Abstract
Use of vertical farms is increasing rapidly as it enables year-round crop production, made possible by fully controlled growing environments situated within supply chains. However, intensive planting and high relative humidity make such systems ideal for the proliferation of fungal pathogens. Thus, despite the use of bio-fungicides and enhanced biosecurity measures, contamination of crops does happen, leading to extensive crop loss, necessitating the use of high-throughput monitoring for early detection of infected plants. In the present study, progression of foliar symptoms caused by Pythium irregulare-induced root rot was monitored for flat-leaf parsley grown in an experimental hydroponic vertical farming setup. Structural and spectral changes in plant canopy were recorded non-invasively at regular intervals using a 3D multispectral scanner. Five morphometric and nine spectral features were selected, and different combinations of these features were subjected to multivariate data analysis via principal component analysis to identify temporal trends for early segregation of healthy and infected samples. Combining morphometric and spectral features enabled a clear distinction between healthy and diseased plants at 4–7 days post inoculation (DPI), whereas use of only morphometric or spectral features allowed this at 7–9 DPI. Minimal datasets combining the six most effective features also resulted in effective grouping of healthy and diseased plants at 4–7 DPI. This suggests that selectively combining morphometric and spectral features can enable accurate early identification of infected plants, thus creating the scope for improving high-throughput crop monitoring in vertical farms.
Similar content being viewed by others
Introduction
Vertical farming facilitates year-round crop production, with higher yields per unit area and more efficient utilisation of resources such as land, water, and nutrients than conventional farming (Rajan et al., 2019; Specht et al., 2014). Such systems employ artificial lighting and soilless culture techniques to maximize crop production under controlled environmental conditions (van Delden et al., 2021). Since vertical farms do not require large expanses of land, production points can be located closer to consumers on brownfield sites, potentially reducing the carbon footprint resulting from food transport (van Delden et al., 2021). Further, physical isolation from the external environment provides a high level of biosecurity, reducing the risk of plant pests, diseases, and pathogens associated with human food-borne illness compared to open field production systems.
Despite the presence of physical barriers, achieving complete exclusion of pathogens from vertical farms is practically challenging. Consequently, crop losses from unintentional bacterial or fungal contamination may occur due to inefficient phytosanitary measures, inadequately treated seeds and substrates, or air-borne spores introduced through the ventilation system (Roberts et al., 2020). In addition, the combination of high-density production environment with recirculation of nutrient solutions within vertical farms exacerbates the situation by facilitating rapid pathogen spread. Furthermore, high humidity and favourable ambient temperature supports pathogen proliferation in such systems (Paulitz, 1997; Roberts et al., 2020). Thus, there is a high risk of extensive crop loss when biosecurity measures are breached in vertical farms. Hence, early detection of diseased plants is crucial for controlling pathogen spread and minimising the risk of crop loss in vertical farms.
Conventionally, identification and scoring of plant diseases is performed by experts via visual inspection of samples (Bock et al., 2010). However, this approach is reliant on the presence of visual symptoms, which can be subjective, inconsistent across practitioners, and inadequate for detecting diseases during their latent period when the pathogens are spreading without causing visible symptoms. Further, the process is time-consuming and labour-intensive, and is thus, practically feasible only for small sample sizes. In addition, large-scale hydroponic farms often maintain strict restrictions on the access of personnel around growing benches to ensure biosafety, making visual scoring highly improbable. As an advancement to manual inspection, molecular methods have revolutionized plant disease diagnosis, accelerating pathogen detection and facilitating the screening of asymptomatic plants, making the process less subjective (Martinelli et al., 2015). However, such techniques require specialised infrastructure and experienced laboratory personnel, and are expensive to perform routinely, making it difficult for widespread use in commercial cultivation. In addition, the process would be destructive, and would be implementable on just a few samples in every batch of plants. Hence, the necessity of an alternative easy-to-use/automatable technology for crop monitoring has driven the development of machine vision-based techniques for identifying characteristic stress symptoms in plants to allow real-time assessment of diseases and health status. These methods can be deployed to collect data in real-time without generating significant additional costs.
Plant diseases typically induce various visible morphological changes, such as stunted growth and reduced canopy cover, as well as spectral aberrations associated with disease symptoms, such as generalised or localised lesions, tissue chlorosis, and necrosis (Mutka & Bart, 2015). Plant image analysis for disease detection focuses on finding these irregularities in growth patterns and spectral properties. In this context, multispectral imaging, especially RGB imaging, has emerged as a low-cost, high-throughput alternative for assessing plant health by identifying abnormalities in leaf colour features (Agarwal et al., 2021; Martinelli et al., 2015; Waiphara et al., 2022). In addition, studies have also attempted to highlight the efficacy of 3D imaging for monitoring structural changes in plants under stress (Husin et al., 2020; Su et al., 2019).
Application of advanced data processing techniques such as multivariate data analysis (MDA) and machine learning (ML) for plant image analysis has played a crucial role making in-depth assessment of image features feasible (Singh et al., 2021). Further, tremendous progress has been made over the past decade in the domain of image-based plant disease detection by the implementation of deep-learning, a more advanced information processing tool that can “learn” to recognise data patterns like a human brain (Nagaraju & Chawla, 2020). Although highly accurate, this approach involves computationally intensive algorithms that process thousands of pre-existing plant images to “train” the prediction model, and have predominantly been tested with RGB data. Further, suitability of such plant image analysis approaches for non-destructive monitoring of plant responses to biotic stressors has been investigated in detail mostly for crops grown in field conditions (Mutka & Bart, 2015; Singh et al., 2021; Zhang et al., 2019), while reports for hydroponic vertical farming systems are very limited.
Prevalence of root diseases is a major concern for commercial hydroponic operations (Suárez-Cáceres et al., 2021). Zoosporic and wilt pathogen species such as Pythium, Phytophthora, and Fusarium have been most frequently reported to cause vascular and root diseases in various crops grown hydroponically (Paulitz, 1997; Suárez-Cáceres et al., 2021). In particular, root rot and damping-off due to various Pythium spp. is a serious issue affecting hydroponic production of various Apiaceae crops, including flat-leaf parsley (Petroselinum crispum var. neapolitanum), a popular culinary herb (Minchinton et al., 2013). Early detection of root diseases is more challenging compared to foliar diseases as it is difficult to actively monitor roots, where the symptoms initially occur. However, morphological and spectral changes that appear on the canopy could be used as indirect indicators of root infections and be monitored for detecting such diseases (Salgadoe et al., 2018).
In this study, we examined changes in morphometric and spectral attributes of flat-leaf parsley upon infection with Pythium irregulare in an experimental hydroponic vertical farming system. We aimed to ascertain the post-infection window for reliable early distinction of diseased plants using concurrent 3D and multispectral imaging. As an exploratory assessment, the scope of segregating infected plants from the healthy ones by using morphometric and spectral attributes independently and simultaneously was evaluated via multivariate data analysis using principal component analysis (PCA), along with step-wise variable reduction to determine the efficacy of minimal datasets for reliable comparison between infected and control samples at early stages of infection.
Material and methods
Plant growth trials
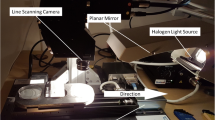
Flat-leaf parsley seedlings were raised from seeds in coco-peat plugs (Van der Knapp, The Netherlands) following dark germination under sterile conditions in a nursery chamber (Aralab-InFarm UK Ltd., London, UK). A seedling density of ~ 25 seedlings/plug was maintained to replicate commercial production standards. After reaching a height of ca. 2 cm, the seedlings were transferred to a customised experimental vertical farming setup (Fig. 1) having “deep water culture” hydroponic units in a growth chamber with regulated environment (Newcastle University, Newcastle upon Tyne, UK). Each hydroponic unit comprised of a polypropylene container (dark grey, opaque; inner dimensions: L × W × H 56 × 36 × 11 cm) with a tray-lid (white, opaque) having a 7 × 4 array of circular empty slots. The container was filled with 18 L commercial hydroponics solution. Each unit received 26 seedling plugs, and a submersible water pump was used for root aeration along with a circulating water bath for maintaining constant water temperature. Plants were grown for 20 days at a temperature of 22 ± 1 °C and 75 ± 5% relative humidity, under 300–350 µmol.m−2 s−1 broad spectrum LED lighting (L28–NS12, Valoya Ltd., Finland) following a 16/8 h day-night cycle. Two experimental trials were carried out as follows: 1) preliminary trial to identify effects of infection on the morphometric and spectral attributes over time; 2) main trial to select the features and perform multivariate data analysis for temporal plant monitoring. In each trial, two units received P. irregulare inoculum (described later), and two were used as control.
Pathogen isolation and inoculation
P. irregulare was isolated from diseased plants growing in a commercial hydroponic vertical farm, and cultured on PARP + B (corn meal agar with pimaricin, 5 mg/L; ampicillin, 250 mg/L; rifampicin, 10 mg/L; pentachloronitrobenzene, 50 mg/L; and benomyl 10 mg/L) semi-selective medium (Matthiesen et al., 2016). Identity of the isolate was confirmed following PCR-based amplification of specific ITS and COX II genes (Online resource 1: Table S1) as reported earlier (Martin, 2000; White et al., 1990). The pathogen was grown in bulk using clarified-V8 broth, with minor modifications to the protocol described by McGehee et al. (2019). Briefly, a 4-mm plug of the PARP+B medium inoculated with the pathogen was transferred to a sterile Petri dish. Subsequently, the Petri dish was filled with 20 mL of V8 broth, and incubated in the dark for 5 days at 25 °C. Subsequently, the mycelial mats were liquified in ddH2O for 2 min to obtain a final concentration of ~ 1 × 104 mycelial fragments/mL. The resulting slurry was used to inoculate the seedling plugs at 1 mL/plug at 5 days post nursery for the preliminary trial, and at 4 days post nursery in the main trial to induce the symptoms of infection earlier. Further, in the main trial two samples were randomly selected from each inoculated tray at 8 days post inoculation (DPI) to confirm the presence of pathogen using the same PCR-based method.
Multispectral 3D scanning
Sample trays were individually scanned using a PlantEye F500 multispectral LiDAR scanner (Phenospex, The Netherlands, www. phenospex.com) to simultaneously record the spectral reflectance and 3D features of the plant canopy at 8, 11, and 15 DPI in the preliminary trial, and at 2, 4, 7, 9, and 11 DPI in the main trial. The device comprised of an overhead scanning unit equipped with in-built blue (B; λ = 460–485 nm), green (G; λ = 530–540 nm), red (R; λ = 620–645 nm), and near-infrared (NIR; λ = 720–750 nm) LEDs for sample illumination along with corresponding sensors for multispectral imaging, placed adjacent to a LiDAR laser source (λ = 940 nm) with a sensor for 3D imaging, and a horizontal platform for placing the sample trays (Fig. 2). The scanner moved along a horizontal track from one end of the platform to the other while scanning. A white metallic reference plate supplied with the device was placed at the starting point to assist in spectral and LiDAR sensor calibration. Plants were scanned maintaining a fixed distance of 100 cm between the scanner and the tray top, and the scanner moved at a fixed speed of 50 mm/s (Y-axis). This provided an approximate resolution of 0.7 mm X-axis, 1 mm Y-axis, and 0.2 mm Z-axis. The scans were processed immediately by the in-built HortControl software (Phenospex), wherein the spectral and LiDAR information were automatically superimposed based on internal calibrations, creating point-cloud (.ply) data files which contained the spatial (X, Y, Z) and spectral (R, G, B, NIR) values of each pixel. No additional light sources were used while scanning to ensure uniformity in spectral information recorded on different days, and nullifying the need for further spectral calibrations.
Schematic layout of the imaging setup for 3D-cum-multispectral scanning (left) and a sample output point-cloud image (right). The imaging setup comprised of an overhead scanner equipped with red, green, blue, and near-infrared LEDs along with corresponding sensors for multispectral imaging, as well as a LiDAR sensor for 3D imaging. The scanner moved along a horizontal track from one end of the platform to the other while scanning the samples, and the reference plate assisted in spectral and positional calibration. Boxes shown in the point-cloud image (right) correspond to individual samples (~ 10,000 data points) with a false-colour scheme being used to depict the data from one spectral channel
Data pre-processing and feature extraction
Each scan was divided into a 7 × 4 array of identical sectors within the HortControl software as per the tray layout (Figs. 1 and 2) for obtaining sample-wise morphometric and spectral data. Each sector covered ca. 10,000 data points pertaining to each sample, of which at least 2,000 data points were used for calculating each plant feature depending on sample canopy size. The software automatically triangulated the spatial coordinates to calculate morphometric features, and the multispectral data to calculate various spectral indices. This generated nine morphometric parameters as follows: mean plant height, maximum plant height, total leaf area (TLA), leaf area index (LAI), projected leaf area (PLA), digital biomass (DB), leaf angle, leaf inclination (LInc), and light penetration depth (LPD) (Table 1). Likewise, five spectral indices were generated by the software as follows: Green Leaf Index (GLI, [(2 × G)-R-B]/[(2 × G) + R + B]), Hue, Normalised Difference Vegetation Index (NDVI, [NIR-R]/[NIR+R]), Normalized Pigment Chlorophyll ratio Index (NPCI, [R-B]/[R+B]), and Plant Senescence Reflectance Index (PSRI, [R-G]/NIR). Spectral information was augmented by extracting the raw R, G, B, and NIR reflectance data from the point-cloud files using Python programming (www.python.org) and by calculating R/G, G/R, R+G+B, R+G-B, R+G, G-minus-R (GMR) (Agarwal & Dutta Gupta, 2018), and augmented green-red index (AGRI, [G-R] × G/R).
PCA for analysing temporal patterns in plant features
PCA was performed using diverse datasets for each interval individually to visualise the temporal trends in different features and how it influenced the segregation of control and infected samples. As a preliminary step for reducing the number of PCA features, all twenty-five morphometric and spectral attributes were subjected to correlation analysis using the Data Analysis ToolPak (Microsoft Excel 365, Microsoft Corp., USA) to identify attributes exhibiting identical linear trends. Attributes with strong linear relations (r < -0.95, r > 0.95) were selectively omitted to minimise redundancy. The attributes that were retained, henceforth referred to as “selected features”, were subjected to PCA to ascertain the possibility of distinguishing between healthy and infected plants at different intervals post infection. PCA was performed in Python using the Scikit-learn module (Pedregosa et al., 2011), at a threshold of > 99% variance explained. The following subsets of selected features were used for PCA at each imaging interval: 1) all morphometric attributes (M_all); 2) all spectral attributes (S_all); 3) all selected features (MS_all). Eigen values and percentage of variance explained were recorded for each principal component (PC), along with the PC loadings of each feature. PC loadings obtained from the MS_all dataset were used to calculate the weighted loading (WL) for each feature at every interval as follows:
Here, Exi and Li indicate proportion of variance explained (0–1) and loading of the feature for the ith PC, respectively, and n indicates the total number of PCs obtained. Features were assigned ranks based on WL values to indicate their performance at each interval; higher WL earned a better rank, and vice versa. Overall performance ranks were assigned as per the mean of ranks obtained across all intervals. Based on this, PCA was performed using the best ranked variables collated in two subsets containing < 25% of the actual features (henceforth referred to as “minimal datasets”) as follows: 1) three best morphometric and three best spectral attributes (MS_3-3); and 2) top six attributes (MS_top6). This was done to assess the impact of total number of features and the balance between morphometric and spectral features on the analysis.
Biplots indicating the loadings and scores for the first two PCs were plotted for each PCA along with the 95% confidence ellipse for each class, i.e., healthy and infected, and the Euclidean distance between the centroids for all five feature subsets at each imaging interval to visualise the temporal trends in unsupervised (without prior labelling) segregation of the two classes.
Statistical analysis
One-way ANOVA was performed using the main trial dataset for all selected features to ascertain the significance of difference between the means of control and infected samples at individual intervals. The process was repeated with PC1 and PC2 scores for each grouped dataset, i.e., M_all, S_all, MS_all, MS_3-3, and MS_top6, following dataset transformation by PCA at each interval to obtain the dissimilarity of both classes in terms of F-statistic. Further, statistical difference of data distribution between the PCA scores of control and infected samples was also assessed by two-sample Kolmogorov–Smirnov (KS) test using the scipy.stats.ks_2samp library in Python (https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.ks_2samp.html), wherein KS = 1 implies no similarity and KS = 0 indicates identical data distribution. Overlap between the PCA scores of control and infected sample clusters was further assessed by calculating the Jaccard index (JI) and the Szymkiewicz–Simpson overlap coefficient (SS) using standard functions in Microsoft Excel 365 (Microsoft Corp.) as follows:
Here, RC and RI indicate the ranges for PCA scores of control and infected samples, respectively, the |R| function calculates the range size, ∩ and ∪ operators find the intersection and union of datasets, respectively, and the min function identifies the smallest range. JI indicates the proportion of overlap across the entire data distribution, with JI = 0 and JI = 1 implying no and complete overlap, respectively. Similarly, SS indicates the proportion of the dataset with a smaller range overlapping with the dataset with bigger range, where SS = 0 implies no overlap and SS = 1 indicates that the dataset with the smaller range lies entirely within the range of the dataset with the bigger range. Interquartile range was used to exclude any outliers while calculating JI and SS to improve the reliability of the results.
Results
Morphometric and spectral attributes
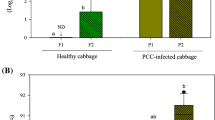
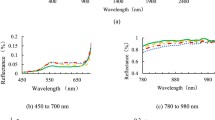
Plant growth and appearance was markedly affected by P. irregulare infection 7 DPI onwards, as observed in both trials (Fig. 3; Online resource 1: Fig. S1). Clear signs of stress and tissue damage were visible on leaves as well as roots (Online resource 1: Fig. S2). Morphometric attributes such as DB, mean plant height, and LAI indicated that growth was stunted in the infected plants, with the difference compared to control becoming more prominent in height and DB with the progression of infection (Fig. 3a–c). Spectral attributes such as Hue, GLI, NDVI, NPCI, and AGRI remained higher in healthy plants relative to the infected samples (Fig. 3f–h, j, n), whereas the opposite trend was observed for PSRI, G, and NIR, i.e., the values for the healthy samples were generally lower than the infected plants (Fig. 3i, k, l). Other features, viz., LInc, LPD, and GMR exhibited significant similarity between the control and infected samples especially at ~ 7 DPI (Fig. 3d, e, m).
Digital biomass (a), plant height (b), leaf area index (c), leaf inclination (LInc; d), light penetration depth (LPD; e), Hue (f), Green Leaf Index (GLI; g), Normalised Difference Vegetation Index (NDVI; h), Plant Senescence Reflectance Index (PSRI; i), Normalised Pigment Chlorophyll ratio Index (NPCI; j), green reflectance (G; k), near-infrared reflectance (NIR; l), green-minus-red reflectance (GMR; m), and Augmented Green-Red Index (AGRI; n) of flat-leaf parsley infected with P. irregulare (main trial). Values have been expressed as mean ± SD (Control: n = 52; Infected: n = 52 for 2–7 DPI, n = 48 for 9 and 11 DPI). “n.s.” indicates no statistically significant difference (p > 0.05) between the mean values of control and infected samples at the specified interval (DPI) following one-way ANOVA
Feature selection for MDA
As per the results of linear correlation analysis (Online resource 1: Table S2), five morphometric and nine spectral features, viz., DB, mean plant height, LAI, LInc, LPD, GLI, Hue, NDVI, NPCI, PSRI, G, NIR, GMR and AGRI, were selected for MDA via PCA. Eleven features, i.e., maximum plant height, leaf angle, TLA, PLA, R, B, R/G, G/R, R+G+B, R+G-B, and R+G, were omitted from subsequent analyses based on their strong correlation (r < -0.95, r > 0.95) with at least one of the selected features (Online resource 1: Table S3).
MDA of all selected features
PCA biplots for analyses using morphometric features (M_all), spectral features (S_all), and both datasets combined (MS_all) revealed the scope of accurately segregating healthy and infected plants by machine vision at later stages of infection, i.e., 7 DPI and higher (Fig. 4; Online resource 1: Fig. S3). Analysing the MS_all dataset enabled better differentiation between the healthy and diseased samples than the M_all and S_all datasets at 7 DPI, as indicated by the 95% confidence ellipse and higher centroid distance. All biplots for subsequent intervals showed this trend, suggesting that M_all and S_all could also be used to reliably identify infected plants at 9 and 11 DPI, i.e., when the effects of infection became more prominent. Cumulative variance explained (CVE) by PC1 and PC2 for the M_all dataset did not exhibit a clear trend, and varied between 76–83% for the five intervals (Online resource 1: Table S4). In contrast, CVE by PC1 and PC2 increased from 2 to 11 DPI for S_all and MS_all datasets, i.e., from 77.09% to 94.54% and from 64.71% to 81.6%, respectively (Online resource 1: Tables S5 and S6), suggesting steady trends with disease progression.
PCA of morphometric and spectral features for healthy (green, diamonds) and infected (red, triangles) plants at 2, 4, and 7 days post inoculation (DPI). Biplots represent the first two principal components (PCs) for the analyses with morphometric attributes (a; M_all, 5 features), spectral attributes (b; S_all, 9 features), and both datasets combined (c; MS_all, 14 features). Values in parentheses indicate the percentage of variance explained by the corresponding PC. Ellipses represent the 95% confidence interval for each class, viz. healthy and infected (n = 52). ΔC indicates the Euclidean distance between the centroids of both groups. Order of the features: DB, Height, LAI, LInc, LPD (a; morphometric); GLI, Hue, NDVI, NPCI, PSRI, G, NIR, GMR, AGRI (b; spectral); morphometric followed by spectral in the same order (c). Plots for 9 and 11 DPI are presented in Online Resource 1: Fig. S3
MDA of minimal datasets
WL values calculated based on the PCA with MS_all dataset revealed a clear variation in the performance of all the selected features at different stages of infection (Table 2). The magnitudes of WL were close to 0.11 and 0.15 at 2 DPI for the lowest and highest ranked features, respectively, indicating relatively small variability in performance amongst the features. The range gradually increased with the duration of infection, and WL values reached 0.08 and 0.22 at 11 DPI for the weakest and best features, respectively, suggesting a clear improvement in the performance of some features. Rankings based on WL values (Table 2) indicated that features such as GLI and NDVI performed well consistently (rank < 5), whereas LPD and LInc had consistently poor performance (rank > 11); the performance of all other features varied markedly at each interval.
Considering the overall performance and ranking, three morphometric features, viz., DB, mean plant height, and LAI, and four spectral features, viz., GLI, NDVI, AGRI, and G, were selected for further analyses using minimal datasets as follows: the MS_3-3 dataset consisted of DB, mean plant height, LAI, GLI, NDVI, and AGRI, whereas the MS_top6 dataset had G instead of LAI. PCA using the minimal datasets yielded effective segregation (95% confidence interval) of the healthy and infected samples at 7 DPI (Fig. 5). Notably, the MS_3-3 dataset resulted in a more compact clustering of the two sample classes compared to the MS_top6 at 9 and 11 DPI (Online resource 1: Fig. S4). CVE by PC1 and PC2 increased from 83.82% to 95.29% over 2 to 11 DPI for MS_3-3 (Online resource 1: Table S7), whereas the values increased from 78.67% to 95.11% for the MS_top6 dataset over the same interval (Online resource 1: Table S8), indicating steady improvement in sample clustering with disease progression.
PCA of minimal datasets with specific morphometric and spectral attributes of healthy (green, diamonds) and infected (red, triangles) plants at 2, 4, and 7 days post inoculation (DPI). Biplots represent the first two principal components (PCs) for analyses performed by combining the three best morphometric and three best spectral features (a, MS_3-3), as well as the top six features (b, MS_top6). Values in parentheses indicate the percentage of variance explained by the corresponding PC. Ellipses represent the 95% confidence interval for each class, viz., healthy and infected (n = 52). ΔC indicates the Euclidean distance between the centroids of both groups. Features are in the order: DB, Height, LAI, GLI, NDVI, AGRI (a, MS_3-3); DB, Height, GLI, NDVI, G, AGRI (b, MS_top6). Plots for 9 and 11 DPI are presented in Online Resource 1: Fig. S4
Similarity and overlap of PCA clusters
Results of the two similarity and two overlap tests indicated that distinctiveness between the PC1 scores of control and infected samples increased from 2 to 11 DPI for all datasets, whereas no clear trend was visible for PC2 scores likely due to its highly variable nature (Table 3). Considering the early stages of infection, F-values of PC1 for M_all and S_all were closer to the F-critical value of 3.938 as compared to the MS_all, MS_3-3, and MS_top6 datasets at 2 DPI, whereas at 4 DPI it was highest for MS_all, followed by MS_top6. Moreover, at 7 DPI all three datasets with morphometric and spectral features combined had F-values > 1.16 times higher than the M_all and S_all datasets. Similarly, a distinct increase in the dissimilarity of data distribution between control and infected samples from 2 to 7 DPI was also evident from the KS values, with a higher KS value indicating greater dissimilarity. Notably, while the S_all dataset had the lowest KS value at 2 DPI, it was lowest for M_all at 4 DPI, although the dissimilarity became very high (KS > 0.9) for all datasets at 7 DPI. Although overlap in data range at 4 DPI was relatively lower for the MS_all and MS_top6 datasets (JI < 0.2, SS < 0.4), it was least for MS_3-3 and MS_top6 datasets for 7 DPI (JI = 0.06, SS < 0.2).
Discussion
Application of machine vision for real-time high-throughput non-invasive monitoring of plant health status and growth has been explored for a wide variety of crops (Bock et al., 2010; Singh et al., 2021; Waiphara et al., 2022; Zhang et al., 2019). In this context, 3D scanners have been implemented for characterising growth and stress-related structural changes in various crops, including maize (Friedli et al., 2016; Su et al., 2019), wheat, soybean (Friedli et al., 2016), peanut (Yuan et al., 2019), oil palm (Husin et al., 2020), sugar beet (Xiao et al., 2020), and potato (Mulugeta Aneley et al., 2022). However, the use of 3D imaging for crop disease detection is still in the conceptualisation stage (Zhang et al., 2019). In contrast, use of multispectral sensors has been investigated in depth for detecting various crop diseases, including sheath blight (Qin & Zhang, 2005), powdery mildew, leaf rust (Franke & Menz, 2007), root rot (Yang et al., 2010), late blight (Sugiura et al., 2016), mosaic virus disease (Raji et al., 2016), huanglongbing (DadrasJavan et al., 2019), and light leaf spot (Veys et al., 2019). Notably, only a small percentage of such studies have attempted to amalgamate the information obtained via simultaneous implementation of both types of sensors (Lazarević et al., 2021; Manavalan et al., 2021).
Herein, we aim to highlight the potential of machine-vision for co-monitoring morphometric and spectral attributes using P. irregulare infection in flat-parsley as a model system to assess the health of crops grown hydroponically in a vertical farm, with emphasis on better segregation of samples based on disease symptoms using MDA following temporal data acquisition. In the current study, multiple exploratory trials were initially conducted using different inoculation methods, inoculum concentrations, and inoculation intervals (data not shown), and were tested to establish the pathogenesis model to be used for the intended analysis. The design yielding the best result (described in the methods section) was considered as the preliminary trial, and was followed in the main trial for temporal MDA. The preliminary trial presented clear indications of shoot and root tissue damage in infected plant samples, and provided a tentative timeframe for attempting early monitoring of disease symptoms (Online resource 1: Fig. S1). In this trial, data was collected from 8 DPI onwards when disease symptoms such as leaf yellowing and slower growth compared to the control became very clear upon visual inspection.
In the main trial, recording morphometric and spectral data intermittently from 2 DPI onwards helped provide a clear picture of how the infection affected the plants over time (Fig. 3). While some parameters such as DB and plant height increased steadily in healthy samples from 2 to 11 DPI, attributes such as LAI, GLI, NDVI, and PSRI plateaued around 4 to 7 DPI. In contrast, the infected samples did not exhibit any considerable change in DB and plant height, whereas a steady decline in GLI and AGRI was recorded after 4 DPI. As recorded in the healthy plants, LAI, NDVI, and PSRI showed plateauing around 4 to 7 DPI in the infected plants as well, although the magnitudes differed considerably between the two groups. Such differences in relative temporal trends of the different imaging attributes highlight the complexity in automatic identification of diseased plants using individual parameters, because the numerical trends and efficacy of disease detection using each attribute may vary with time. However, co-interpretation of feature trends following MDA by PCA helped overcome this limitation.
The selected features were subjected to PCA to obtain a holistic overview of the temporal trends via dimensional reduction. As an initial feature elimination step prior to PCA, eleven out of the twenty-five recorded attributes were omitted based on their strong correlation (r < -0.95, r > 0.95) with one or more selected features (Online resource 1: Tables S2 and S3). This helped minimise redundancy in information, reduced the computational load, and simplified data analysis. In addition, removing redundant variables also reduced the likelihood of biasing feature rankings without affecting the quality of analysis, which would otherwise have a negative impact on subsequent computations and interpretations. This was confirmed by performing PCA with all twenty-five features (data not shown), wherein extensive overlap between highly correlated features was observed, with no significant improvement in the clustering of healthy and infected samples.
Results of PCA using all data subsets, viz., M_all, S_all, MS_all (Fig. 4; Online resource 1: Fig. S3), MS_3-3, and MS_top6 (Fig. 5; Online resource 1: Fig. S4), revealed that the infected samples could be segregated from the control most reliably at later stages, i.e., 7 DPI onwards, when the symptoms became more prominent. Since members of the Pythium genera are root necrotrophs, they preferentially attack the root tissue and cause root rot (Okubara & Paulitz, 2005). This leads to a gradual systematic decline in overall plant health and possibly even death. Considering the indirect impact of Pythium on above-ground plant parts, a delay in the appearance of foliar symptoms is highly likely. Hence, spectral attributes such as GLI, Hue, and AGRI (Fig. 3) present a decline in infected plants from around 4 DPI, which indicates deteriorating plant health. Although a steady increase in the morphometric attributes was noticeable for healthy plants 4 DPI onwards, these attributes did not change considerably in the infected plants (Fig. 3). Nevertheless, multivariate analysis of all datasets enabled a considerable proportion of the infected samples to be distinguished from the healthy samples even at 2 DPI (Figs. 4 and 5). This suggests that samples which got affected more severely could be distinguished via MDA of spectral and morphometric attributes as early as 2 DPI.
Distinguishing infected samples from healthy ones using the absolute values of individual attributes would have been particularly challenging at 2 DPI owing to the considerable overlap between the range of values between the two classes (Fig. 3). In contrast, multivariate data analysis via PCA allowed the variations for all selected features to be assessed simultaneously and be concatenated to provide an overall depiction of the difference between the samples (Figs. 4, 5, Table 3). Dimensionality reduction by PCA further simplified data interpretation by generating PCs, i.e., hypothetical variables formed by linear combinations of all features, which allowed the information from numerous real variables (in this case five to fourteen) to be presented in a two-dimensional biplot. Thus, co-assessing morphometric and spectral attributes via PCA aggregated even minor differences between the healthy and diseased plant samples, which could potentially enable isolation of infected plants based on pre-symptomatic changes.
Abnormalities in plant health may not be clearly perceptible very early if only morphological traits are being used. In addition, a major conundrum related to this is that morphological features such as plant height, leaf area, and dry biomass tend to remain unchanged in stressed plants, which necessitates the presence of a healthy plant for reference. In contrast, alterations in leaf pigmentation, which closely represent changes in plant physiological status, start occurring following the onset of stress and lead to characteristic deviations in spectral properties (Nilsson, 1995). Thus, temporal monitoring of spectral attributes in conjunction with morphometric measurements could provide a better insight into plant health status. This is indicated by the PCA biplots for 4 and 7 DPI (Fig. 4) as well as the comparative statistics for data similarity and overlap (Table 3), wherein MS_all and S_all datasets result in better separation of the healthy and infected plants than M_all.
It is worth mentioning that multispectral measurements allow numerous hypothetical spectral indices to be generated from only a few wavebands, which creates more potential features to be analysed. For instance, in this study reflectance values from four spectral regions, viz., R, G, B, and NIR, were used to calculate twelve theoretical spectral indices. In contrast, since the morphometric features represent “physical” or “tangible” traits, the same level of flexibility for generating hypothetical morphometric features may not be present due to the likelihood of new hypothetical morphometric traits being physically unrealistic or ambiguous. Hence, this limits the scope of expanding the morphometric feature subset by creating additional traits with the available morphometric measurements, as was done for the spectral features.
In the present study, the smaller number of morphometric features (n = 5) compared to spectral features (n = 9) used for MDA could have arguably biased the results in favour of spectral attributes. In addition, the MS_all dataset had a high proportion of spectral attributes, which could have skewed the outcome as well. Thus, in addition to testing the possibility of using a smaller number of features for isolating infected plants accurately, analyses with fewer features (n = 6), i.e., MS_3-3 and MS_top6, were performed to better understand how the total number of features and the balance between morphometric and spectral features impacted the analysis. Similar to the test with MS_all, PCA with both minimal datasets produced strongly non-overlapping clusters (JI = 0.06, SS = 0.13–0.14; Table 3) for the healthy and infected samples at 7 DPI, albeit using less than half the features as MS_all (n = 14 features). Since features having better overall WL in previous analyses (Table 2) were selected for this step, it may be inferred that the quality of features being used was more important than the total number of features.
Notably, the best-ranked features, viz., GLI, NDVI, AGRI, G, DB, and mean plant height (Table 2), showed distinct temporal trends individually, with limited overlap between the two classes (Fig. 3). In contrast, features such as NPCI, LPD, and LInc, which were ranked the lowest based on WL (Table 2), exhibited irregular trends or had considerable overlap between the healthy and infected plants (Fig. 3), which corroborates the selection and exclusion of features based on WL. Further, the analysis was not affected significantly by the ratio of morphometric and spectral features (Fig. 5) as long as features exhibiting clear trends were being used. Notably, all three datasets combining both types of features, i.e., MS_all, MS_3-3, and MS_top6, resulted in better segregation of healthy and infected samples, as indicated by the biplots and cluster comparisons for 7 DPI (Figs. 4, 5, Table 3). This clearly indicates that using a combination of morphometric and spectral attributes improves the scope for early identification of unhealthy plants. Hence, inclusion of morphometric data in addition to multispectral (RGB) data could improve plant disease detection via ML by bringing in a totally diverse set of informative attributes representing plant health status, which could enable more accurate analysis even with fewer total features.
Earlier studies have performed plant image analysis via various ML methods for disease detection (Ahmad et al., 2023; Singh et al., 2021). For instance, ML approaches such as Support Vector Machines, Random Forest, Decision Trees, Naïve Bayes, and K-Nearest Neighbours have been deployed for the identification of diseased samples using images of bell-pepper (Anjna et al., 2020), maize (Panigrahi et al., 2020), tomato (Agarwal et al., 2020; Harakannanavar et al., 2022), and rice (Shrivastava & Pradhan, 2021; Zamani et al., 2022). Further, application of more advanced computational tools such as deep-learning has greatly improved information processing for plant health assessment following data acquisition via machine vision (Ghosal et al., 2018; Nagasubramanian et al., 2019; Yamamoto et al., 2017). A wide variety of deep-learning algorithms implementing unique iterations of convolutional neural networks have also been successfully tested for plant disease detection in various crops, such as cassava (Sambasivam & Opiyo, 2021), tomato (Abbas et al., 2021; Agarwal et al., 2020; Chowdhury et al., 2021; Harakannanavar et al., 2022), maize (Li et al., 2020), peach (Bedi & Gole, 2021), and strawberry (Shin et al., 2021). Moreover, studies have also been carried out to compare the performance of various ML and deep-learning approaches (Harakannanavar et al., 2022; Sujatha et al., 2021). While all these studies focussed on implementing different learning methods for processing RGB images, other studies have reported the use of Linear Discriminant Analysis along with Support Vector Machines for disease detection using thermal and hyperspectral images of winter-wheat (Zhang et al., 2012), olive (Calderón et al., 2015), and almond (López-López et al., 2016).
Although very advantageous for high-throughput plant stress detection, application of intensive ML and deep-learning modelling has certain limitations as well. For instance, various supervised learning models require training samples to be manually annotated by an expert, making the method labour-intensive, subjective, and prone to errors (Singh et al., 2021), and focus on less labour-intensive ML methods is gaining interest. Moreover, a majority of the reports on ML and deep-learning mentioned earlier implemented huge datasets with thousands of images from pre-existing repositories to generate the prediction models, making the process less conducive if a previously-unexamined plant or disease was being monitored. As an example, Sambasivam and Opiyo (2021) and Chowdhury et al. (2021) utilised 10,000+ images in their deep-learning tests to generate robust prediction models. On the other hand, Bedi and Gole (2021) reported a method for reducing the number of useful training parameters to 9914, compared to other studies where more than a million parameters had been used to generate high-performance prediction models (Mohanty et al., 2016; Shin et al., 2021). Further, such reports on the application of deep-learning algorithms implemented “black-box” models (Saleem et al., 2020; Chowdhury et al., 2021; Sambasivam & Opiyo, 2021; Shin et al., 2021; Tiwari et al., 2021), which are difficult to depict, making their interpretation challenging for plant scientists with limited knowledge of advanced ML (Singh et al., 2021).
Owing to the bottlenecks of these computationally intensive learning approaches adopted for plant monitoring, alternative analytical approaches that are more flexible and explicit must be chosen for exploratory studies. Hence, in the present study, a simplified model-free approach employing MDA via PCA was adopted for identifying temporal trends in the canopy features of diseased plants, bypassing the issues mentioned previously. Since the present study was exploratory, with the main aim of understanding and depicting temporal changes in sample segregation by the use of diverse datasets from multiple sensors, PCA-based analyses were chosen to represent the entire process in a more comprehensible manner. A fundamental difference between PCA and intensive supervised classification modelling is that while the former finds the maximum variance in data across multiple variables, the latter can be used to predict sample category (healthy or infected) based on previous datasets, i.e., a trained model. Hence, at least one adequate sample dataset is needed for reference in all supervised learning operations, which may not always be available for all crops and/or diseases, as was the case here. In contrast, PCA condenses the readily available information for numerous variables into fewer dimensions, and displays the alignment of each sample with respect to the different features in the biplot. Thus, PCA was used here to create a qualitative gradient based on numerous parameters instead of directly determining the fate of a sample.
Salient characteristics of the data analysis method adopted herein are as follows: 1) combining the information from diverse sensors improved data segregation; 2) unsupervised data processing allowed fully objective analysis, with no human intervention and minimal preprocessing; 3) it did not require elaborate training datasets for model generation; 4) could be implemented and interpreted with limited computational expertise; 5) could be implemented with a limited sample size. Moreover, as the method described here is totally non-specific, i.e., it is model-free and does not rely on any previously-generated information, it could be easily adapted or customised for other crops and diseases as well. Nonetheless, this method does yield PCs, i.e., linear combinations of the original features, which could be used for further characterisation of new samples from the same crop imaged under the same conditions.
Since the method presented in the study is unsupervised, it focused on maximising separability between samples based on multiple canopy features, but did not tag samples as “healthy” or “infected”. Thus, in practice users could follow this approach to locate divergent samples by using a set of known reliable features to identify the cluster of stressed plants based on their relative location on the biplot. For example, as observed in our analyses, features such as NDVI or AGRI always pointed towards the healthier samples, whereas G was directed towards the unhealthier samples (Fig. 5b). However, knowledge of feature trends for healthy and unhealthy samples would be needed to correctly interpret the clustering.
Inability to inherently account for feature redundancy might bias the results of PCA, necessitating stringent feature selection prior to plant monitoring via the presented method. Another limitation of the PCA-based approach presented here is that it would not be able to perform well if all the samples being analysed were infected and were showing very similar canopy symptoms as the process does not rely on earlier trials or datasets. In addition, since this study aimed at identifying disease symptoms in an indoor farming system with controlled lighting, data pre-processing was minimal, i.e., only linear correlation analysis was performed for reducing redundancy. However, if the protocol is adapted for operations with a variable light source (e.g., sunlight) as in the field or a glasshouse, colour balancing would be needed as the first data pre-processing step to compensate for variations in light environment at each imaging interval. Hence, relevant considerations would be required before implementing the proposed method to achieve better results for different crops, diseases, cultivation systems, and sensors. Use of supervised ML and deep-learning tools for multi-sensor dataset analysis could also expand the scope of implementing such crop monitoring systems.
Although machine vision is very advantageous for crop monitoring, practical application of the technology in vertical farms poses certain challenges that are not encountered in field studies (Tian et al., 2022). For instance, installation of imaging sensors within the growing area is not feasible because such crop production systems aim at maximising vertical space utilisation and have crops growing in multiple tiers. Additionally, illumination spectrum could be another limiting factor for in situ imaging in vertical farms that employ non-white LED lighting, as leaf spectral reflectance characteristics would change according to incident light, necessitating reconfiguration of spectral indices that have been established using white light. A solution to these issues is the use of mobile cultivation beds that may be transferred intermittently to a fixed imaging platform for crop monitoring. Use of a fixed and optimised lighting regime during imaging would allow better standardisation of spectral data collection in such systems. Camera resolution and planting density would also affect the overall accuracy of the process, and image analysis protocols would require careful spatial calibration as per planting layout to identify trends at individual plant level. The present study highlights the efficacy of unsupervised and simultaneous implementation of spectral and morphometric features for early detection of root rot in hydroponics using flat-parsley as a model system. Investigations with other plant species and diseases are needed to further expand this knowledge base, and use of other imaging systems such as thermal, hyperspectral, and fluorescence cameras along with multispectral and 3D imaging will be especially helpful in providing further insights into the scope of applying machine vision for early disease detection in vertical farms.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Abbas, A., Jain, S., Gour, M., & Vankudothu, S. (2021). Tomato plant disease detection using transfer learning with C-GAN synthetic images. Computers and Electronics in Agriculture, 187, 106279. https://doi.org/10.1016/j.compag.2021.106279
Agarwal, A., Dongre, P. K., & Dutta Gupta, S. (2021). Smartphone-assisted real-time estimation of chlorophyll and carotenoid concentrations and ratio using the inverse of red and green digital color features. Theoretical and Experimental Plant Physiology, 33, 293–302. https://doi.org/10.1007/s40626-021-00210-4
Agarwal, A., & Dutta Gupta, S. (2018). Assessment of spinach seedling health status and chlorophyll content by multivariate data analysis and multiple linear regression of leaf image features. Computers and Electronics in Agriculture, 152, 281–289. https://doi.org/10.1016/j.compag.2018.06.048
Agarwal, M., Gupta, S. K., & Biswas, K. K. (2020). Development of efficient CNN model for tomato crop disease identification. Sustainable Computing: Informatics and Systems, 28, 100407. https://doi.org/10.1016/j.suscom.2020.100407
Ahmad, A., Saraswat, D., & El Gamal, A. (2023). A survey on using deep learning techniques for plant disease diagnosis and recommendations for development of appropriate tools. Smart Agricultural Technology, 3, 100083. https://doi.org/10.1016/j.atech.2022.100083
Anjna, Sood, M., & Singh, P. K. (2020). Hybrid system for detection and classification of plant disease using qualitative texture features analysis. Procedia Computer Science, 167, 1056–1065. https://doi.org/10.1016/j.procs.2020.03.404
Bedi, P., & Gole, P. (2021). Plant disease detection using hybrid model based on convolutional autoencoder and convolutional neural network. Artificial Intelligence in Agriculture, 5, 90–101. https://doi.org/10.1016/j.aiia.2021.05.002
Bock, C. H., Poole, G. H., Parker, P. E., & Gottwald, T. R. (2010). Plant disease severity estimated visually, by digital photography and image analysis, and by hyperspectral imaging. Critical Reviews in Plant Sciences, 29, 59–107. https://doi.org/10.1080/07352681003617285
Calderón, R., Navas-Cortés, J. A., & Zarco-Tejada, P. J. (2015). Early detection and quantification of Verticillium wilt in olive using hyperspectral and thermal imagery over large areas. Remote Sensing, 7, 5584–5610. https://doi.org/10.3390/rs70505584
Chowdhury, M. E., Rahman, T., Khandakar, A., Ayari, M. A., Khan, A. U., Khan, M. S., Al-Emadi, N., Reaz, M. B. I., Islam, M. T., & Ali, S. H. M. (2021). Automatic and reliable leaf disease detection using deep learning techniques. AgriEngineering, 3, 294–312. https://doi.org/10.3390/agriengineering3020020
DadrasJavan, F., Samadzadegan, F., Seyed Pourazar, S. H., & Fazeli, H. (2019). UAV-based multispectral imagery for fast Citrus Greening detection. Journal of Plant Diseases and Protection, 126, 307–318. https://doi.org/10.1007/s41348-019-00234-8
Franke, J., & Menz, G. (2007). Multi-temporal wheat disease detection by multi-spectral remote sensing. Precision Agriculture, 8, 161–172. https://doi.org/10.1007/s11119-007-9036-y
Friedli, M., Kirchgessner, N., Grieder, C., Liebisch, F., Mannale, M., & Walter, A. (2016). Terrestrial 3D laser scanning to track the increase in canopy height of both monocot and dicot crop species under field conditions. Plant Methods, 12, 9. https://doi.org/10.1186/s13007-016-0109-7
Ghosal, S., Blystone, D., Singh, A. K., Ganapathysubramanian, B., Singh, A., & Sarkar, S. (2018). An explainable deep machine vision framework for plant stress phenotyping. Proceedings of the National Academy of Sciences of the United States of America, 115, 4613–4618. https://doi.org/10.1073/pnas.1716999115
Harakannanavar, S. S., Rudagi, J. M., Puranikmath, V. I., Siddiqua, A., & Pramodhini, R. (2022). Plant leaf disease detection using computer vision and machine learning algorithms. Global Transitions Proceedings, 3, 305–310. https://doi.org/10.1016/j.gltp.2022.03.016
Husin, N. A., Khairunniza-Bejo, S., Abdullah, A. F., Kassim, M. S. M., Ahmad, D., & Azmi, A. N. N. (2020). Application of ground-based LiDAR for analysing oil palm canopy properties on the occurrence of basal stem rot (BSR) disease. Scientific Reports, 10, 6464. https://doi.org/10.1038/s41598-020-62275-6
Lazarević, B., Šatović, Z., Nimac, A., Vidak, M., Gunjača, J., Politeo, O., & Carović-Stanko, K. (2021). Application of phenotyping methods in detection of drought and salinity stress in basil (Ocimum basilicum L.). Frontiers in Plant Science, 12, 629441. https://doi.org/10.3389/fpls.2021.629441
Li, Y., Nie, J., & Chao, X. (2020). Do we really need deep CNN for plant diseases identification? Computers and Electronics in Agriculture, 178, 105803. https://doi.org/10.1016/j.compag.2020.105803
López-López, M., Calderón, R., González-Dugo, V., Zarco-Tejada, P. J., & Fereres, E. (2016). Early detection and quantification of almond red leaf blotch using high-resolution hyperspectral and thermal imagery. Remote Sensing, 8, 276. https://doi.org/10.3390/rs8040276
Manavalan, L. P., Cui, I., Ambrose, K. V., Panjwani, S., DeLong, S., Mleczko, M., & Spetsieris, K. (2021). Systematic approach to validate and implement digital phenotyping tool for soybean: A case study with PlantEye. Plant Phenome Journal, 4, e20025. https://doi.org/10.1002/ppj2.20025
Martin, F. N. (2000). Phylogenetic relationships among some Pythium species inferred from sequence analysis of the mitochondrially encoded cytochrome oxidase II gene. Mycologia, 92, 711–727. https://doi.org/10.1080/00275514.2000.12061211
Martinelli, F., Scalenghe, R., Davino, S., Panno, S., Scuderi, G., Ruisi, P., Villa, P., Stroppiana, D., Boschetti, M., Goulart, L. R., Davis, C. E., & Dandekar, A. M. (2015). Advanced methods of plant disease detection. A review. Agronomy for Sustainable Development, 35, 1–25. https://doi.org/10.1007/s13593-014-0246-1
Matthiesen, R. L., Ahmad, A. A., & Robertson, A. E. (2016). Temperature affects aggressiveness and fungicide sensitivity of four Pythium spp. That cause soybean and corn damping off in Iowa. Plant Disease, 100, 583–591. https://doi.org/10.1094/PDIS-04-15-0487-RE
McGehee, C. S., Raudales, R. E., Elmer, W. H., & McAvoy, R. J. (2019). Efficacy of biofungicides against root rot and damping-off of microgreens caused by Pythium spp. Crop Protection, 121, 96–102. https://doi.org/10.1016/j.cropro.2018.12.007
Minchinton, E., Petkowski, J., deBoer, D., Thomson, F., Trapnell, L., Tesoriero, L., Forsyth, L., Parker, J., Pung, H., & McKay, A. (2013). Identification of IPM strategies for Pythium induced root rots in Apiacae vegetable crops. Horticulture Australia Ltd., Sydney.
Mohanty, S. P., Hughes, D. P., & Salathé, M. (2016). Using deep learning for image-based plant disease detection. Frontiers in Plant Science, 7, 1419. https://doi.org/10.3389/fpls.2016.01419
Mulugeta Aneley, G., Haas, M., & Köhl, K. (2022). LIDAR-based phenotyping for drought response and drought tolerance in potato. Potato Research. https://doi.org/10.1007/s11540-022-09567-8
Mutka, A. M., & Bart, R. S. (2015). Image-based phenotyping of plant disease symptoms. Frontiers in Plant Science, 5, 734. https://doi.org/10.3389/fpls.2014.00734
Nagaraju, M., & Chawla, P. (2020). Systematic review of deep learning techniques in plant disease detection. International Journal of System Assurance Engineering and Management, 11, 547–560. https://doi.org/10.1007/s13198-020-00972-1
Nagasubramanian, K., Jones, S., Singh, A. K., Sarkar, S., Singh, A., & Ganapathysubramanian, B. (2019). Plant disease identification using explainable 3D deep learning on hyperspectral images. Plant Methods, 15, 98. https://doi.org/10.1186/s13007-019-0479-8
Nilsson, H.-E. (1995). Remote sensing and image analysis in plant pathology. Annual Review of Phytopathology, 15, 489–527. https://doi.org/10.1146/annurev.py.33.090195.002421
Okubara, P. A., & Paulitz, T. C. (2005). Root defense responses to fungal pathogens: A molecular perspective. Plant and Soil, 274, 215–226. https://doi.org/10.1007/1-4020-4099-7_11
Panigrahi, K. P., Das, H., Sahoo, A. K., & Moharana, S. C. (2020). Maize leaf disease detection and classification using machine learning algorithms. In Progress in Computing, Analytics and Networking: Proceedings of ICCAN 2019, pp. 659–669. https://doi.org/10.1007/978-981-15-2414-1_66
Paulitz, T. C. (1997). Biological control of root pathogens in soilless and hydroponic systems. HortScience, 32, 193–196. https://doi.org/10.21273/HORTSCI.32.2.193
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V., Thirion, B., Grisel, O., Blondel, M., Prettenhofer, P., Weiss, R., Dubourg, V., Vanderplas, J., Passos, A., Cournapeau, D., Brucher, M., Perrot, M., & Duchesnay, É. (2011). Scikit-learn: Machine learning in Python. Journal of Machine Learning Research, 12, 2825–2830.
Qin, Z., & Zhang, M. (2005). Detection of rice sheath blight for in-season disease management using multispectral remote sensing. International Journal of Applied Earth Observation and Geoinformation, 7, 115–128. https://doi.org/10.1016/j.jag.2005.03.004
Rajan, P., Lada, R. R., & MacDonald, M. T. (2019). Advancement in indoor vertical farming for microgreen production. American Journal of Plant Sciences, 10, 1397–1408. https://doi.org/10.4236/ajps.2019.108100
Raji, S. N., Subhash, N., Ravi, V., Saravanan, R., Mohanan, C. N., MakeshKumar, T., & Nita, S. (2016). Detection and classification of mosaic virus disease in cassava plants by proximal sensing of photochemical reflectance index. Journal of the Indian Society of Remote Sensing, 44, 875–883. https://doi.org/10.1007/s12524-016-0565-6
Roberts, J. M., Bruce, T. J. A., Monaghan, J. M., Pope, T. W., Leather, S. R., & Beacham, A. M. (2020). Vertical farming systems bring new considerations for pest and disease management. Annals of Applied Biology, 176, 226–232. https://doi.org/10.1111/aab.12587
Saleem, M. H., Khanchi, S., Potgieter, J., & Arif, K. M. (2020). Image-based plant disease identification by deep learning meta-architectures. Plants, 9, 1451. https://doi.org/10.3390/plants9111451
Salgadoe, A. S. A., Robson, A. J., Lamb, D. W., Dann, E. K., & Searle, C. (2018). Quantifying the severity of phytophthora root rot disease in avocado trees using image analysis. Remote Sensing, 10, 226. https://doi.org/10.3390/rs10020226
Sambasivam, G., & Opiyo, G. D. (2021). A predictive machine learning application in agriculture: Cassava disease detection and classification with imbalanced dataset using convolutional neural networks. Egyptian Informatics Journal, 22, 27–34. https://doi.org/10.1016/j.eij.2020.02.007
Shin, J., Chang, Y. K., Heung, B., Nguyen-Quang, T., Price, G. W., & Al-Mallahi, A. (2021). A deep learning approach for RGB image-based powdery mildew disease detection on strawberry leaves. Computers and Electronics in Agriculture, 183, 106042. https://doi.org/10.1016/j.compag.2021.106042
Shrivastava, V. K., & Pradhan, M. K. (2021). Rice plant disease classification using color features: A machine learning paradigm. Journal of Plant Pathology, 103, 17–26. https://doi.org/10.1007/s42161-020-00683-3
Singh, A., Jones, S., Ganapathysubramanian, B., Sarkar, S., Mueller, D., Sandhu, K., & Nagasubramanian, K. (2021). Challenges and opportunities in machine-augmented plant stress phenotyping. Trends in Plant Science, 26, 53–69. https://doi.org/10.1016/j.tplants.2020.07.010
Specht, K., Siebert, R., Hartmann, I., Freisinger, U. B., Sawicka, M., Werner, A., Thomaier, S., Henckel, D., Walk, H., & Dierich, A. (2014). Urban agriculture of the future: An overview of sustainability aspects of food production in and on buildings. Agriculture and Human Values, 31, 33–51. https://doi.org/10.1007/s10460-013-9448-4
Su, Y., Wu, F., Ao, Z., Jin, S., Qin, F., Liu, B., Pang, S., Liu, L., & Guo, Q. (2019). Evaluating maize phenotype dynamics under drought stress using terrestrial lidar. Plant Methods, 15, 11. https://doi.org/10.1186/s13007-019-0396-x
Suárez-Cáceres, G. P., Pérez-Urrestarazu, L., Avilés, M., Borrero, C., Lobillo Eguíbar, J. R., & Fernández-Cabanás, V. M. (2021). Susceptibility to water-borne plant diseases of hydroponic vs. aquaponics systems. Aquaculture, 544, 737093. https://doi.org/10.1016/j.aquaculture.2021.737093
Sugiura, R., Tsuda, S., Tamiya, S., Itoh, A., Nishiwaki, K., Murakami, N., Shibuya, Y., Hirafuji, M., & Nuske, S. (2016). Field phenotyping system for the assessment of potato late blight resistance using RGB imagery from an unmanned aerial vehicle. Biosystems Engineering, 148, 1–10. https://doi.org/10.1016/j.biosystemseng.2016.04.010
Sujatha, R., Chatterjee, J. M., Jhanjhi, N. Z., & Brohi, S. N. (2021). Performance of deep learning vs machine learning in plant leaf disease detection. Microprocessors and Microsystems, 80, 103615. https://doi.org/10.1016/j.micpro.2020.103615
Tian, Z., Ma, W., Yang, Q., & Duan, F. (2022). Application status and challenges of machine vision in plant factory—A review. Information Processing in Agriculture, 9, 195–211. https://doi.org/10.1016/j.inpa.2021.06.003
Tiwari, V., Joshi, R. C., & Dutta, M. K. (2021). Dense convolutional neural networks based multiclass plant disease detection and classification using leaf images. Ecological Informatics, 63, 101289. https://doi.org/10.1016/j.ecoinf.2021.101289
van Delden, S. H., SharathKumar, M., Butturini, M., Graamans, L. J. A., Heuvelink, E., Kacira, M., Kaiser, E., Klamer, R. S., Klerkx, L., Kootstra, G., Loeber, A., Schouten, R. E., Stanghellini, C., van Ieperen, W., Verdonk, J. C., Vialet-Chabrand, S., Woltering, E. J., van de Zedde, R., Zhang, Y., & Marcelis, L. F. M. (2021). Current status and future challenges in implementing and upscaling vertical farming systems. Nature Food, 2, 944–956. https://doi.org/10.1038/s43016-021-00402-w
Veys, C., Chatziavgerinos, F., AlSuwaidi, A., Hibbert, J., Hansen, M., Bernotas, G., Smith, M., Yin, H., Rolfe, S., & Grieve, B. (2019). Multispectral imaging for presymptomatic analysis of light leaf spot in oilseed rape. Plant Methods, 15, 4. https://doi.org/10.1186/s13007-019-0389-9
Waiphara, P., Bourgenot, C., Compton, L. J., & Prashar, A. (2022). Optical imaging resources for crop phenotyping and stress detection. In Duque, P., Szakonyi, D. (Eds.), Methods in molecular biology (Vol. 2494 pp. 255–265). Humana, New York. https://doi.org/10.1007/978-1-0716-2297-1_18
White, T. J., Bruns, T., Lee, S., & Taylor, J. W. (1990). Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In M. A. Innis, D. H. Gelfand, J. J. Sninsky, & T. J. White (Eds.), PCR Protocols: A Guide to Methods and Applications (pp. 315–322). Academic Press, Inc., New York
Xiao, S., Chai, H., Shao, K., Shen, M., Wang, Q., Wang, R., Sui, Y., & Ma, Y. (2020). Image-based dynamic quantification of aboveground structure of sugar beet in field. Remote Sensing, 12, 269. https://doi.org/10.3390/rs12020269
Yamamoto, K., Togami, T., & Yamaguchi, N. (2017). Super-resolution of plant disease images for the acceleration of image-based phenotyping and vigor diagnosis in agriculture. Sensors, 17, 2557. https://doi.org/10.3390/s17112557
Yang, C., Everitt, J. H., & Fernandez, C. J. (2010). Comparison of airborne multispectral and hyperspectral imagery for mapping cotton root rot. Biosystems Engineering, 107, 131–139. https://doi.org/10.1016/j.biosystemseng.2010.07.011
Yuan, H., Bennett, R. S., Wang, N., & Chamberlin, K. D. (2019). Development of a peanut canopy measurement system using a ground-based lidar sensor. Frontiers in Plant Science, 10, 203. https://doi.org/10.3389/fpls.2019.00203
Zamani, A. S., Anand, L., Rane, K. P., Prabhu, P., Buttar, A. M., Pallathadka, H., Raghuvanshi, A., & Dugbakie, B. N. (2022). Performance of machine learning and image processing in plant leaf disease detection. Journal of Food Quality, 2022, 1598796. https://doi.org/10.1155/2022/1598796
Zhang, J., Huang, Y., Pu, R., Gonzalez-Moreno, P., Yuan, L., Wu, K., & Huang, W. (2019). Monitoring plant diseases and pests through remote sensing technology: A review. Computers and Electronics in Agriculture, 165, 104943. https://doi.org/10.1016/j.compag.2019.104943
Zhang, J., Pu, R., Wang, J., Huang, W., Yuan, L., & Luo, J. (2012). Detecting powdery mildew of winter wheat using leaf level hyperspectral measurements. Computers and Electronics in Agriculture, 85, 13–23. https://doi.org/10.1016/j.compag.2012.03.006
Acknowledgements
We thank the InFarm UK team for supplying seedlings and providing technical support along with InFarm Crop Science team for their support. We acknowledge all the partners (RoboScientific, Marks and Spencer, and InFarm) for their feedback and support in the project. We also thank the staff at Newcastle University for their technical, administrative, and logistical support.
Funding
This work was funded by Innovate UK (Technology Strategy Board – CR&D) [grant number: TS/V002880/1].
Author information
Authors and Affiliations
Contributions
NB, VACG, TH, and AP conceived the project. NB, AP, VACG, FdJC, and JBR designed the study. AA, FdJC, and JBR performed the experiments. JBR and SS performed pathogen strain identification. AA performed image and data analysis. AA drafted the manuscript with key inputs from SS on pathology work. All authors contributed to amendments and revisions. All authors approved the final submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare that there are no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Agarwal, A., de Jesus Colwell, F., Bello Rodriguez, J. et al. Monitoring root rot in flat-leaf parsley via machine vision by unsupervised multivariate analysis of morphometric and spectral parameters. Eur J Plant Pathol (2024). https://doi.org/10.1007/s10658-024-02834-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s10658-024-02834-z