Abstract
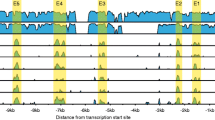
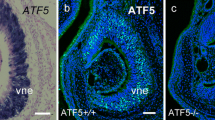
Activating transcription factor 5 (ATF5) is a transcription factor that belongs to the cAMP-response element-binding protein/ATF family and is essential for the differentiation and survival of sensory neurons in mouse olfactory organs. However, transcriptional target genes for ATF5 have yet to be identified. In the present study, chromatin immunoprecipitation-quantitative polymerase chain reaction (ChIP-qPCR) experiments were performed to verify ATF5 target genes in the main olfactory epithelium and vomeronasal organ in the postnatal pups. ChIP-qPCR was conducted using hemagglutinin (HA)-tagged ATF5 knock-in olfactory organs. The results obtained demonstrated that ATF5-HA fusion proteins bound to the CCAAT/enhancer-binding protein-ATF response element (CARE) site in the enhancer region of nescient helix-loop-helix 1 (Nhlh1), a transcription factor expressed in differentiating olfactory and vomeronasal sensory neurons. Nhlh1 mRNA expression was downregulated in ATF5-deficient (ATF5−/−) olfactory organs. The LIM/homeobox protein transcription factor Lhx2 co-localized with ATF5 in the nuclei of olfactory and vomeronasal sensory neurons and bound to the homeodomain site proximal to the CARE site in the Nhlh1 gene. The CARE region of the Nhlh1 gene was enriched by the active enhancer marker, acetyl-histone H3 (Lys27). The present study identified Nhlh1 as a novel target gene for ATF5 in murine olfactory organs. ATF5 may upregulate Nhlh1 expression in concert with Lhx2, thereby promoting the differentiation of olfactory and vomeronasal sensory neurons.




Similar content being viewed by others
Data availability
The datasets generated and/or analyzed in the present study are available from the corresponding author upon reasonable request.
Change history
26 March 2024
A Correction to this paper has been published: https://doi.org/10.1007/s00441-024-03891-w
References
Begley CG, Lipkowitz S, Göbel V, Mahon KA, Bertness V, Green AR, Gough NM, Kirsch IR (1992) Molecular characterization of NSCL, a gene encoding a helix-loop-helix protein expressed in the developing nervous system. Proc Natl Acad Sci USA 89:38–42
Castro-Mondragon JA, Riudavets-Puig R, Rauluseviciute I, Lemma RB, Turchi L, Blanc-Mathieu R, Lucas J, Boddie P, Khan A, Manosalva Pérez N, Fornes O, Leung TY, Aguirre A, Hammal F, Schmelter D, Baranasic D, Ballester B, Sandelin A, Lenhard B, Vandepoele K, Wasserman WW, Parcy F, Mathelier A (2022) JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res 50(D1):D165–D173
Cau E, Gradwohl G, Fode C, Guillemot F (1997) Mash1 activates a cascade of bHLH regulators in olfactory neuron progenitors. Development 124:1611–1621
Chang I, Parrilla M (2016) Expression patterns of homeobox genes in the mouse vomeronasal organ at postnatal stages. Gene Expr Patterns 21:69–80
Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, Boyer LA, Young RA, Jaenisch R (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci USA 107:21931–21936
Dalton RP, Lyons DB, Lomvardas S (2013) Co-opting the unfolded protein response to elicit olfactory receptor feedback. Cell 155:321–332
Dalton RP (2018) Shared genetic requirements for ATF5 translation in the vomeronasal organ and main olfactory epithelium [version 1; peer review: 3 approved with reservations]. F1000Research 7:73. https://doi.org/10.12688/f1000research.13659.1
Hansen MB, Mitchelmore C, Kjaerulff KM, Rasmussen TE, Pedersen KM, Jensen NA (2002) Mouse Atf5: molecular cloning of two novel mRNAs, genomic organization, and odorant sensory neuron localization. Genomics 80:344–350
Hatano M, Umemura M, Kimura N, Yamazaki T, Takeda H, Nakano H, Takahashi S, Takahashi Y (2013) The 5′-untranslated region regulates ATF5 mRNA stability via nonsense-mediated mRNA decay in response to environmental stress. FEBS J 280:4693–4707
Hirota J, Mombaerts P (2004) The LIM-homeodomain protein Lhx2 is required for complete development of mouse olfactory sensory neurons. Proc Natl Acad Sci USA 101:8751–8755
Hirota J, Omura M, Mombaerts P (2007) Differential impact of Lhx2 deficiency on expression of class I and class II odorant receptor genes in mouse. Mol Cell Neurosci 34:679–688
Ibarra-Soria X, Levitin MO, Saraiva LR, Logan DW (2014) The olfactory transcriptomes of mice. PLoS Genet 10:e1004593
Katreddi RR, Forni PE (2021) Mechanisms underlying pre- and postnatal development of the vomeronasal organ. Cell Mol Life Sci 78:5069–5082
Kim WY (2012) NeuroD1 is an upstream regulator of NSCL1. Biochem Biophys Res Commun 419:27–31
Kolterud A, Alenius M, Carlsson L, Bohm S (2004) The Lim homeobox gene Lhx2 is required for olfactory sensory neuron identity. Development 131:5319–5326
Krüger M, Braun T (2002) The neuronal basic helix-loop-helix transcription factor NSCL-1 is dispensable for normal neuronal development. Mol Cell Biol 22:792–800
Magklara A, Yen A, Colquitt BM, Clowney EJ, Allen W, Markenscoff-Papadimitriou E, Evans ZA, Kheradpour P, Mountoufaris G, Carey C, Barnea G, Kellis M, Lomvardas S (2011) An epigenetic signature for monoallelic olfactory receptor expression. Cell 145:555–570
Masuda A, Ajima R, Saga Y, Hirata T, Zhu Y (2022) Nhlh1 and Nhlh2, a global transcriptional mechanism regulating commissural axon projection via Robo3 activation. Preprint at https://doi.org/10.1101/2022.09.23.509112
Monahan K, Lomvardas S (2015) Monoallelic expression of olfactory receptors. Annu Rev Cell Dev Biol 31:721–740
Monahan K, Schieren I, Cheung J, Mumbey-Wafula A, Monuki ES, Lomvardas S (2017) Cooperative interactions enable singular olfactory receptor expression in mouse olfactory neurons. Elife 6:e28620
Monahan K, Horta A, Lomvardas S (2019) LHX2- and LDB1-mediated trans interactions regulate olfactory receptor choice. Nature 565:448–453
Murdoch JN, Eddleston J, Leblond-Bourget N, Stanier P, Copp AJ (1999) Sequence and expression analysis of Nhlh1: a basic helix-loop-helix gene implicated in neurogenesis. Dev Genet 24:165–177
Nakano H, Iida Y, Suzuki M, Aoki M, Umemura M, Takahashi S, Takahashi Y (2016) Activating transcription factor 5 (ATF5) is essential for the maturation and survival of mouse basal vomeronasal sensory neurons. Cell Tissue Res 363:621–633
Nakano H, Iida Y, Murase T, Oyama N, Umemura M, Takahashi S, Takahashi Y (2019) Co-expression of C/EBPγ and ATF5 in mouse vomeronasal sensory neurons during early postnatal development. Cell Tissue Res 378:427–440
Nakano H, Kawai S, Ooki Y, Chiba T, Ishii C, Nozawa T, Utsuki H, Umemura M, Takahashi S, Takahashi Y (2021) Functional validation of epitope-tagged ATF5 knock-in mice generated by improved genome editing of oviductal nucleic acid delivery (i-GONAD). Cell Tissue Res 385:239–249
Saito H, Kubota M, Roberts RW, Chi Q, Matsunami H (2004) RTP family members induce functional expression of mammalian odorant receptors. Cell 119:679–691
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682
Schmid T, Boehm U, Braun T (2020) GnRH neurogenesis depends on embryonic pheromone receptor expression. Mol Cell Endocrinol 518:111030
Shayya HJ, Kahiapo JK, Duffié R, Lehmann KS, Bashkirova L, Monahan K, Dalton RP, Gao J, Jiao S, Schieren I, Belluscio L, Lomvardas S (2022) ER stress transforms random olfactory receptor choice into axon targeting precision. Cell 185:3896-3912.e22
Suzuki Y, Tsuruga E, Yajima T, Takeda M (2003) Expression of bHLH transcription factors NSCL1 and NSCL2 in the mouse olfactory system. Chem Senses 28:603–608
Umemura M, Tsunematsu K, Shimizu YI, Nakano H, Takahashi S, Higashiura Y, Okabe M, Takahashi Y (2015) Activating transcription factor 5 is required for mouse olfactory bulb development via interneuron. Biosci Biotechnol Biochem 79:1082–1089
Wang SZ, Ou J, Zhu LJ, Green MR (2012) Transcription factor ATF5 is required for terminal differentiation and survival of olfactory sensory neurons. Proc Natl Acad Sci USA 109:18589–18594
Watatani Y, Ichikawa K, Nakanishi N, Fujimoto M, Takeda H, Kimura N, Hirose H, Takahashi S, Takahashi Y (2008) Stress-induced translation of ATF5 mRNA is regulated by the 5′-untranslated region. J Biol Chem 283:2543–2553
Wodrich APK, Scott AW, Shukla AK, Harris BT, Giniger E (2022) The unfolded protein responses in health, aging, and neurodegeneration: recent advances and future considerations. Front Mol Neurosci 15:831116
Zhang G, Titlow WB, Biecker SM, Stromberg AJ, McClintock TS (2016) Lhx2 determines odorant receptor expression frequency in mature olfactory sensory neurons. eNeuro 3:ENEURO.0230–16.2016
Zhou D, Palam LR, Jiang L, Narasimhan J, Staschke KA, Wek RC (2008) Phosphorylation of eIF2 directs ATF5 translational control in response to diverse stress conditions. J Biol Chem 283:7064–7073
Acknowledgements
We are grateful to the members of Yuji Takahashi’s lab for their support on this project.
Author information
Authors and Affiliations
Contributions
HN conceived the study and designed the project with CI, ST, and YT. MU maintained the ATF5-knockout mouse line. CI, HN, RH, and YO performed the experiments and analyzed the data. CI and HN wrote the manuscript, and all authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
All mouse studies were approved by the Institutional Animal Experiment Committee of the university and were performed in accordance with institutional and governmental guidelines.
Informed consent
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The authors regret that the version of the supplementary file that appears in the original article is incorrect. The correct supplementary figure is provided in the erratum article. The original article has been corrected.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ishii, C., Nakano, H., Higashiseto, R. et al. Nescient helix-loop-helix 1 (Nhlh1) is a novel activating transcription factor 5 (ATF5) target gene in olfactory and vomeronasal sensory neurons in mice. Cell Tissue Res 396, 85–94 (2024). https://doi.org/10.1007/s00441-024-03871-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-024-03871-0