikoff, S. Multifactorial profiling of epigenetic landscapes at single-cell resolution using MulTI-Tag. Nat. Biotechnol. 41, 708–716 (2023).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat. Rev. Genet. 24, 550–572 (2023).
Di, L. et al. RNA sequencing by direct tagmentation of RNA/DNA hybrids. Proc. Natl Acad. Sci. USA 117, 2886–2893 (2020).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Musinova, Y. R. et al. Nucleolar localization/retention signal is responsible for transient accumulation of histone H2B in the nucleolus through electrostatic interactions. Biochim. Biophys. Acta 1813, 27–38 (2011).
Li, W. et al. Quality control, modeling, and visualization of CRISPR screens with MAGeCK-VISPR. Genome Biol. 16, 281 (2015).
Wang, B. et al. Integrative analysis of pooled CRISPR genetic screens using MAGeCKFlute. Nat. Protoc. 14, 756–780 (2019).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Schnitzler, G. R. Isolation of histones and nucleosome cores from mammalian cells. Curr. Protoc. Mol. Biol. 50, 21.5.1–21.5.12 (2000).
Schägger, H. Tricine–SDS-PAGE. Nat. Protoc. 1, 16–22 (2006).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Bonhoure, N. et al. Quantifying ChIP–seq data: a spiking method providing an internal reference for sample-to-sample normalization. Genome Res. 24, 1157–1168 (2014).
Hu, B. et al. Biological chromodynamics: a general method for measuring protein occupancy across the genome by calibrating ChIP–seq. Nucleic Acids Res. 43, e132 (2015).
Orlando, D. A. et al. Quantitative ChIP–seq normalization reveals global modulation of the epigenome. Cell Rep. 9, 1163–1170 (2014).
Montefiori, L. et al. Reducing mitochondrial reads in ATAC-seq using CRISPR/Cas9. Sci. Rep. 7, 1–9 (2017).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Park, S. J. ATACprofWS.vR2. Zenodo https://doi.org/10.5281/zenodo.10417228 (2023).
Acknowledgements
We appreciate all members of Miyanari lab for their kind discussion, M. Hirabayashi and T. Kobayashi (National Institute for Physiological Sciences) for help with FACS analysis, T. Hiyama and M. Noda (National Institute for Basic Biology, NIBB) for providing HEK293T cells, T. Nishiyama (Nagoya University) and T. Kuraishi (Kanazawa University) for providing S2 cells, E. V. Sheval (Moscow State University) for providing pEGFP–H2B (21–35), K. Yamaguchi, A. Akita, S. Shigenobu and M. Matsumoto (NIBB) for technical support for deep sequencing, K. Naruse and A. Kato (NIBB) for Sanger sequencing, M. E. Torres-Padilla (Helmholtz Munich), T. Ishiuchi (Yamanashi University), M. A. Koyanagi, T. Aoi (Kobe University) and K. Nishimura (Tsukuba University) for comments on the manuscript, the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley, NYU Genome Technology Center for their deep sequencing service, NIBB Data Integration and Analysis Facility, the NIG supercomputer at ROIS National Institute of Genetics for the computational resources, and NIBB Bioimaging Facility for their imaging supports. S.I. and T.K. are supported by JSPS Grant-in-Aid for Young Scientists. S.-J.P. is supported by JSPS KAKENHI (18H04710 and 20H05940). Y.O. is supported by JSPS KAKENHI (JP23H00372, JP22H04676 and JP22K19275), AMED BINDS (JP22ama121017j0001), the Medical Research Center Initiative for High Depth Omics and the Cooperative Research Project Program of the Medical Institute of Bioregulation at Kyushu University. Y.M. is supported by JSPS KAKENHI (16H06279 (PAGS), 22H04925 (PAGS), 18H04722, 22H04688 and 21H04765), World Premier International Research Center Initiative (WPI, MEXT), Okazaki Orion Project, Astellas Foundation, JST/PRESTO, Takeda Science Foundation, the Cell Science Research Foundation, Mochida Memorial Foundation for Medical and Pharmaceutical Research, the Naito Foundation, the Cannon Foundation and Ohsumi Frontier Science Foundation.
Author information
Authors and Affiliations
Contributions
S.I. and Y.M. designed and performed experiments and wrote the paper. T.K. set up the optimized ATAC-see. A.T. supported cell sorting. C.S. performed the construction of plasmids. S.-J.P. and Y.M. conducted bioinformatic analyses. K.N. supported the bioinformatic analyses. H.T. provided a resource and technical advice for DNA FISH analysis. Y.O. contributed to the initial setup for deep sequencing. M.N. provided a resource for iPS cell reprogramming.
Corresponding author
Ethics declarations
Competing interests
M.N. is the founder and Chief Technology Officer of Tokiwa-Bio, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Andrey Krokhotin and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of optimized ATAC-see.
a, Mouse genomic DNA (gDNA) were tagmented using Tn5 transposome with Cy3-adaptor DNAs (ME19 or MEA34) with or without MgCl2. Agarose gel images for ethidium bromide (EtBr) and Cy3 channels are displayed. b, Representative images of NIH3T3 cells stained for ATAC-see, DAPI and H3K9me3 under the non-crosslinked condition. scale bar, 2 μm. c, Representative images of NIH3T3 cells stained for ATAC-see and DAPI under indicated crosslink conditions (the left panels). Green dashed circles, mitotic cells. scale bars, 5 μm. d, A bar plot shows intensity of ATAC-see on cells at metaphase relative to that at G2 phase. Data are presented as mean values +/− SD. n, number of cells analyzed. Statistics: a two-tailed unpaired Student’s t-test. e, Representative images of female mouse embryonic fibroblasts (MEFs) stained for ATAC-see, H3K27me3, DAPI, and X chromosome (Chr X, DNA-FISH) without cell-crosslinking (the top panels). Xi and Xa indicate inactive and active Chr X, respectively. Relative fluorescent intensities for indicated marks in Chr X territories are displayed at bottom panels. Box plots (the Bottom panels) are shown as Fig. 1c. n, number of cells analyzed. Scale bars, 5 μm. Statistics: a two-tailed unpaired Student’s t-test. f, Images for ATAC-see and H3K27me3 on female MEFs crosslinked with indicated concentration of PFA. Arrowheads and green dashed circles, inactive Chr X. Scale bars, 5 μm. g, Representative images of NIH3T3 cells stained for ATAC-see, DAPI, phospho histone H3 Ser10 (H3S10P) under the non-crosslinked condition (the upper panels, scale bar, 30 µm). Arrows, H3S10P-positive mitotic cells. Lower panels show higher magnification images of indicated cell-cycle stages. Scale bars, 5 μm. h, Scatter plot and histograms representing intensities for DAPI and ATAC-see in cells at i n, number of cells analyzed. Experiments in a, b, c, and g were reproduced three times.
Extended Data Fig. 2 Validation of optimized ATAC-see by ATAC-seq analysis.
a, Schematic illustrations displaying Tn5 transposome assembled with indicated adaptor DNAs. b, Representative images of NIH3T3 cells analyzed by ATAC-see using indicated transposomes (the left panels, scale bars, 30 μm). n, number of cells analyzed. A boxplot in the right panel shows relative intensity of ATAC-see. Statistics and definition of the box plot are the same as Fig. 1i. Experiments were reproduced three times. c, Representative genome browser tracks of ATAC-seq analysis with mES cells using indicated transposomes. d, A correlation matrix between ATAC-seq using indicated transposomes. A heat map represents clustering of three independent biological replicates for each experimental condition based on read enrichments in merged ATAC-seq peaks. e, Pairwise scatter plots of signals in the merged peaks. Values of correlation coefficients (Pearson) for each comparison are indicated. f, Insert-size distributions of ATAC-seq using indicated adaptor DNAs. g, Metagene profiles for read enrichments around the transcriptional start sites (TSS). h, Signal extraction scaling (SES) computed by using deeptools plotFingerprint represents cumulative read coverages.
Extended Data Fig. 3 CRISPR-mediated depletion of chromatin related genes.
eHAP cells expressing Cas9 were transfected with a plasmid encoding two pairs of sgRNAs against indicated genes and analyzed by either western blotting (a) or immunostaining (b) on day 4-5 post-transfection. a, Specific bands using indicated antibodies are shown with arrowheads. Size markers (kDa) are indicated in left side of each panel. These experiments were reproduced twice. b, Immunofluorescence images with indicated antibodies (gray) and DNA staining (green). scale bars, 30 μm. c, Knockout efficiency calculated based on immunostaining data. n; number of cells analyzed. Scale bars, 10 μm.
Extended Data Fig. 4 Extended results of the CRISPR screening.
a, A gating strategy for flow cytometry to collect cells used for the CRISPR screening. Cells stained for ATAC-see, DAPI, and phospho Histone H3 were analyzed for a DAPI channel to gate cells at G2/M phase. The cells were subsequently gated out based on the signals of phospho Histone H3 to eliminate the mitotic fraction. A cell population displaying the highest or lowest 5% of ATAC-see signals was collected, respectively. Immunostaining images for phospho H3 and DAPI before and after the cell-sorting (the bottom panels) confirm the high purity of G2 phase. Scale bars, 15 μm. b, Pearson correlations of CRISPR screen data between biological replicates from the high and low fractions. Data from day 5 or day 7 after transduction were analyzed by MAGeCK-VISPR. c, The beta score and rank of genes analyzed in Fig. 1i. Only the data from day 7 is shown here. d, A volcano plot for gene hits from the screening result at day 5. Data is shown as in Fig. 2d (the left panel). Lists of chromatin regulating enzymes highlighted in the plot (the right panel). Genes are included in these categories only in the case where they are expressed in eHAP cells (TPM ≧10) and corresponding sgRNAs are not depleted at day 7.
Extended Data Fig. 5 Validation of screen hits.
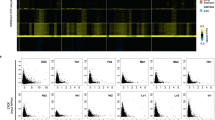
a, ATAC-see analysis with indicated cell lines as shown in Fig. 2c. b, Distribution of essential scores and proportion of the essential genes (the left and right panels) for all genes and indicated screen hits. c, ATAC-see analysis measured by FACS (the left panel), annexin V staining (the middle panel), and bright field images with 30 µm scale bars (the right panel) under indicated conditions. Ctrl: control, H2O2: Hydrogen peroxide, Rap: Raptinal, Stau: Staurosporine. d, eHAP cells were cultured with or without QVQ-OPh during the knockout, and subsequently assayed for ATAC-see and survival rates (the left and right panels). e, ATAC-see analysis measured by FACS on knockouts for indicated non-essential and essential genes selected from the non-hit group. Essential score for each gene is displayed at bottom. All the samples showed statistically non-significant (P > 0.05). f, eHAP or HeLa cells were treated with 1 µM SAHA, for 8 hrs and subjected to ATAC-see measured by FACS (the top panel) and western blotting with indicated antibodies (the bottom panel). g, Representative genome browser tracks of spike-in calibrated ATAC-seq on control and SMARCA4 knockout eHAP cells. h, Relative mean ATAC-seq signals around merged ATAC-seq peaks between control and SMARCA4 KO. i, The left panel, the total number of ATAC-seq peaks detected in indicated knockouts. The right panels, schematic illustration showing classification of differentially accessible regions (DARs, top) and the number of DARs in SMARCA4 knockout eHAP cells (bottom). j, Gene enrichment analyses with gene hits. Statistics, hypergeometric test computed using g:Profiler. n in a, c-f indicates number of cells analysed. x in c and d indicates number of independent experiments. Statistics in the right panels of c and d: a two-tailed unpaired Student’s t-test. Statistics and the definition of box plots for ATAC-see assays in a, c-f, and i are the same as Fig. 1i. Data in the right panels of c and d are presented as mean values +/− SD. *: P < 0.05.
Extended Data Fig. 6 Validation of ATAC-seq analyses for knockout of the selected screen hits.
a, Knockouts of indicated genes (colored in red) were confirmed by western blotting. Arrowheads indicate specific bands and asterisks indicate non-specific bands. These experiments were reproduced twice. b, Results of CRISPR screen for the selected screen hits at day 5 and day 7. Left panels for each day show log2 fold changes of 4 sgRNAs designed to the indicated positive (blue) and negative regulators (red) analyzed by ATAC-seq in Fig. 3. Volcano plots for these genes are shown as Fig. 2d (the right panels). c, Schematic illustrations displaying the quantitative ATAC-seq with spike-in control. d, Numbers of total ATAC-seq peaks in the indicated knockout conditions. e, Pearson correlation between two biological replicates. Normalized read enrichments within TSS proximal regions (TSS + /− 1 kb, 22987 RefSeq genes) were analyzed. f, Pairwise Pearson correlation of ATAC-seq samples based on normalized read enrichments in the consensus peak set (n = 256,617). g, PCA clustering analysis of ATAC-seq data. h, ATAC-qPCR analysis for TFDP1 KO (the left panels) and HNRNPU KOs (the right panels) to confirm differential accessibility at DARs identified by ATAC-seq analysis. Genome browser tracks for ATAC-seq of control and the indicated knockout cells at corresponding DARs are shown on the right side. The DAR categories are shown at the bottom of corresponding browser tracks. Statistics: a one-side unpaired Student’s t-test. *: P < 0.05. NS, not significant. n, independent experiments, Data of ATAC-qPCR are presented as mean values +/− SD.
Extended Data Fig. 7 Extended ATAC-seq analyses for DARs of each knockout.
a, Enrichments of genomic elements (upper half) and chromatin signatures (lower half) at DARs categorized as Fig. 3a in indicated knockouts. b, Proportion of genomic features in conserved-up and conserved-down DARs in indicated knockouts. c, Enrichments of transcription factor motifs at the opened and closed DARs in indicated knockouts. d-f, Representative genome browser tracks of ATAC-seq analysis of the control and the indicated knockouts. The DARs are highlighted in grey. The GC contents of the indicated regions are shown in a violin plot (the bottom of d). Red bars show the medians. The box plots are shown as Fig. 1c. Aggregation plots at the bottom of e and f indicate normalized density of insertion sites at the indicated genomic sites. g, Box plots show proportion of GC content of indicated DARs in each knockout. The width of the boxes indicates a relative number of DARs. The box plots are shown as Fig. 1c. n, number of DARs. h, Motif enrichment analysis of the DARs. Each dot represents significantly enriched motif. x-axis, logFC of ATAC-seq signals at the DARs associated with enriched motif. y-axis, significance of motif enrichment. Motifs with similar sequences are marked with dashed circles.
Extended Data Fig. 8 Characterization of TFDP1 knockout cells.
a, Growth curves of eHAP cells transfected with either non-targeting (control) or TFDP1 sgRNAs. b, Western blotting with the indicated cell lines transfected with sgRNA (for eHAP, HeLa and 293 T cells) and siRNA (for TIG3, NIH3T3, and mES cells) against non-targeting control (Ctrl), TFDP1 (TFDP1-depletion) or TFDP2 (TFDP2-depletion). Arrowheads indicate specific bands and asterisks indicate non-specific bands. These experiments were reproduced twice. c, Growth curves of eHAP cells transfected with either non-targeting (control) or both TFDP1 and TFDP2 sgRNAs (the left panel). Relative cell number of mES cells 4 days after the transfection with indicated siRNA (the right panel). Statistics: a two-tailed unpaired Student’s t-test. NS: non-significant. d, ChIP-qPCR analysis for the putative E2F4 binding sites of indicated gene loci. eHAP cells were cultured with or without 5 µM HLM006474. (the left panel). Growth curves of eHAP cells treated with the indicated concentration of HLM006474 (the right panel). Statistics: a two-tailed unpaired Student’s t-test. e, Averaged nucleosome occupancy of the control and TFDP1 KO at positions across 300% of coding regions (the left panels) or within 2 kb of TSS of all genes, high-, low-, and non-expressed genes (the right panels). The expression status (High, Low, or no expression) is categorized based on transcripts per kilobase million (TPM). f, Normalized MNase-seq read counts of distances between neighboring nucleosomes (the top panel). Linear fits to the phase peaks calculated from the data in the top are shown with resulting NRLs (the bottom panel). g, Gene enrichment analysis on RNA-seq data for differentially expressed genes upon TFDP1 KO. P-values were computed by representation analysis using Metascape. h, A MA plot for the comparison between TFDP1 KO and control cells with gene spots for up- (red spots), down-regulated genes (blue spots), screen hit genes (green spots), and TFDP1 (an orange spot). Data in a, c-d are presented as mean values +/− SD. n in a, c and d, independent experiments.
Extended Data Fig. 9 Characterizations of FLAG or mAID knock-in TFDP1 eHAP cells.
a, Schematic diagram showing the generation of FLAG-TFDP1 knock-in eHAP cells. The targeting cassette was knocked-in to TFDP1 locus by CRISPR-Cas9 mediated homologous recombination. Representative clone was further transfected with a plasmid expressing Cre recombinase to flip out Puromycin resistance cassette, resulting in FLAG-TFDP1 knock-in cells. b, Western blotting for indicated proteins in wild-type and FLAG-TFDP1 eHAP cells. c, TFDP1 knockout eHAP cells (TFDP1 KO) were transfected with either an empty or FLAG-TFDP1 expressing plasmid. 3 days after the transfection, cells were analyzed by ATAC-see (the top panel). The statistical test and definition of the box plot are the same as in Fig. 1i. *: P < 0.05. Western blotting for TFDP1 and α-Tubulin are shown in the bottom panels. d, Top 5 transcription factor-binding motifs enriched in TFDP1 binding sites. Statistics, Hypergeometric test computed using Pscan-ChIP. e, Heat maps of the indicated chromatin profiles aligned at ± 5,000 bp of TSSs in 22,987 individual RefSeq genes, sorted by TFDP1 ChIP-seq enrichment levels. Expression levels of corresponding genes analysed by RNA-seq are shown in the right side. f, Pearson correlation between enrichments of indicated factors on ATAC-seq peaks. g, Schematic diagram representing the generation of mAID-TFDP1 knock-in cells (the left panel). Acute depletion of TFDP1 upon adding 5Ph-IAA for 6 hrs was confirmed by western blotting (the right panel). Experiments in b and g were reproduced three times.
Extended Data Fig. 10 TFDP1 transcriptionally regulates canonical histones.
a, Relative expression levels of canonical histones calculated from total RNA-seq of wild-type and TFDP1 KO eHAP cells. Only expressed histone genes (LogCPM>2) are shown here. b, Quantitative fluorescent western blotting analyses (left) and CBB staining quantification (right) in either whole cell extracts or chromatin fractions of wild-type and TFDP1 KO eHAP cells. Amount of genomic DNA was used as an internal control for calibrating sample variations. Statistical comparison; a two-side unpaired Student’s t-test. *: P < 0.05. n, independent experiments. Data are presented as mean values +/− SD. c, Relative expression levels of mRNA of the indicated non-canonical histone variants in control and TFDP1 KO cells. Statistics: a two-tailed unpaired Student’s t-test. NS: non-significant. *: P < 0.05. n, independent experiments. Data are presented as mean values +/− SD. d, Genome browser track of TFDP1 ChIP-seq at a histone cluster HIST1 locus in the indicated human cell lines. e, Proportion of the indicated canonical histone gene loci occupied by TFDP1. The total numbers of analyzed genes for each canonical histone are shown in the right side.
Supplementary information
Supplementary Information
Supplementary Tables 1–3, a list of source data, Notes 1–3, Extended Data figure captions, Methods and References.
Supplementary Table 1
Materials used in this study, including oligonucleotides, adapter DNAs, sgRNAs, viruses, siRNAs, plasmid DNAs, antibodies and cell lines.
Supplementary Table 2
Results of CRISPR screening.
Supplementary Table 3
Results of RNA-seq on TFDP1 knockout.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots and CBB-stained gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ishii, S., Kakizuka, T., Park, SJ. et al. Genome-wide ATAC-see screening identifies TFDP1 as a modulator of global chromatin accessibility. Nat Genet 56, 473–482 (2024). https://doi.org/10.1038/s41588-024-01658-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-024-01658-1
This article is cited by
-
The regulatory landscape of chromatin accessibility
Nature Reviews Genetics (2024)