Abstract
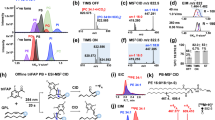
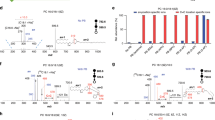
Mass spectrometry-based quantitative lipidomics is an emerging field aiming to uncover the intricate relationships between lipidomes and disease development. However, quantifying lipidomes comprehensively in a high-throughput manner remains challenging owing to the diverse lipid structures. Here we propose a diazobutanone-assisted isobaric labelling strategy as a rapid and robust platform for multiplexed quantitative lipidomics across a broad range of lipid classes, including various phospholipids and glycolipids. The diazobutanone reagent is designed to conjugate with phosphodiester or sulfate groups, while accommodating various functional groups on different lipid classes, enabling subsequent isobaric labelling for high-throughput multiplexed quantitation. Our method demonstrates excellent performance in terms of labelling efficiency, detection sensitivity, quantitative accuracy and broad applicability to various biological samples. Finally, we performed a six-plex quantification analysis of lipid extracts from lean and obese mouse livers. In total, we identified and quantified 246 phospholipids in a high-throughput manner, revealing lipidomic changes that may be associated with obesity in mice.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study are available in the Supplementary Information and source data of this article. Raw data files were uploaded to the MassIVE database (massive.ucsd.edu/ProteoSAFe/static/massive.jsp) under accession number MSV000091941. The phospholipid database, LipidBlast, is available at prime.psc.riken.jp/compms/msdial/main.html#MSP. Source data are provided with this paper.
Code availability
Scripts are available on Zenodo (https://doi.org/10.5281/zenodo.8417892). The script was utilized for the identification and quantification of phospholipids and sulfated glycolipids. Briefly, the converted raw data in mzXML format undergo an exact mass match against a lipid database. The MS2 spectra of the matched precursors are then extracted and screened for both diagnostic and acylium ions. Finally, the abundance of reporter ions is extracted from the MS2 spectra of the identified lipids.
References
Blanksby, S. J. & Mitchell, T. W. Advances in mass spectrometry for lipidomics. Annu. Rev. Anal. Chem. 3, 433–465 (2010).
Meer, G., van, Voelker, D. R. & Feigenson, G. W. Membrane lipids: where they are and how they behave. Nat. Rev. Mol. Cell. Biol. 9, 112–124 (2008).
Tobias, F., Pathmasiri, K. C. & Cologna, S. M. Mass spectrometry imaging reveals ganglioside and ceramide localization patterns during cerebellar degeneration in the Npc1−/− mouse model. Anal. Bioanal. Chem. 411, 5659–5668 (2019).
Barrientos, R. C. & Zhang, Q. Recent advances in the mass spectrometric analysis of glycosphingolipidome – a review. Anal. Chim. Acta 1132, 134–155 (2020).
Fu, S. et al. Aberrant lipid metabolism disrupts calcium homeostasis causing liver endoplasmic reticulum stress in obesity. Nature 473, 528–531 (2011).
Fadeel, B. & Xue, D. The ins and outs of phospholipid asymmetry in the plasma membrane: roles in health and disease. Crit. Rev. Biochem. Mol. Biol. 44, 264–277 (2009).
Salita, T., Rustam, Y. H., Mouradov, D., Sieber, O. M. & Reid, G. E. Reprogrammed lipid metabolism and the lipid-associated hallmarks of colorectal cancer. Cancers 14, 3714 (2022).
Han, X. The emerging role of lipidomics in prediction of diseases. Nat. Rev. Endocrinol. 18, 335–336 (2022).
Danne-Rasche, N., Coman, C. & Ahrends, R. Nano-LC/NSI MS refines lipidomics by enhancing lipid coverage, measurement sensitivity, and linear dynamic range. Anal. Chem. 90, 8093–8101 (2018).
Unsihuay, D. et al. Imaging and analysis of isomeric unsaturated lipids through online photochemical derivatization of carbon–carbon double bonds. Angew. Chem. Int. Ed. Engl. 60, 7559–7563 (2021).
Randolph, C. E., Blanksby, S. J. & McLuckey, S. A. Toward complete structure elucidation of glycerophospholipids in the gas phase through charge inversion ion/ion chemistry. Anal. Chem. 92, 1219–1227 (2020).
Ma, X. et al. Identification and quantitation of lipid C=C location isomers: a shotgun lipidomics approach enabled by photochemical reaction. Proc. Natl Acad. Sci. USA 113, 2573–2578 (2016).
Jung, H. R. et al. High throughput quantitative molecular lipidomics. Biochim. Biophys. Acta 1811, 925–934 (2011).
Leng, J. et al. A highly sensitive isotope-coded derivatization method and its application for the mass spectrometric analysis of analytes containing the carboxyl group. Anal. Chim. Acta 758, 114–121 (2013).
Sivanich, M. K., Gu, T., Tabang, D. N. & Li, L. Recent advances in isobaric labeling and applications in quantitative proteomics. Proteomics 22, 2100256 (2022).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Xiang, F., Ye, H., Chen, R. B., Fu, Q. & Li, L. J. N,N-Dimethyl leucines as novel isobaric tandem mass tags for quantitative proteomics and peptidomics. Anal. Chem. 82, 2817–2825 (2010).
Hahne, H. et al. Carbonyl-reactive tandem mass tags for the proteome-wide quantification of N-linked glycans. Anal. Chem. 84, 3716–3724 (2012).
Gu, T.-J., Feng, Y., Wang, D. & Li, L. Simultaneous multiplexed quantification and C=C localization of fatty acids with LC-MS/MS using isobaric multiplex reagents for carbonyl-containing compound (SUGAR) tags and C=C epoxidation. Anal. Chim. Acta 1225, 340215 (2022).
Yang, T. et al. Lipid mass tags via aziridination for probing unsaturated lipid isomers and accurate relative quantification. Angew. Chem. Int. Ed. Engl. 61, e202207098 (2022).
Laayoun, A. et al. Aryldiazomethanes for universal labeling of nucleic acids and analysis on DNA chips. Bioconjugate Chem. 14, 1298–1306 (2003).
Bourget, C. et al. Biotin-phenyldiazomethane conjugates as labeling reagents at phosphate in mono and polynucleotides. Bioorgan. Med. Chem. 13, 1453–1461 (2005).
Furuta, T. et al. Versatile synthesis of head group functionalized phospholipids via oxime bond formation. Org. Lett. 10, 4847–4850 (2008).
Mix, K. A., Aronoff, M. R. & Raines, R. T. Diazo compounds: versatile tools for chemical biology. ACS Chem. Biol. 11, 3233–3244 (2016).
Takahashi, T. & Suzuki, T. Role of sulfatide in normal and pathological cells and tissues. J. Lipid Res. 53, 1437–1450 (2012).
Wasslen, K. V., Canez, C. R., Lee, H., Manthorpe, J. M. & Smith, J. C. Trimethylation enhancement using diazomethane (TrEnDi) II: rapid in-solution concomitant quaternization of glycerophospholipid amino groups and methylation of phosphate groups via reaction with diazomethane significantly enhances sensitivity in mass spectrometry analyses via a fixed, permanent positive charge. Anal. Chem. 86, 9523–9532 (2014).
Wei, F. et al. Rapid profiling and quantification of phospholipid molecular species in human plasma based on chemical derivatization coupled with electrospray ionization tandem mass spectrometry. Anal. Chim. Acta 1024, 101–111 (2018).
Neumann, E. K., Comi, T. J., Rubakhin, S. S. & Sweedler, J. V. Lipid heterogeneity between astrocytes and neurons revealed by single‐cell MALDI‐MS combined with immunocytochemical classification. Angew. Chem. Int. Ed. Engl. 131, 5971–5975 (2019).
Kind, T. et al. LipidBlast in silico tandem mass spectrometry database for lipid identification. Nat. Methods 10, 755–758 (2013).
Saito, K. et al. Characterization of hepatic lipid profiles in a mouse model with nonalcoholic steatohepatitis and subsequent fibrosis. Sci. Rep. 5, 12466 (2015).
Peng, H. et al. Offspring NAFLD liver phospholipid profiles are differentially programmed by maternal high-fat diet and maternal one carbon supplement. J. Nutr. Biochem. 111, 109187 (2023).
Vucenik, I. & Stains, J. P. Obesity and cancer risk: evidence, mechanisms, and recommendations. Ann. N.Y. Acad. Sci. 1271, 37–43 (2012).
Arendt, B. M. et al. Nonalcoholic fatty liver disease is associated with lower hepatic and erythrocyte ratios of phosphatidylcholine to phosphatidylethanolamine. Appl. Physiol. Nutr. Metab. 38, 334–340 (2013).
Koekemoer, A. L. et al. Large-scale analysis of determinants, stability, and heritability of high-density lipoprotein cholesterol efflux capacity. Arterioscler. Thromb. Vasc. Biol. 37, 1956–1962 (2017).
Jordy, A. B. et al. Analysis of the liver lipidome reveals insights into the protective effect of exercise on high-fat diet-induced hepatosteatosis in mice. Am. J. Physiol. Endocrinol. Metab. 308, E778–E791 (2015).
Nam, M. et al. Lipidomic profiling of liver tissue from obesity-prone and obesity-resistant mice fed a high fat diet. Sci. Rep. 5, 16984 (2015).
Matsuzaka, T. et al. Crucial role of a long-chain fatty acid elongase, Elovl6, in obesity-induced insulin resistance. Nat. Med. 13, 1193–1202 (2007).
Scheja, L. et al. Liver TAG transiently decreases while PL n-3 and n-6 fatty acids are persistently elevated in insulin resistant mice. Lipids 43, 1039–1051 (2008).
Ellett, J. D., Evans, Z. P., Zhang, G., Chavin, K. D. & Spyropoulos, D. D. A rapid PCR‐based method for the identification of ob mutant mice. Obesity 17, 402–404 (2009).
Acknowledgements
This study was supported in part by grant funding from the NIH (R01AG052324, R01AG078794, P41GM108538, and R01DK071801). We also appreciate funding support from a Washington University/University of Wisconsin Diabetes Research Center pilot and feasibility grant (P30DK020579). L.L. acknowledges the funding support of NIH shared instrument grants (NIH-NCRR S10RR029531, S10OD028473 and S10OD025084), a Vilas Distinguished Achievement Professorship and the Charles Melbourne Johnson Distinguished Chair Professorship with funding provided by the Wisconsin Alumni Research Foundation and University of Wisconsin-Madison School of Pharmacy. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the paper.
Author information
Authors and Affiliations
Contributions
T.-J.G. and L.L. developed the initial concept. T.-J.G. and P.-K.L. designed the research and performed most of the experiments and data analysis. Y.-W.W. and T.-J.G. wrote R code and conducted statistical analysis. M.T.F. and D.B.D. prepared mouse disease models. Y.L. prepared cell cultures. S.X. provided experimental feedback and assistance. T.-J.G., P.-K.L. and L.L. discussed the results and wrote the paper. M.T.F., S.X. and Y.L. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The University of Wisconsin-Madison has filed a patent application (18/518,234) based on this work. L.L., T.-J.G. and P.-K.L. are named as inventors on this provisional patent application. The other authors declare no competing interests.
Peer review
Peer review information
Nature Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14, Tables 1 and 2 and Methods.
Supplementary Data 1
Details of PL acyl chain and species.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Intensity of MS1 and MS2 spectra.
Source Data Fig. 5
Statistical source data and details of PL species.
Source Data Fig. 6
Statistical source data and details of PL species.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gu, TJ., Liu, PK., Wang, YW. et al. Diazobutanone-assisted isobaric labelling of phospholipids and sulfated glycolipids enables multiplexed quantitative lipidomics using tandem mass spectrometry. Nat. Chem. (2024). https://doi.org/10.1038/s41557-023-01436-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41557-023-01436-2