Abstract
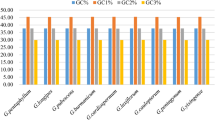
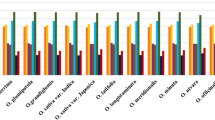
Gracilariaceae is a group of marine large red algae and main source of agar with important economic and ecological value. The codon usage patterns of chloroplast genomes in 36 species from Graciliaceae show that GC range from 0.284 to 0.335, the average GC3 range from 0.135 to 0.243 and the value of ENC range from 35.098 to 42.327, which indicates these genomes are rich in AT and prefer to use codons ending with AT in these species. Nc plot, PR2 plot, neutrality plot analyses and correlation analysis indicate that these biases may be caused by multiple factors, such as natural selection and mutation pressure, but prolonged natural selection is the main driving force influencing codon usage preference. The cluster analysis and phylogenetic analysis show that the differentiation relationship of them is different and indicate that codons with weak or unbiased preferences may also play an irreplaceable role in these species’ evolution. In addition, we identified 26 common high-frequency codons and 8–18 optimal codons all ending in A/U in these 36 species. Our results will not only contribute to carrying out transgenic work in Gracilariaceae species to maximize the protein yield in the future, but also lay a theoretical foundation for further exploring systematic classification of them.












Similar content being viewed by others
Data availability
The taxonomic information and chloroplast genomes data that support the findings of this study are openly available in NCBI (https://www.ncbi.nlm.nih.gov/). Other data generated or analysed during this research are available within the article and its supplementary materials.
References
Abreu MH, Pereira R, Yarish C, Buschmann AH, Sousa-Pinto I (2011) IMTA with gracilaria vermiculophylla: productivity and nutrient removal performance of the seaweed in a land-based pilot scale system. Aquaculture 312(1–4):77–87
Adl SM, Simpson AG, Farmer MA, Andersen RA, Anderson OR, Barta JR, Bowser SS, Brugerolle G, Fensome RA, Fredericq S et al (2005) The new higher level classification of eukaryotes with emphasis on the taxonomy of protists. J Eukaryot Microbiol 52(5):399–451
Andargie M, Zhu CY (2022) Genome-wide analysis of codon usage in sesame (Sesamum indicum L.). Heliyon 8(1):e08687
Archetti M (2004) Codon usage bias and mutation constraints reduce the level of error minimization of the genetic code. J Mol Evol 59(2):258–266
Bixler HJ, Porse H (2011) A decade of change in the seaweed hydrocolloids industry. J Appl Phycol 23(3):321–335
Blouin NA, Brodie JA, Grossman AC, Xu P, Brawley SH (2011) Porphyra: a marine crop shaped by stress. Trends Plant Sci 16(1):29–37
Brown CM, Stockwell PA, Trotman CN, Tate WP (1990) Sequence analysis suggests that tetra-nucleotides signal the termination of protein synthesis in eukaryotes. Nucleic Acids Res 18(21):6339–6345
Chakraborty S, Yengkhom S, Uddin A (2020) Analysis of codon usage bias of chloroplast genes in oryza species : codon usage of chloroplast genes in oryza species. Planta 252(4):67
Chaney JL, Clark PL (2015) Roles for synonymous codon usage in protein biogenesis. Annu Rev Biophys 44:143–166
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol Plant 13(8):1194–1202
Clarke RT, Greenacre MJ (1985) Theory and applications of correspondence analysis. J Anim Ecol 54:1031
Coghlan A, Wolfe KH (2000) Relationship of codon bias to mRNA concentration and protein length in Saccharomyces cerevisiae. Yeast 16(12):1131–1145
Cole KM, Sheath RG (1991) Biology of the red algae
Collen J, Porcel B, Carre W, Ball SG, Chaparro C, Tonon T, Barbeyron T, Michel G, Noel B, Valentin K et al (2013) Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the archaeplastida. Proc Natl Acad Sci U S A 110(13):5247–5252
Daniell H, Lin CS, Yu M, Chang WJ (2016) Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol 17(1):134
Daniell H, Jin S, Zhu XG, Gitzendanner MA, Soltis DE, Soltis PS (2021) Green giant-a tiny chloroplast genome with mighty power to produce high-value proteins: history and phylogeny. Plant Biotechnol J 19(3):430–447
Darriba D, Posada D, Kozlov AM, Stamatakis A, Morel B, Flouri T (2020) ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol Biol Evol 37(1):291–294
Davoult D, Surget G, Stiger-Pouvreau V, Noisette F, Riera P, Stagnol D, Androuin T, Poupart N (2017) Multiple effects of a gracilaria vermiculophylla invasion on estuarine mudflat functioning and diversity. Mar Environ Res 131:227–235
Di T, Chen GJ, Sun Y, Ou SY, Zeng XX, Ye H (2017) Antioxidant and immunostimulating activities in vitro of sulfated polysaccharides isolated from Gracilaria rubra. J Funct Foods 28:64–75
dos Reis M, Wernisch L, Savva R (2003) Unexpected correlations between gene expression and codon usage bias from microarray data for the whole Escherichia coli K-12 genome. Nucleic Acids Res 31(23):6976–6985
Duret L, Mouchiroud D (1999) Expression pattern and surprisingly, gene length shape codon usage in caenorhabditis, drosophila, and arabidopsis. Proc Natl Acad Sci U S A 96(8):4482–4487
Fages-Lartaud M, Hundvin K, Hohmann-Marriott MF (2022) Mechanisms governing codon usage bias and the implications for protein expression in the chloroplast of Chlamydomonas reinhardtii. Plant J 112(4):919–945
Fan YL, Wang WH, Song W, Chen HS, Teng AG, Liu AJ (2012) Partial characterization and anti-tumor activity of an acidic polysaccharide from Gracilaria lemaneiformis. Carbohyd Polym 88(4):1313–1318
Frumkin I, Lajoie MJ, Gregg CJ, Hornung G, Church GM, Pilpel Y (2018) Codon usage of highly expressed genes affects proteome-wide translation efficiency. Proc Natl Acad Sci U S A 115(21):E4940–E4949
Ghaemmaghami S, Huh WK, Bower K, Howson RW, Belle A, Dephoure N, O’Shea EK, Weissman JS (2003) Global analysis of protein expression in yeast. Nature 425(6959):737–741
Goetz RM, Fuglsang A (2005) Correlation of codon bias measures with mRNA levels: analysis of transcriptome data from Escherichia coli. Biochem Biophys Res Commun 327(1):4–7
Gouy M, Gautier C (1982) Codon usage in bacteria: correlation with gene expressivity. Nucleic Acids Res 10(22):7055–7074
Grantham R, Gautier C, Gouy M, Mercier R, Pave A (1980) Codon catalog usage and the genome hypothesis. Nucleic Acids Res 8(1):r49–r62
Gurgel CFD, Fredericq S (2004) Systematics of the gracilariaceae (gracilariales, rhodophyta): a critical assessment based on rbcL sequence analyses. J Phycol 40(1):138–159
Gustafsson C, Minshull J, Govindarajan S, Ness J, Villalobos A, Welch M (2012) Engineering genes for predictable protein expression. Protein Expr Purif 83(1):37–46
He B, Dong H, Jiang C, Cao F, Tao S, Xu LA (2016) Analysis of codon usage patterns in Ginkgo biloba reveals codon usage tendency from a/u-ending to G/C-ending. Sci Rep 6:35927
Herrero R, Lodeiro P, Rojo R, Ciorba A, Rodríguez P (2008) Sastre de Vicente ME: the efficiency of the red alga mastocarpus stellatus for remediation of cadmium pollution. Biores Technol 99(10):4138–4146
Hershberg R, Petrov DA (2008) Selection on codon bias. Annu Rev Genet 42:287–299
Ibrahim WM (2011) Biosorption of heavy metal ions from aqueous solution by red macroalgae. J Hazard Mater 192(3):1827–1835
Ikemura T (1985) Codon usage and tRNA content in unicellular and multicellular organisms. Mol Biol Evol 2(1):13–34
Ingvarsson PK (2008) Molecular evolution of synonymous codon usage in populus. BMC Evol Biol 8:307
Iriarte A, Lamolle G, Musto H (2021) Codon usage bias: an endless tale. J Mol Evol 89(9–10):589–593
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780
Keeling PJ (2004) Diversity and evolutionary history of plastids and their hosts. Am J Bot 91(10):1481–1493
Keeling PJ (2010) The endosymbiotic origin, diversification and fate of plastids. Philos Trans R Soc Lond B Biol Sci 365(1541):729–748
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35(21):4453–4455
Ku C, Nelson-Sathi S, Roettger M, Garg S, Hazkani-Covo E, Martin WF (2015) Endosymbiotic gene transfer from prokaryotic pangenomes: inherited chimerism in eukaryotes. Proc Natl Acad Sci U S A 112(33):10139–10146
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549
Li G, Pan Z, Gao S, He Y, Xia Q, Jin Y, Yao H (2019) Analysis of synonymous codon usage of chloroplast genome in Porphyra umbilicalis. Genes Genomics 41(10):1173–1181
Li GLPZL, He YY, Gao SC, Xia QY, Jin Y, Yao HP (2020) Analysis of codon bias of chloroplast genome in Porphyra yezoensis. Jiyinzuxue Yu Yingyong Shengwuxue (genomics and Applied Biology) 39(12):5789–5795 ((in Chinese))
Li G, Zhang L, Xue P, Zhu M (2022) Comparative Analysis on the Codon Usage Pattern of the Chloroplast Genomes in Malus Species. Biochem Genet 61:1050
Liu XY, Li Y, Ji KK, Zhu J, Ling P, Zhou T, Fan LY, Xie SQ (2020) Genome-wide codon usage pattern analysis reveals the correlation between codon usage bias and gene expression in Cuscuta australis. Genomics 112(4):2695–2702
Liu Y (2020) A code within the genetic code: codon usage regulates co-translational protein folding. Cell Commun Signal 18(1):145
Lyra GM, Gurgel CF, Costa ED, de Jesus PB, Oliveira MC, Oliveira EC, Davis CC, Nunes JM (2016) Delimitating cryptic species in the Gracilaria domingensis complex (gracilariaceae, rhodophyta) using molecular and morphological data. J Phycol 52(6):997–1017
Lyra GD, Iha C, Grassa CJ, Cai LM, Zhang HR, Lane C, Blouin N, Oliveira MC, Nunes JMD, Davis CC (2021) Phylogenomics, divergence time estimation and trait evolution provide a new look into the Gracilariales (Rhodophyta). Molecular Phylogenetics and Evolution 165:107295
Lyra Gde M, Costa Eda S, de Jesus PB, de Matos JC, Caires TA, Oliveira MC, Oliveira EC, Xi Z, Nunes JM, Davis CC (2015) Phylogeny of gracilariaceae (rhodophyta): evidence from plastid and mitochondrial nucleotide sequences. J Phycol 51(2):356–366
Ma L, Cui P, Zhu J, Zhang Z, Zhang Z (2014) Translational selection in human: more pronounced in housekeeping genes. Biol Direct 9:17
Mawi S, Krishnan S, Din MFMD, Arumugam N, Chelliapan S (2020) Bioremediation potential of macroalgae Gracilaria edulis and gracilaria changii co-cultured with shrimp wastewater in an outdoor water recirculation system. Environ Technol Innov 17:100571
McFadden GI (2001) Chloroplast origin and integration. Plant Physiol 125(1):50–53
Mohanta TK, Mishra AK, Khan A, Hashem A, Abd Allah EF, Al-Harrasi A (2020) Gene Loss and Evolution of the Plastome. Genes (Basel) 11(10):1133
Morton BR (1998) Selection on the codon bias of chloroplast and cyanelle genes in different plant and algal lineages. J Mol Evol 46(4):449–459
Morton BR (2003) The role of context-dependent mutations in generating compositional and codon usage bias in grass chloroplast DNA. J Mol Evol 56(5):616–629
Nakamura-Gouvea N, Alves-Lima C, Benites LF, Iha C, Maracaja-Coutinho V, Aliaga-Tobar V, Carneiro M, Yokoya NS, Marinho-Soriano E, Graminha MAS et al (2022) Insights into agar and secondary metabolite pathways from the genome of the red alga Gracilaria domingensis (rhodophyta, gracilariales). J Phycol 58(3):406–423
Nie XJ, Deng PC, Feng KW, Liu PX, Du XH, You FM, Song WN (2014) Comparative analysis of codon usage patterns in chloroplast genomes of the Asteraceae family. Plant Mol Biol Rep 32(4):828–840
Osteryoung KW, Weber AP (2011) Plastid biology: focus on the defining organelle of plants. Plant Physiol 155(4):1475–1476
Palmer JD, Soltis DE, Chase MW (2004) The plant tree of life: an overview and some points of view. Am J Bot 91(10):1437–1445
Parvathy ST, Udayasuriyan V, Bhadana V (2022) Codon usage bias. Mol Biol Rep 49(1):539–565
Peden JF(1999) Analysis of codon usage [PhD dissertation]
Pinto RM, Bosch A (2021) The codon usage code for cotranslational folding of viral capsids. Genome Biol Evol 13(9):evab089
Poblete R, Cortes E, Macchiavello J, Bakit J (2018) Factors influencing solar drying performance of the red algae Gracilaria chilensis. Renew Energ 126:978–986
Pogson BJ, Ganguly D, Albrecht-Borth V (2015) Insights into chloroplast biogenesis and development. Bba-Bioenergetics 1847(9):1017–1024
Price DC, Chan CX, Yoon HS, Yang EC, Qiu H, Weber AP, Schwacke R, Gross J, Blouin NA, Lane C et al (2012) Cyanophora paradoxa genome elucidates origin of photosynthesis in algae and plants. Science 335(6070):843–847
Rodriguez-Ezpeleta N, Brinkmann H, Burey SC, Roure B, Burger G, Loffelhardt W, Bohnert HJ, Philippe H, Lang BF (2005) Monophyly of primary photosynthetic eukaryotes: green plants, red algae, and glaucophytes. Curr Biol 15(14):1325–1330
Ścieszka S, Klewicka E (2019) Algae in food: a general review. Crit Rev Food Sci Nutr 59(21):3538–3547
Sharp PM, Li WH (1986) An evolutionary perspective on synonymous codon usage in unicellular organisms. J Mol Evol 24(1–2):28–38
Sharp PM, Li WH (1987) The codon adaptation index–a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15(3):1281–1295
Sharp PM, Cowe E, Higgins DG, Shields DC, Wolfe KH, Wright F (1988) Codon usage patterns in Escherichia coli, Bacillus subtilis, Saccharomyces cerevisiae, schizosaccharomyces pombe, Drosophila melanogaster and Homo sapiens; a review of the considerable within-species diversity. Nucleic Acids Res 16(17):8207–8211
Sharp PM, Bailes E, Grocock RJ, Peden JF, Sockett RE (2005) Variation in the strength of selected codon usage bias among bacteria. Nucleic Acids Res 33(4):1141–1153
Sharp PM, Emery LR, Zeng K (2010) Forces that influence the evolution of codon bias. Philos Trans R Soc Lond B Biol Sci 365(1544):1203–1212
Smith DR, Keeling PJ (2015) Mitochondrial and plastid genome architecture: reoccurring themes, but significant differences at the extremes. P Natl Acad Sci USA 112(33):10177–10184
Stegemann S, Hartmann S, Ruf S, Bock R (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci U S A 100(15):8828–8833
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci U S A 85(8):2653–2657
Sueoka N (1995) Intrastrand parity rules of DNA base composition and usage biases of synonymous codons. J Mol Evol 40(3):318–325
Sueoka N (1999a) Translation-coupled violation of parity rule 2 in human genes is not the cause of heterogeneity of the DNA G+C content of third codon position. Gene 238(1):53–58
Sueoka N (1999b) Two aspects of DNA base composition: G+C content and translation-coupled deviation from intra-strand rule of a = T and G = C. J Mol Evol 49(1):49–62
Sueoka N (2001) Near homogeneity of PR2-bias fingerprints in the human genome and their implications in phylogenetic analyses. J Mol Evol 53(4–5):469–476
Sun J, Chen M, Xu J, Luo J (2005) Relationships among stop codon usage bias, its context, isochores, and gene expression level in various eukaryotes. J Mol Evol 61(4):437–444
Sun X, Yang Q, Xia X (2013) An improved implementation of effective number of codons (nc). Mol Biol Evol 30(1):191–196
Suzuki H, Morton BR (2016) Codon adaptation of plastid genes. PLoS ONE 11(5):e0154306
Suzuki H, Brown CJ, Forney LJ, Top EM (2008) Comparison of correspondence analysis methods for synonymous codon usage in bacteria. DNA Res 15(6):357–365
Takaichi S, Yokoyama A, Mochimaru M, Uchida H, Murakami A (2016) Carotenogenesis diversification in phylogenetic lineages of rhodophyta. J Phycol 52(3):329–338
Tonti-Filippini J, Nevill PG, Dixon K, Small I (2017) What can we do with 1000 plastid genomes? Plant J 90(4):808–818
Trench RK (1981) Chloroplasts: presumptive and de facto organelles. Ann N Y Acad Sci 361:341–355
Wang Z, Xu B, Li B, Zhou Q, Wang G, Jiang X, Wang C, Xu Z (2020) Comparative analysis of codon usage patterns in chloroplast genomes of six euphorbiaceae species. PeerJ 8:e8251
Wang Z, Cai Q, Wang Y, Li M, Wang C, Wang Z, Jiao C, Xu C, Wang H, Zhang Z (2022) Comparative analysis of codon bias in the chloroplast genomes of theaceae species. Front Genet 13:824610
Wang XL, Guo ML, Yan SS, Wang YQ, Sun ZM, Xia BM, Wang GC (2023) Diversity of Gracilariaceae (Rhodophyta) in China: An integrative morphological and molecular assessment including a description of Gracilaria tsengii sp. nov. Algal Res 71:103074
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci U S A 84(24):9054–9058
Wright F (1990) The “effective number of codons” used in a gene. Gene 87(1):23–29
Yao H, Li T, Ma Z, Wang X, Xu L, Zhang Y, Cai Y, Tang Z (2023) Codon usage pattern of the ancestor of green plants revealed through rhodophyta. BMC Genomics 24(1):538
Yoon HS, Müller K, Sheath R, Ott F, Bhattacharya D (2006) Defining the major lineages of red algae (rhodophyta). J Phycol 42(2):482–492
Yu TH, Li JS, Yang Y, Qi L, Chen BB, Zhao FQ, Bao QY, Wu JY (2012) Codon usage patterns and adaptive evolution of marine unicellular cyanobacteria synechococcus and prochlorococcus. Mol Phylogenet Evol 62(1):206–213
Zhang Z, Li J, Cui P, Ding F, Li A, Townsend JP, Yu J (2012) Codon deviation coefficient: a novel measure for estimating codon usage bias and its statistical significance. BMC Bioinformatics 13:43
Zhaxybayeva O, Doolittle WF, Papke RT, Gogarten JP (2009) Intertwined evolutionary histories of marine synechococcus and Prochlorococcus marinus. Genome Biol Evol 1:325–339
Acknowledgements
We are grateful to all those who have supported this work.
Funding
This work was jointly funded by the 2023 Provincial Science and Technology Plan (2023YFH0103) and the 2022 Double Support Project of Sichuan Agricultural University.
Author information
Authors and Affiliations
Contributions
TTL and HPY identified the research topic and conceived the experiments. FYL and XC collected and download the data. TMD, YXY, FW and XJW participated in the data processing of experiments. TTL and ZM performed and finished the data analysis for experiments. TTL wrote the manuscript. HPY was responsible for the entire article revision, and finalization. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
There is no any research involving humans or animals in this article.
Consent for publication
Not applicable.
Competing interests
The authors declare that there is no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10142_2024_1316_MOESM1_ESM.pdf
Supplementary file1 The high-frequency codons (codons with RSCU>1) of 36 Gracilariaceae species. The darker the color of the grid, the greater the value of RSCU. (PDF 51 KB)
10142_2024_1316_MOESM2_ESM.pdf
Supplementary file2 The highly expressed codons (codons with △RSCU>0.08) of 36 Gracilariaceae species. The darker the color of the grid, the greater the value of △RSCU. (PDF 240 KB)
10142_2024_1316_MOESM3_ESM.pdf
Supplementary file3 The accession numbers, sequence length, number of CDS before processing, number of CDS after processing and detailed information of the classification of each species for 36 Gracilariaceae species. (PDF 11 KB)
10142_2024_1316_MOESM4_ESM.xlsx
Supplementary file4 Codon usage statistics in the coding sequence of 36 Gracilariaceae species. SD represents the standard deviation of the data. N/A indicates that there is no optimal preferred stop codon in the species. The slope of regression line represents the lope of regression line in neutrality plot analysis. (XLSX 15 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, T., Ma, Z., Ding, T. et al. Codon usage bias and phylogenetic analysis of chloroplast genome in 36 gracilariaceae species. Funct Integr Genomics 24, 45 (2024). https://doi.org/10.1007/s10142-024-01316-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-024-01316-z