Abstract
Amino acid transporters (AATs) are essential integral membrane proteins that serve multiple roles, such as facilitating the transport of amino acids across cell membranes. They play a crucial role in the growth and development of plants. Phaseolus vulgaris, a significant legume crop, serves as a valuable model for studying root symbiosis. In this study, we have conducted an exploration of the AAT gene family in P. vulgaris. In this research, we identified 84 AAT genes within the P. vulgaris genome sequence and categorized them into 12 subfamilies based on their similarity and phylogenetic relationships with AATs found in Arabidopsis and rice. Interestingly, these AAT genes were not evenly distributed across the chromosomes of P. vulgaris . Instead, there was an unusual concentration of these genes located toward the outer edges of chromosomal arms. Upon conducting motif analysis and gene structural analysis, we observed a consistent presence of similar motifs and an intron-exon distribution pattern among the subfamilies. When we analyzed the expression profiles of PvAAT genes, we noted tissue-specific expression patterns. Furthermore, our investigation into AAT gene expression under rhizobial and mycorrhizal symbiotic conditions revealed that certain genes exhibited high levels of expression. Specifically, ATLa5 and LHT2 was notably upregulated under both symbiotic conditions. These findings point towards a potential role of AATs in the context of rhizobial and mycorrhizal symbiosis in P. vulgaris, in addition to their well-established regulatory functions.
Similar content being viewed by others
Introduction
Transporters are a class of membrane proteins that transport nutrients, hormones, and metabolites for the purposes of growth, development, and adaptation to stresses across membranes (Bo et al. 2017). Transmembrane transporters facilitate substrate transport across biological membranes through their transmembrane segments (TMS), which are also known as transmembrane domains (TMDs). Nitrogen is one of the most important nutrients for plant growth and is a major component of many cellular compounds. Plants absorb organic and inorganic nitrogen from the rhizosphere through the root system (Smith and Chalk 2021). In soil, inorganic nitrogen is usually found in the forms of nitrate and ammonium, while organic nitrogen usually exists in the forms of free amino acids, urea, and short peptides (Shen et al. 2003; Kojima et al. 2007; Tegeder and Rentsch 2010; Kotur et al. 2013; Forde 2014).
Plants acquire inorganic nitrogen, rapidly incorporating it into amino acids like glutamine or glutamic acid. Amino acid synthesis occurs in plastids of mesophyll cells and are transported through xylem sap and phloem to support the protein biosynthesis, growth and reproduction (Rentsch et al. 2007; Masclaux-Daubresse et al. 2010; Pratelli and Pilot 2014). Amino acid transporters play a crucial role in regulating processes like leaf senescence, seed germination, and stress response (Frommer et al. 1993; Tegeder and Masclaux-Daubresse 2018).
The uptake and transport of nitrogen-containing molecules by plants is facilitated by an array of amino acid transporters, which have also been known to play a major role in distributing nitrogen throughout the whole plant (Tegeder 2012; Dinkeloo et al. 2018). Most of the amino acid transporters described in plants are proton–amino acid symporters, which is an active transport (Bush 1993). The majority of active transport is driven by proton gradient across the cellular/subcellular membrane. Hence, pH changes in subcellular compartments could regulate the transport activities and determine the direction of transport. The pH change also implies a change in electrical potential difference across the biological membrane. The amino acid uptake is coupled to the proton electrochemical potential difference that is maintained through p-type proton-pumping ATPase (Sze et al. 1999). Taken together these factors influence the transport efficiency and, thus, bring forth the physiological regulations.
Amino acid transporters (AATs) are studied extensively in yeast, mammals and plants are found to be encoded by multiple gene families that encode different classes of amino acid transporters. In plants AATs include two main families, classified based on sequence similarity and uptake properties. The amino acid/auxin permease (AAAP) family, also called the amino acid transporter (ATF) family, and the Amino Acid-Polyamine-Organocation (APC) family (Fischer et al. 1998; Okumoto and Pilot 2011). AAAP family includes 6 subfamilies, amino acid permeases (AAPs), lysine and histidine transporters (LHTs), proline transporters (ProTs), γ-aminobutyric acid transporters (GATs), auxin transporters (AUXs), and aromatic and neutral amino acid transporters (ANTs) (Ortiz-Lopez et al. 2000; Saier et al. 2009). APC subfamily is grouped into three subfamilies cationic amino acid transporters (CATs), amino acid/choline transporters (ACTs) and polyamine H+-symporters (PHSs).
The AAT genes are identified by the characteristic domains PF01490 (Aa_trans) and PF00324 (Aa_permease). In the AAT superfamily, Arabidopsis is found to have 8 genes in Amino acid permease (AAP) subfamily and 6 of them have been functionally characterized. AtAAP1-8 have been found to be predominately transporting neutral and acidic amino acids with moderate affinity except AtAAP3 and AtAAP5 which also transport basic amino acids (Fischer et al. 1995; Rentsch et al. 2007; Forsum et al. 2008). AtAAP1 was found to be expressing in epidermis and was essential for amino acid import into embryo, AtAAP8 also was found to be essential for uptake of amino acids into the endosperm during early seed development (Fischer et al. 1995; Hirner et al. 1998; Schmidt et al. 2007; Forsum et al. 2008; Sanders et al. 2009). AtAAP2 and AtAAP6 have been suggested to function in xylem-phloem transfer (Okumoto et al. 2002; Zhang et al. 2010). AtAAP3 and AtAAP5 expressing in the roots and root vasculature could be involved in amino acid uptake from the rhizosphere (Birnbaum et al. 2003; Okumoto et al. 2004). Furthermore, noteworthy members of the AAP subfamily from different species have been documented, including S. tuberosum AAP1 (Koch et al. 2003), PvAAP1 (Tan et al. 2008), OsAAP8, and OsAAP15 (Zhao et al. 2012). The potential significance of P. trichocarpa AAP11 in xylogenesis, possibly involving proline provision in Populus, has been suggested (Couturier et al. 2010).
Lysine-histidine transporters (LHTs) facilitate the transport of lysine, histidine, as well as neutral and acidic amino acids (Perchlik et al. 2014). AUX subfamilies are major auxin influx carriers, AUX1 is found to be involved in regulating root gravitropism and root hair development. AtANT1 is an aromatic and neutral amino acid transporter, ProT are Proline specific transporters were found be expressing in flowers and under salt-stress respectively in Arabidopsis (Chen et al. 2001; Birnbaum et al. 2003; Lee and Tegeder 2004; Swarup et al. 2004; Hirner et al. 2006; Foster et al. 2008; Zhang et al. 2010; Péret et al. 2012). Among APC family, various CAT subfamily members have been functionally characterized in Arabidopsis. AtCAT5, AtCAT8, AtCAT1 and AtCAT2 have been implicated in uptake of amino acids at the leaf margin, root, shoot meristems and tonoplasts to regulate soluble leaf amino acid concentration (Frommer et al. 1995; Su et al. 2004; Yang et al. 2014).
Though, functional characterization of AAT genes is limited to few plant species, genome wide identification studies have been carried out extensively and a varied number of genes were identified in Arabidopsis, rice, Populus, wheat, potato, rapeseed, soybean, strawberry, cotton, tomato and poplar (Rentsch et al. 2007; Zhao et al. 2012; Wu et al. 2015; Ma et al. 2016; Cheng et al. 2016; Wan et al. 2017; Tian et al. 2020; Liang et al. 2020; Yang et al. 2021; Omari 2021; Kong et al. 2022; Du et al. 2022). Genome wide identification of the gene families and annotation plays a pioneering step towards functional characterization of these genes in a plant species. When we investigate the list of species where AATs have been identified show a glaring lack of such studies in legumes. Until now AAT gene family was identified in G. max (Liang et al. 2020) and the current study adds Phaseolus vulgaris as the second to the list. Research on crucial gene families like AATs holds significance because, as transporters, they may play a vital role in transporting nutrients associated with Biological Nitrogen Fixation (BNF) and mycorrhization.. The biological nitrogen fixation initiates by the infection of legume host by bacteria rhizobia, resulting formation of root nodules. Bacteroids, the differentiated form of rhizobia reduces N2 to ammonia in the symbiosome which is later assimilated to glutamine, asparagine, and other amino acids (Atkins et al. 1982; White et al. 2007). While temperate legumes have aspargine as the major N transport form, tropical legumes produce ureides. Thus, produced amino acids are exported from nodules via xylem to the shoot or through phloem to the root for metabolism and growth (Pate et al. 1969, 1979; Atkins et al. 1982; Tegeder 2014). To transport these amino acids the role of AATs is indispensable. Moreover, arbuscular mycorrhizal (AM) fungi, besides transporting inorganic phosphate (Pi), also contribute to soil nutrient cycling by acquiring both organic and inorganic nitrogen (N) from the soil (Lanfranco et al. 2011). They absorb inorganic N compounds like ammonium, facilitated by various identified ammonium transporters (Gomez et al. 2009; Guether et al. 2009b; Kobae et al. 2010). However, besides LHT1.2 in Lotus japonicus (Guether et al. 2011), no other AAT has been recognized for amino acid transport during AM symbiosis. Hence, in our present investigation we carried out an extensive analysis to identify AATs in P. vulgaris and classified them into 12 subfamilies based on their sequence and structural conservation as compared to Arabidopsis AATs. Further, we have researched into the differential expression patterns of P. vulgaris AATs using the data from the public data base and in house transcriptomic analysis.
Materials and methods
Identification of the AAT family members in P. vulgaris
Phaseolus vulgaris AAT sequences were identified based on the Arabidopsis, Oryza sativa and Glycine max AAT sequences. BLASTN and BLASTP search in Phytozome 13/Phaseolus vulgaris v2.1 genome database (https://phytozome-next.jgi.doe.gov/). In addition to Phytozome, Legume Information System (https://legumeinfo.org) genome databases was used to verify the AAT homologs. The conserved domains of the P. vulgaris AAT genes identified above were analyzed, and the genes that did not contain the PF01490 and PF00324 domains were removed. The nucleic acid and peptide sequences were retrieved from the online tool PhytoMine, accessible through the plant comparative genomics portal Phytozome 13, in preparation for subsequent analysis and annotation. Basic information on physico-chemical properties of AAT genes, including chromosome number, length of gene, CDs, peptide; isoelectric point (pI) and molecular weight (MW) was predicted through the Phytozome and ExPASy (https://web.expasy.org/protparam/). The subcellular localization of the complete protein sequences of AATs was predicted using the in silico tool Plant-mPLoc (version 2.0), available at http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/.
Chromosomal location of Phaseolus vulgaris AAT genes
The chromosomal localization of AAT gene family members was confirmed using the Legume Information System database (accessible at https://legumeinfo.org). Gene nomenclature followed the guidelines set forth by Quezada et al. (2019). Centromere locations and scales were determined based on the methods outlined by Fonsêca et al. (2010) and Wang et al. (2016), respectively.
Phylogenetic analysis of P. vulgaris AAT superfamily genes
We utilized the complete amino acid sequences of AAT protein genes for conducting phylogenetic analysis. Initially, all obtained sequences (A. thaliana, O. sativa, G. max and P. vulgaris) were aligned using the Clustal Omega tool (https://www.ebi.ac.uk/Tools/msa/clustalo/) with default parameters. Using Molecular Evolutionary Genetics Analysis (MEGA 11), we performed phylogenetic analysis on the aligned sequences employing the neighbor-joining (NJ) method. The JTT + I + G substitution model was applied, along with 1000 bootstrap replicates and default parameters. The P. vulgaris AAT genes were classified into different groups according to the topology of the phylogenetic tree.
Gene structure, conserved motif and promoter analysis of AATs
To visualize gene features, we utilized DNA sequences of AATs obtained from Phytozome 13. This included the structural information and positioning of exons, introns, and untranslated regions, which were reconstructed using the Gene Structure Display Server2.0 (GSDS2.0; http://gsds.gao-lab.org/Gsds_about.php) as described by Hu et al. (2015).
We utilized the Multiple Expectation Maximization for Motif Elicitation (MEME Suite 5.5.4) online program (https://meme-suite.org/meme/tools/meme) with specified parameters: number of repetitions = any, maximum number of motifs = 20, and optimal motif length range = 6 to 100 residues, to identify conserved motifs within the amino acid transporter (AAT) protein family in P. vulgaris. Following this, the more pertinent motifs underwent further scrutiny using the MOTIF Search interface (https://www.genome.jp/tools/motif/). Additionally, the respective identification numbers were acquired from the Pfam database (http://pfam.xfam.org/), streamlining the process of pinpointing conserved domains across various subfamilies.
The 2 kb DNA sequences preceding the start codon of amino acid transporter genes were retrieved from the Phytozome 13 database. Plant transcriptional regulatory elements (cis-elements) within the promoter sequences were analyzed using the PlantPAN 3.0 database (Chow et al. 2019) (http://plantpan.itps.ncku.edu.tw/plantpan4/index.html).
Gene Ontology and TMH analysis
To gain functional insights into the identified AAT genes in P. vulgaris, Gene Ontology (GO) analysis was performed. The sequences of the identified AAT genes were submitted to the Blast2GO tool (https://www.blast2go.com/) for GO term annotation. The annotated GO terms provide information on the biological processes, molecular functions, and cellular components associated with each gene, and the results are represented graphically.
We employed the DeepTMHMM server (https://dtu.biolib.com/DeepTMHMM) to predict the topology of transmembrane domains, encompassing both alpha-helical and beta-barrel structures, within the amino acid transporter peptide sequences.
Transcriptomic profiling and quantitative reverse transcription PCR analysis
Data on the differential expression of AAT genes in 24 tissues of P. vulgaris were obtained from both the PvGEA website (https://www.zhaolab.org/PvGEA/ )and Phytozome 13 (P. vulgaris v2.1). Additionally, data on AAT gene expression in arbuscular mycorrhiza (Rhizophagus irregularis) colonized roots was retrieved from our prior publication (Nanjareddy et al. 2017b). The construction of the heat map was performed using Fragments per Kilobase of Exon Model per Million Reads Mapped (FPKM) values for each AAT gene, with the analysis conducted in the R programming language.
To validate the RNA-seq data, we surface-sterilized P. vulgaris L. cv. Negro Jamapa seeds and germinated them as outlined by Nanjareddy et al. (2017a). In the experimental setup, two-day-old germinated seedlings were transferred into sterile vermiculite. Subsequently, inoculation with either R. irregularis or R. tropici was performed to assess AAT gene expression (Table S4) under mycorrhizal or root nodule symbiotic conditions following the procedure described by Nanjareddy et al. (2017b). Total RNA extraction and subsequent RT–qPCR analysis was performed in accordance with the protocol established by Arthikala et al. (2021). Relative gene expression levels were calculated using the 2−ΔCT method, with ΔCT = CTgene – CTreference gene. P. vulgaris EF1α and IDE were used as internal controls (Islas-Flores et al. 2011; Borges et al. 2012).
Results
Identification of PvAAT protein orthologues in P. vulgaris genome
A total of 84 AATs were identified in P. vulgaris genome database using HMM model. We used Arabidopsis, O. sativa and G. max AAT sequences to conduct a BLASTN and BLASTP search in Phytozome 13 database. A total of 84 AAT gene family members were identified based on the presence of PF01490 and PF00324 domains and care was taken to made sure that the sequence was complete when compared to Arabidopsis AATs. In Arabidopsis, O. sativa and G. max, 63, 85 and 189 AATs have been reported respectively (Rentsch et al. 2007; Zhao et al. 2012; Kong et al. 2022). The AATs were classified into subfamilies depending on the sequence homology to other species and naming of the members of each subgroup was based on their appearance in the genome from the chromosomal short arm towards the long arm, starting from proximal to distal ends of the respective arms. We analyzed the physico chemical properties of AAT genes in P. vulgaris using online database. According to the assessment (Table 1), the lengths of AAT genes varied considerably with the largest protein CAT1 (641 aa) with the molecular weight (68.13 kD) and smallest CAT3 (278 aa) and the molecular weight (30.86 kD). Isoelectric point was highest in ProT3 (9.67) and lowest in ATLb6 (4.88) and prediction of subcellular localization showed most of the proteins in the plasma membrane and some proteins in the Golgi apparatus or chloroplast (Table S1).
Phaseolus vulgaris AAT classification and chromosomal localization
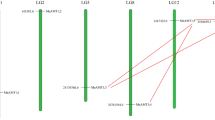
The superfamily of 84 AATs is classified into 12 subfamilies with an unequal distribution in each subfamily as follows, 13 genes in cationic amino acid transporters (CAT), 5 genes in the polyamineH+-symporters (PHS), 3 genes in amino acid/choline transporters (ACT), one gene of tyrosine specific transporter (TTP), 16 genes of amino acid transporter-like proteins (ATL) in turn classified into 2 subfamilies: 8 ATLa y 8 ATLb, 2 genes of aromatic and neutral amino acid transporter (ANT), 7 genes Auxin transporters (AUX), 16 genes of amino acid permeases (AAP) (Tabla 8), 5 genes of γ-aminobutyric acid (GABA) transporter (GAT), 3 genes de proline transporters and finally 13 genes of lysine/histidine transporter (LHT). The AATs in P. vulgaris were unevenly distributed among all the 11 chromosomes (Fig. 1). Chromosome 8 had a maximum of 14 genes followed by Chr1 with 12, Chr9 with 11, Chr2 and 7 with 9 genes each, Chr3 with 7 and Chr10 had a minimum of 1 gene. Most of the chromosomes showed accumulation of the genes towards the outer edges of the chromosomal arms except for the submetacentric chromosome Chr9 where the long arm had 13 genes spread out (Fig. 1).
Phylogenetic analysis of AATs
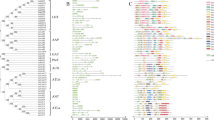
To explore the evolutionary relationship of the AAT superfamily genes, a phylogenetic tree was constructed for 84 P. vulgaris AAT protein sequences. A phylogenetic tree developed using MEGA 11 with the neighbour-joining (NJ) method classified the AAT proteins into 12 clades (Fig. 2) which is like Arabidopsis and other plant species where the AAT superfamily has been identified and annotated. These 12 clades could be broadly classified into 2 families AAAP and APC family were, the AAAP family consists of 54 AATs, including seven distinct subfamilies: 8 genes in ATLb, 2 genes of aromatic and neutral amino acid transporter (ANT), 7 genes Auxin transporters (AUX), 16 genes of amino acid permeases (AAP), 5 genes of γ-aminobutyric acid (GABA) transporter (GAT), 3 genes de proline transporters (ProT) and finally 13 genes de lysine/histidine transporter (LHT). The APC family is comprised of 5 subfamilies including 30 AATs including 13 genes in cationic amino acid transporters (CAT), 5 genes in the polyamine H+-symporters (PHS), 3 genes in amino acid/choline transporters (ACT), one gene of tyrosine specific transporter (TTP), 8 genes of amino acid transporter-like proteins (ATLa) (Fig. 2).
Phylogenetic analysis of the Amino Acid Transporters (AAT) superfamily in Phaseolus vulgaris. The amino acid sequences of 84 AAT genes identified in the Phytozome database. The phylogenetic tree was constructed using MEGA 11 software with the Neighbor-Joining tree method with 1000 bootstrap values. Subfamilies within the AAT superfamily are distinguished by different colors
Further, to establish evolutionary relationship and identify orthologous genes between monocots, dicots and legumes, non-legumes, a phylogenetic tree was developed using protein sequences from A. thaliana, O. sativa, G. max and P. vulgaris (Fig. S1). The combines phylogenetic tree developed the classification of the AAT proteins slightly different than what was seen in P. vulgaris. The classification of subfamilies was similar in all the species compared forming 12 clades indicating the formation of AATs before divergence of the monocots and dicots. There was a considerable variation in the total number of proteins in each of the subfamilies specifically, in AAPs where P. vulgaris has 16 genes, Arabidopsis, O. sativa and G. max genomes had 8, 19 and 34 genes each. Further total number of genes in APC subfamily are 25, 15, 35 and 30 genes in O. sativa, Arabidopsis, G. max and P. vulgaris respectively.
Motif conservation and gene structure
MEME tools were used to identify motifs shared in AATs of Phaseolus. A total of 20 motifs were identified among both AAAP and APC families of AATs. The distribution of motifs in all P. vulgaris AATs are as shown in Fig. 3. Some of the motifs were well conserved and widely distributed among P. vulgaris AATs in AAAP family while some were specific to each subfamily. The frequent motif in most of the AATs was motif 2 followed by motif 1 and 6 which could be found in 7 subfamilies, all of which belonged to subfamilies ATL, ANT, AUX, AAP, GAT, LHT, ProT belonged to the family AAAP. Among AAAP subfamilies AAP had a characteristic motif 7, ATLa, ATLb had motif 14 and AUX had the motifs 10, 12, 13 and 19 restricted to the subfamily alone. Among APC subfamilies, CAT had characteristic motifs 15, 16 and 17, PHS and ACT had only motif 9. In general, the subfamilies in AAAP shared more motifs ranging from ATL with 4 motifs to LHT with 10 motifs. While APC family members had a maximum of 4 motifs in CAT and ATLa and ACT and PHS subfamilies had only one motif (Fig. 3).
The conserved motifs identified through MEME-suite analysis of Amino Acid Transporter (AAT) proteins obtained from Phaseolus vulgaris. A total of 20 distinct motifs are identified across various AAT sub-families. Each color within the predicted protein domains represents a unique motif, providing insights into the structural and functional diversity of AAT proteins in P. vulgaris
To understand the gene structure of AAT transporters in P. vulgaris, we compared the full-length cDNA sequences to the corresponding genomic DNA sequences using GSDS 2.0. Among all the P. vulgaris AATs, 9 genes did not have any introns, which included 4 genes of PHS subfamily, 3 genes from CAT subfamily and one gene each from ATL and ANT subfamilies. Unlike in AAT family genes in other species, the number of introns within the same subfamily varied greatly. Among CAT subfamily, the intron numbers varied from 1 to 13, ATL subfamily had 0-10 introns, AAP and GAT had 4-6, LHT from 4-7 introns (Fig. 4).
Gene structure analysis of the Amino Acid Transporters (AAT) superfamily genes in Phaseolus vulgaris organized by subfamilies. Coding sequences (CDs) and exons are represented by green bars, introns with black lines and untranslated regions (UTR's) in the ´upstream´ (5')/´downstream´ (3') direction with red boxes. The scale bar is shown at the bottom (Kb). Gene structure maps were drawn with the Gene Structure Display Server 2.0. The scale bar is shown at the bottom (Kb)
AATs being the membrane bound proteins, it was crucial to understand the distribution of transmembrane domains among the subfamilies. We developed the transmembrane domain structures using DeepTMHMM software. The results showed that CAT subfamily had lowest of 6 TMDs in CAT3 and a maximum of 15 TMDs were found in CAT7 and 8 with the remaining genes having 10-14 TMDs. PHS and ACT subfamily had 10-12 TMDs, TTP subfamily had 11, ATLa subfamily had 11-12, ATLb ranged between 6 to 11 and ANT, AUX had 10 TMDs each. The multiple sequence alignment of subfamilies showed highly conserved nature of TM domains in all the subfamilies (Fig. S2).
Cis – acting elements identification and gene ontology
To identify the putative cis-element in the promoter region of the AAT genes, the first 2.0 kb of AAT genes of cis-elements was analyzed using the Plantpan database. The highest number of cis-elements were WRKY, AT-Hook, MYB and Homeodomain which are involved in regulation of gene expression, influence in biological processes such as growth, development, and response to abiotic and biotic stresses. Apart from these, several other cis-elements such as stress inducible AP2; ERF, lateral organ growth regulating B3, ABA-responsive element (ABRE) binding bZIP, plant growth and development and stress responsive C2H2, cell elongation regulating bHLH, calcium signal responsive CG-1 are found frequently. Further, ERFs involved in regulation of developmental processes in response to stimuli, and NAC, C2H2 and bHLH involved in the pathogen response, cell proliferation and development are some of the important cis - elements found on the promoter regions of AATs (Table S2). Eight of the identified cis elements, including AP2, ERF, bHLH, bZIP, C2H2, MYB/SANT, NAC, and WRKY, have previously been documented as specific to root nodule symbiosis (RNS). According to the GO molecular functional analysis and biological processes, 96% of them to be involved in activities related to transmembrane transporters involved in transport of amino acids, organic acids, nitrogenous compounds. Some are involved in response to various stimuli such as hormones, more specifically to auxins, organic compounds, chemicals etc. (Fig. 5).
Expression analysis of AAT genes in different Phaseolus vulgaris tissues
To principally investigate the probable gene function of AATs in various Phaseolus tissues, the tissue−specific expression data of AATs were first downloaded from the PvGEA: Common bean Gene expression atlas and Network analysis, which includes the transcriptional profiles of different P. vulgaris tissues (Table S3) including young leaves, mature leaves, roots, flower, pods, seeds and nodules. As it is evident from the heat map (Fig. 6) most of the AATs had low to very low expression levels in all the tissues studied. Among APC family, CAT subfamily genes had very low expression except for CAT8 and CAT10 which showed high expression specifically in seeds, CAT7 was found to be expressing in pods. Among PHS subfamily, PHS3 had very high expression in leaves derived from Rhizobium inoculated plants. ACT subfamily showed low expression in leaf tissues, ACT2 specifically showed very high expression in young leaf and ACT3 in young roots. ANT1 showed pleotropic expression in all the tissues listed. ATLa5 gene showed elevated expression in leaves and young root, while other ATLa and ATLb genes didn’t show significant expression. AAP1 and AUX3 were the genes showing positive expression in most of the tissues studied. AUX genes were positively regulated in pods and ProT subfamily genes expressed mostly in young shoots, pods, and leaves. LHT2 and LHT13 showed very high expression in young root and leaves respectively. GAT subfamily had very high expression root tissues (Fig. 6).
In silico expression profiles of P. vulgaris AATs. Heat map expression profiles of AAT family genes in various tissues of P. vulgaris. Transcriptome data spanning diverse tissues (Table S3) were extracted from both Phytozome 13 (P. vulgaris v2.1) and the P. vulgaris Gene Expression Atlas (PvGEA). The generation of the heat map was facilitated by utilizing the Fragments per Kilobase of Exon Model per Million Reads Mapped (FPKM) values for each AAT gene, a process executed through the programming language R
Expression analysis of AAT genes during rhizobial and mycorrhizal symbiosis
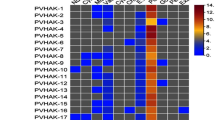
Further, analysis of AAT gene expression specifically in Rhizobium inoculated and mycorrhiza colonized P. vulgaris tissues was done using the PvGEA: Common bean Gene expression atlas and Network analysis database and the data from our previous transcriptomic analysis (Nanjareddy et al. 2017b). Gene expression data for root nodule symbiosis were downloaded from the PvGEA: Common bean Gene expression atlas and Network analysis for 5 days post inoculation (dpi) and 21 dpi. The data revealed very interesting findings as presented in shown in the Fig. 7a. At 5 dpi, among 84 AATs, 36 genes had no, or very low expression (< 2) and 10 genes were highly expressed. However, AAP14 (60.9) and LHT2 had the highest expression (136.3) when compared to any other gene. At 21 dpi, 49 genes had no, or very low expression (< 2) which amounts to more than half of ATT genes. The highest gene expression at 21 dpi was observed in ATLa5 (108.6), followed by LHT2 (80.3) among the top 9 expressed genes. To analyze the differential expression of AATs under mycorrhizal symbiotic condition we used the transcriptomic analysis previously published from our group (Fig. 7a). The data shows that LHT2 as very highly expressed in comparison to all the other AATs followed by ATLa5, AAP3, AAP13, AAP16 and GAT2. Among all the AATs, 33 genes did not or very low express (< 2) in mycorrhizal symbiotic condition.
Analysis of AAT gene expression in response to root symbionts in P. vulgaris. a) Heat maps showing the differencial AAT gene expression patterns specific to rhizobial and mycorrhizal colonization nodules and roots, respectively. Colour bar shows the fold-change range, with yellow and blue representing upregulation and downregulation, respectively. b) Validation of the transcriptomic data by RT-qPCR analysis. The expression of 8 AAT genes in nodules, and mycorrhized roots was quantified by RT-qPCR. The data are the averages of three biological replicates (n > 9). Error bars represent means ± Standard error mean (SEM)
Subsequently, RT−qPCR was performed to validate the expression data retrieved from PvGEA and RNA-seq in P. vulgaris. The transcript accumulation of random eight genes viz., CAT4, PHS5, ATLa5, AUX6, AAP14, GAT2, ProT3 and LHT2 were analyzed at 5 dpi and 21 dpi and 14day mycorrhizal colonized root tissues of P. vulgaris (Fig. 7b). The expression of the chosen genes was consistent with the RNA-seq data (Fig. 7a, b). Taken together, we propose that LHT2 and ATLa5 might be involved in root nodule symbiosis both at early and late stages and in mycorrhizal symbiosis. Further, AAP subfamily genes could be significant to carry out functional genomic studies during P. vulgaris symbiotic associations.
Discussion
Amino acid transporters in plants are one of the largest gene families playing crucial roles during seed germination, growth and development of the plants, seed formation, biotic and abiotic stress through transport of amino acids (Yao et al. 2020). The inorganic salts of nitrogen acquired from the soil solution, these compounds are incorporated into amino acids in root and mature leaves in all the plant species. In legumes, through biological nitrogen fixation a variety of amino acids are synthesized and are transported to the sink organs through xylem and phloem. In any of these scenarios the role of amino acid transporters is indispensable. Though there has been extensive research on genome wide identification of AATs in plants, there is limited research in legumes. Herein, we identified and characterized AAT gene family members in P. vulgaris through genome wide analysis and differential gene expression in P. vulgaris tissues, finally AAT expression pattern in root nodule and mycorrhizal symbiotic condition.
AAT gene family in plants is classified as AAAP and APC families, with 54 and 30 genes under each family. AAAP family is further divided into 7 subfamilies, ATLb, ANT, AUX, AAP, GAT, ProT and LHT. APC family is divided into CAT, PHS, ACT, TTP and ATLa subfamilies. The characteristic motifs for identification of AAT gene family members were PF01490 (Aa_trans) and PF00324 (Aa_permease). Studies published so far have identified a varied number of AAT gene family members such as Arabidopsis (63), rice (85), wheat (296), potato (72), rapeseed (203), soybean (189), strawberry (45), cotton (Gossypium hirsutum - 190, G. barbadense - 190, G. arboretum - 101, and G. raimondii - 94), tomato (88) and poplar (83) (Rentsch et al. 2007; Zhao et al. 2012; Wu et al. 2015; Cheng et al. 2016; Ma et al. 2016; Wan et al. 2017; Liang et al. 2020; Tian et al. 2020; Yang et al. 2021; Omari 2021; Kong et al. 2022; Du et al. 2022). In the current study we have identified 84 AAT genes in P. vulgaris which is like rice, tomato and poplar. These genes were found to be distributed on all the 11 chromosomes of the P. vulgaris genome however, a characteristic clustering of genes towards the outer edges of most of the chromosomes was noticed. Phylogenetic relationships with orthologs in comparison to Arabidopsis and rice were useful to identify the genes in subfamilies. The major division of the AATs in P. vulgaris was into 2 families as AAAP and APC which were further classified into 12 clades conserving the major clade structure as in other species. The number of genes in each subfamily varied as compared to other.
A total of 20 conserved motifs were identified in P. vulgaris AAT family. Among these, motif 1, 2 and 6 were found in most of the subfamilies belonging to AAAP subfamily. Among APC subfamilies, the motif conservation was rather specific to subfamilies than to the group of subfamilies under APCs. Gene structure analysis plays a crucial role in understanding the functional role of the genes. Gene family evolution is well represented through variation in gene structure (Javelle et al. 2011; Du et al. 2012; Hudson and Hudson 2015). Our findings show that multiple AAT genes were devoid of introns. The feature is common to other AAT genes in Arabidopsis, rice and soyabean. In summary, most members of the subfamilies shared similar intron/exon structures and gene length. Such conservation of the gene structure in subfamilies imply evolutionary relationships among them. Reports suggest that either complete absence or presence of shorter and fewer introns could be an indication of genes that are effectively expressed under stressful conditions (Heyn et al. 2015). These findings imply a probable rapid induction of ANT2, ATLa4, CAT11-13 and PHS subfamily genes to stress conditions. Further, analysis of trans membrane domains revealed a highly conserved nature of TMDs in each subfamily and this was consistent with the other species compared.
An analysis of cis-elements in the promoter region of Phaseolus AAT transporters showed presence of many cis-elements such as WRKY, AT-HOOK and MYB involved in growth, development, and abiotic and biotic stress response. Other frequently occurring cis-elements were ABA responsive (ABRE), cell elongation and cell proliferation implying the important role played by the amino acid transporters majorly in growth and development related aspects of P. vulgaris. Gene ontology studies further confirmed the biological role of ATTs as transporters involved in transport of amino acids, nitrogenous compounds among others.
When we analyzed the expression pattern of AATs in various tissues of P. vulgaris, we found most of the CAT subfamily genes expressing in shoot tissues, PHS showed very low expression in most of the tissues and PHS genes expressed very high either in roots or leaf tissues. Among AAAP subfamilies, ANT1 expressed in all the tissues, AUX genes had very low expression in most of the tissues, GATs mostly expressed in roots, ProT genes highly expressed in leaves and pods, LHT, AAP and ATL were the least expressed genes in Phaseolus tissues. In contrast, studies on rice revealed LHT1 expression in roots, leaves, and flowers (Wang et al. 2019). Expression analysis of AAP1 in Vicia faba, P. vulgaris, and Pisum sativum indicated specificity to cotyledons, while V. faba AAP3 was expressed in roots, shoots, and pods (Tegeder et al. 2000; Miranda et al. 2001; Tan et al. 2008).
The significance of AAT gene expression in root nodule symbiotic conditions is noteworthy, given the role in biological nitrogen fixation. Limited studies have explored the involvement of AATs in symbiotic processes. Our analysis focused on 5 dpi and 21 dpi, representing early and late stages of RNS. Herein, our analysis showed LHT2 as a gene with high expression at 5 dpi and 21 dpi during rhizobial symbiosis and 3 weeks post inoculation mycorrhizal symbiosis, followed by ATLa5 as a next highly expressed gene. Under rhizobial symbiotic conditions, a total of 37 genes and 48 genes did not show positive regulation at 5 dpi and 21 dpi respectively. Similarly, 20 AAT genes were unresponsive under mycorrhizal symbiosis. In P. sativum PsAAP6 is reported to be a key player in nitrogen retrieval from the apoplasm into inner cortex cells for nodule export (Garneau et al. 2018) Response of AAT gene expression to rhizobial symbiosis is crucial to understand as during biological nitrogen fixation bacteroids, the differentiated rhizobia reduce N2 to ammonia and is later assimilated to glutamine, asparagine, and other amino acids or as ureides (Atkins et al. 1982; White et al. 2007). On the other hand, arbuscular mycorrhizal (AM) fungi can acquire both inorganic and organic nitrogen (N), transporting them through arginine from the extra- to the intraradical mycelium where N is transferred to the plant as inorganic N compounds such as ammonium. Previous studies identified altered expression of various amino acid transporters (AATs) in Lotus japonicus mycorrhizal roots through microarray analysis (Guether et al. 2009a) and functional characterization of LHT1.2 in L. japonicus had demonstrated its role in intricate mechanisms for amino acid reuptake and recycling, specifically in mycorrhizal roots (Guether et al. 2011). While the exploration of amino acid transporters (AATs) in understanding their role in symbiotic associations has been limited, genome-wide identification studies could establish a crucial foundation for selecting candidate genes for further functional characterization.
Data availability
Not applicable
References
Arthikala MK, Nanjareddy K, Blanco L, Affantrange XA, Lara M (2021) Target of rapamycin, PvTOR, is a key regulator of arbuscule development during mycorrhizal symbiosis in Phaseolus. Sci Rep 11:11319
Atkins CA, Pate JS, Ritchie A, Peoples MB (1982) Metabolism and translocation of allantoin in ureide-producing grain legumes. Plant Physiol 70:476–482
Birnbaum K, Shasha DE, Wang JY, Jung JW, Lambert GM, Galbraith DW, Benfey PN (2003) A gene expression map of the Arabidopsis root. Science 302:1956–1960
Bo L, Xu D, Ann HB, Hassan NEH (2017) Advances in methods for identification and characterization of plant transporter function. J Exp Bot 68:4045–4056
Borges A, Tsai SM, Caldas DG (2012) Validation of reference genes for RT-qPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep 31:827–838
Bush DR (1993) Proton-coupled sugar and amino acid transporters in plants. Annu Rev Plant Physiol Plant Mol Biol 44:513–542
Chen LS, Ortiz-Lopez A, Jung A, Bush DR (2001) ANT1, an aromatic and neutral amino acid transporter in Arabidopsis. Plant Physiol 125:1813–1820
Cheng L, Yuan HY, Ren R, Zhao SQ, Han YP, Zhou QY, Ke DX, Wang YX, Wang L (2016) Genome-wide identification, classification, and expression analysis of amino acid transporter gene family in Glycine max. Front Plant Sci 7:515
Chow CN, Lee TY, Hung YC, Li GZ, Tseng KC, Liu YH, Kuo PL, Zheng HQ, Chang WC (2019) PlantPAN3.0: a new and updated resource for reconstructing transcriptional regulatory networks from ChIP-seq experiments in plants. Nucleic Acids Res 47(D1):D1155–D1163
Couturier J, De FE, Fitz M, Wipf D, Blaudez D, Chalot M (2010) PtAAP11, a high affinity amino acid transporter specifically expressed in differentiating xylem cells of poplar. J Exp Bot 61:1671–1682
Dinkeloo K, Boyd S, Pilot G (2018) Update on amino acid transporter functions and on possible amino acid sensing mechanisms in plants. Semin Cell Dev Biol 74:105–113
Du H, Yang SS, Liang Z, Feng BR, Liu L, Huang YB, Tang YX (2012) Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biol 12:106
Du J, Du C, Ge X, Wen S, Zhou X, Zhang L, Hu J (2022) Genome-wide analysis of the AAAP gene family in Populus and functional analysis of PsAAAP21 in root growth and amino acid transport. Int J Mol Sci 24:624
Fischer WN, Andre B, Rentsch D, Krolkiewicz S, Tegeder M et al (1998) Amino acid transport in plants. Trends Plant Sci 3:188–195
Fischer WN, Kwart M, Hummel S, Frommer WB (1995) Substrate specificity and expression profile of amino acid transporters (AAPs) in Arabidopsis. J Biol Chem 270:16315–16320
Fonsêca A, Ferreira J, dos Santos TRB, Mosiolek M, Bellucci E, Kami J, Gepts P, Geffroy V, Schweizer D, Dos Santos KGB et al (2010) Cytogenetic map of common bean (Phaseolus vulgaris L.). Chromosom Res 18:487–502
Forde BG (2014) Glutamate signalling in roots. J Exp Bot 65:779–787
Forsum O, Svennerstam H, Ganeteg U, Naesholm T (2008) Capacities and constraints of amino acid utilization in Arabidopsis. New Phytol 179:1058–1069
Foster J, Lee YH, Tegeder M (2008) Distinct expression of members of the LHT amino acid transporter family in flowers indicates specific roles in plant reproduction. Sex Plant Reprod 21:143–152
Frommer WB, Hummel S, Riesmeier JW (1993) Expression cloning in yeast of a cDNA encoding a broad specificity amino acid permease from Arabidopsis thaliana. Proc Natl Acad Sci USA 90:5944–5948
Frommer WB, Hummel S, Unseld M, Ninnemann O (1995) Seed and vascular expression of a high-affinity transporter for cationic amino acids in Arabidopsis. Proc Natl Acad Sci USA 92:12036–12040
Garneau MG, Tan Q, Tegeder M (2018) Function of pea amino acid permease AAP6 in nodule nitrogen metabolism and export, and plant nutrition. J Exp Bot 69(21):5205–5219
Gomez SK, Javot H, Deewatthanawong P, Torres-Jerez I, Tang Y, Blancaflor EB, Udvardi MK, Harrison MJ (2009) Medicago truncatula and Glomus intraradices gene expression in cortical cells harboring arbuscules in the arbuscular mycorrhizal symbiosis. BMC Plant Biol 9:1–19
Guether M, Balestrini R, Hannah M, He J, Udvardi MK, Bonfante P (2009a) Genome-wide reprogramming of regulatory networks, transport, cell wall and membrane biogenesis during arbuscular mycorrhizal symbiosis in L otus japonicus. New Phytol 182:200–212
Guether M, Neuhäuser B, Balestrini R, Dynowski M, Ludewig U, Bonfante P (2009b) A mycorrhizal-specific ammonium transporter from Lotus japonicus acquires nitrogen released by arbuscular mycorrhizal fungi. Plant Physiol 150:73–83
Guether M, Volpe V, Balestrini R, Requena N, Wipf D, Bonfante P (2011) LjLHT1.2 A mycorrhiza-inducible plant amino acid transporter from Lotus japonicas. Biol Fertil Soils 47:925–936
Heyn P, Kalinka A, Tomancak P, Neugebauer K (2015) Introns and gene expression: cellular constraints, transcriptional regulation and evolutionary consequences. Bio Essays 37:148–154
Hirner A, Ladwig F, Stransky H, Okumoto S, Keinath M, Harms A, Frommer WB, Koch W (2006) Arabidopsis LHT1 is a high-affinity transporter for cellular amino acid uptake in both root epidermis and leaf mesophyll. Plant Cell 18:1931–1946
Hirner B, Fischer WN, Rentsch D, Kwart M, Frommer WB (1998) Developmental control of H+/amino acid permease gene expression during seed development of Arabidopsis. Plant J 14:535–544
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31:1296–1297
Hudson KA, Hudson ME (2015) A classification of basic helix-loop-helix transcription factors of soybean. Int J Genomics 2015:603182
Islas-Flores T, Guillén G, Alvarado-Affantranger X, Lara-Flores M, Sánchez F, Villanueva MA (2011) PvRACK1 loss-of-function impairs cell expansion and morphogenesis in Phaseolus vulgaris L. root nodules. Mol Plant-Microbe Interact 24:819–826
Javelle M, Klein-Cosson C, Vernoud V, Boltz V, Maher C, Timmermans M, Depege-Fargeix N, Rogowsky PM (2011) Genome-wide characterization of the HD-ZIP IV transcription factor family in maize: preferential expression in the epidermis. Plant Physiol 157:790–803
Kobae Y, Tamura Y, Takai S, Banba M, Hata S (2010) Localized expression of arbuscular mycorrhiza-inducible ammonium transporters in soybean. Plant Cell Physiol 51:1411–1415
Koch W, Kwart M, Laubner M, Heineke D, Stransky H, Frommer WB, Tegeder M (2003) Reduced amino acid content in transgenic potato tubers due to antisense inhibition of the leaf H+ /amino acid symporter StAAP1. Plant J 33:211–220
Kojima S, Bohner A, Gassert B, Yuan L, von Wirén N (2007) AtDUR3 represents the major transporter for high-affinity urea transport across the plasma membrane of nitrogen-deficient Arabidopsis roots. Plant J 52:30–40
Kong XC, Yue CP, Zhang TY, Zhou T, Huang JY, Hua YP (2022) Genome-wide identification of 12 amino acid transporter families and molecular characterization of their transcriptional responses to various nutrient stresses in allotetraploid rapeseed. Online PREPRINT. https://doi.org/10.21203/rs.3.rs-1435140/v1.
Kotur Z, Siddiqi YM, Glass AD (2013) Characterization of nitrite uptake in Arabidopsis thaliana: evidence for a nitrite-specific transporter. New Phytol 200:201–210
Lanfranco L, Guether M, Bonfante P (2011) Arbuscular mycorrhizas and N acquisition by plants. In: Polacco JC, Todd CD (eds) Ecological aspects of nitrogen metabolism in plants. Wiley-Blackwell, Chichester, pp 52–68
Lee YH, Tegeder M (2004) Selective expression of a novel high-affinity transport system for acidic and neutral amino acids in the tapetum cells of Arabidopsis flowers. Plant J 40:60–74
Liang W, Ling L, Wang M, Du B, Duan Y, Song P, Zhang L, Li P, Ma J, Wu L, Guo C (2020) Genome-wide identification and expression analysis of the AAAP family in Fragaria vesca. Biotechnol Biotechnol Equip 34:790–799
Ma HL, Cao XL, Shi SD, Li SL, Gao JP, Ma YL, Zhao Q, Chen Q (2016) Genome-wide survey and expression analysis of the amino acid transporter superfamily in potato (Solanum tuberosum L.). Plant Physiol Biochem 107:164–177
Masclaux-Daubresse C, Daniel-Vedele F, Julie D, Fabien C, Laure G, Akira S (2010) Nitrogen uptake, assimilation and remobilization in plants: Challenges for sustainable and productive agriculture. Ann Bot 105:1141–1157
Miranda M, Borisjuk L, Tewes A, Heim U, Sauer N, Wobus U, Weber H (2001) Amino acid permeases in developing seeds of Vicia faba L.: Expression precedes storage protein synthesis and is regulated by amino acid supply. Plant J 28:61–71
Nanjareddy K, Arthikala MK, Aguirre AL, Gómez BM, Lara M (2017a) Plant promoter analysis: Identification and characterization of root nodule specific promoter in the common bean. J Vis Exp 130:56140
Nanjareddy K, Arthikala MK, Gómez BM, Blanco L, Lara M (2017b) Differentially expressed genes in mycorrhized and nodulated roots of common bean are associated with defense, cell wall architecture, N metabolism, and P metabolism. PLoS One 12:0182328
Okumoto S, Koch W, Tegeder M, Fischer WN, Biehl A, Leister D, Stierhof YD, Frommer WB (2004) Root phloem-specific expression of the plasma membrane amino acid proton co-transporter AAP3. J Exp Bot 55:2155–2168
Okumoto S, Pilot G (2011) Amino acid export in plants: a missing link in nitrogen cycling. Mol Plant 4:453–463
Okumoto S, Schmidt R, Tegeder M, Fischer WN, Rentsch D, Frommer WB, Koch W (2002) High affinity amino acid transporters specifically expressed in xylem parenchyma and developing seeds of Arabidopsis. J Biol Chem 277:45338–45346
Omari AF (2021) Genome wide analysis of amino acid transporter superfamily in Solanum lycopersicum. Plants (Basel) 10(2):289
Ortiz-Lopez A, Chang HC, Bush DR (2000) Amino acid transporters in plants. Biochim Biophys Acta 1465:275–280
Pate JS, Atkins CA, Hamel K, McNeil DL, Layzell DB (1979) Transport of organic solutes in phloem and xylem of a nodulated legume. Plant Physiol 63:1082–1088
Pate JS, Gunning BE, Briarty LG (1969) Ultrastructure and functioning of the transport system of the leguminous root nodule. Planta 85:11–34
Perchlik M, Foster J, Tegeder M (2014) Different and overlapping functions of Arabidopsis LHT6 and AAP1 transporters in root amino acid uptake. J Exp Bot 65:5193–5204
Péret B, Swarup K, Ferguson A, Seth M, Yang Y, Dhondt S et al (2012) AUX/LAX genes encode a family of auxin influx transporters that perform distinct functions during Arabidopsis development. Plant Cell 24:2874–2885
Pratelli R, Pilot G (2014) Regulation of amino acid metabolic enzymes and transporters in plants. J Exp Bot 65:5535–5556
Quezada EH, García GX, Arthikala MK, Melappa G, Lara M, Nanjareddy K (2019) Cysteine-rich receptor-like kinase gene family identification in the phaseolus genome and comparative analysis of their expression profiles specific to mycorrhizal and rhizobial symbiosis. Genes 10:59
Rentsch D, Schmidt S, Tegeder M (2007) Transporters for uptake and allocation of organic nitrogen compounds in plants. FEBS Lett 581(2281–2289):36
Saier MH, Yen MR, Noto K, Tamang DG, Elkan C (2009) The transporter classification database: recent advances. Nucleic Acids Res 37:D274–D278
Sanders A, Sanders A, Collier R, Trethewy A, Gould G, Sieker R, Tegeder M (2009) AAP1 regulates import of amino acids into developing Arabidopsis embryos. Plant J 59:540–552
Schmidt R, Stransky H, Koch W (2007) The amino acid permease AAP8 is important for early seed development in Arabidopsis thaliana. Planta 226:805–813
Shen QR, Ran W, Cao ZH (2003) Mechanisms of nitrite accumulation occurring in soil nitrification. Chemosphere 50:747–753
Smith CJ, Chalk PM (2021) Organic N compounds in plant nutrition: have methodologies based on stable isotopes provided unequivocal evidence of direct N uptake? Isot Environ Health Stud 57:333–349
Su YH, Frommer WB, Ludewig U (2004) Molecular and functional characterization of a family of amino acid transporters from Arabidopsis. Plant Physiol 136:3104–3113
Swarup R, Kargul J, Marchant A, Zadik D, Rahman A, Mills R, Yemm A, May S, Williams L, Millner P, Tsurumi S, Moore I, Napier R, Kerr ID, Bennett MJ (2004) Structure-function analysis of the presumptive Arabidopsis auxin permease AUX1. Plant Cell 16:3069–3083
Sze H, Li XH, Palmgren MG (1999) Energization of plant cell membranes by H+-pumping ATPases: regulation and biosynthesis. Plant Cell 11:677–689
Tan Q, Grennan AK, Pélissier HC, Rentsch D, Tegeder M (2008) Characterization and expression of French bean amino acid transporter PvAAP1. Plant Sci 174:348–356
Tegeder M (2012) Transporters for amino acids in plant cells: Some functions and many unknowns. Curr Opin Plant Biol 15:315–321
Tegeder M (2014) Transporters involved in source to sink partitioning of amino acids and ureides: opportunities for crop improvement. J Exp Bot 65:1865–1878
Tegeder M, Masclaux-Daubresse C (2018) Source and sink mechanisms of nitrogen transport and use. New Phytol 217:35–53
Tegeder M, Offler CE, Frommer WB, Patrick JW (2000) Amino acid transporters are localized to transfer cells of developing pea seeds. Plant Physiol 122:319–325
Tegeder M, Rentsch D (2010) Uptake and partitioning of amino acids and peptides. Mol Plant 3:997–1011
Tian R, Yang Y, Chen M (2020) Genome-wide survey of the amino acid transporter gene family in wheat (Triticum aestivum L.): Identification, expression analysis and response to abiotic stress. Int J Biol Macromol 162:1372–1387
Wan YF, King R, Mitchell RAC, Hassani-Pak K, Hawkesford MJ (2017) Spatiotemporal expression patterns of wheat amino acid transporters reveal their putative roles in nitrogen transport and responses to abiotic stress. Sci Rep 7:5461
Wang S, Su JH, Beliveau BJ, Bintu B, Moffitt JR, Wu C, Zhuang X (2016) Spatial organization of chromatin domains and compartments in single chromosomes. Science 353:598–602
Wang X, Yang G, Shi M, Hao D, Wei Q, Wang Z, Fu S, SunY XJ (2019) Disruption of an amino acid transporter LHT1 leads to growth inhibition and low yields in rice. BMC Plant Biol 19:268–311
White J, Prell J, James EK, Poole P (2007) Nutrient sharing between symbionts. Plant Physiol 144:604–614
Wu XM, Kou SJ, Liu YL, Fang YN, Xu Q, Guo WW (2015) Genomewide analysis of small RNAs in nonembryogenic and embryogenic tissues of citrus: microRNA and siRNA-mediated transcript cleavage involved in somatic embryogenesis. Plant Biotechnol J 13:383–394
Yang H, Krebs M, Stierhof YD, Ludewig U (2014) Characterization of the putative amino acid transporter genes AtCAT2, 3 & 4: the tonoplast localized AtCAT2 regulates soluble leaf amino acids. J Plant Physiol 171:594–601
Yang Y, Chai Y, Liu J et al (2021) Amino acid transporter (AAT) gene family in foxtail millet (Setaria italica L.): widespread family expansion, functional differentiation, roles in quality formation and response to abiotic stresses. BMC Genomics 22:519
Yao X, Nie J, Bai R, Sui X (2020) Amino acid transporters in plants: identification and function. Plants 9:972
Zhang L, Tan Q, Lee R, Trethewy A, Lee YH, Tegeder M (2010) Altered xylem-phloem transfer of amino acids affects metabolism and leads to increased seed yield and oil content in Arabidopsis. Plant Cell 22:3603–3620
Zhao H, Ma H, Li Y, Xin W, Jie Z (2012) Genome-wide survey and expression analysis of amino acid transporter gene family in rice (Oryza sativa L.). PLoS One 7:e49210
Acknowledgements
We are grateful to the SGC-LII certified lab (ISO 9001: 2015), ENES-Unidad Leon for providing facilities during this work.
Funding
This work was supported by the Dirección General de Asuntos del Personal Académico, DGAPA/PAPIIT-UNAM grant no. IN216321 & IN217724 to K.N and IN213221 & IN208424 to M.-K.A.
Author information
Authors and Affiliations
Contributions
Conceptualization of the project M.-K.A.; experiments are performed and softwares by K.N., M.F.G-C; methodology, K.N., M.L, validation, K.N., M.-K.A; writing original draft K.N., M.L; supervision, review and editing, M.-K.A.; funding acquisition, K.N., M.-K.A.
Corresponding authors
Ethics declarations
Ethical Approval
Not applicable
Conflict of interest
The authors declare no conflict of interest related to this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary file 1
Fig. S1 Phylogenetic Analysis of the Amino Acid Transporters (AAT) Superfamily in Arabidopsis thaliana, Oryza sativa, Glycine max, Phaseolus vulgaris. The amino acid sequences of 63, 85, 189 and 84 AAT genes identified in the Phytozome database respectively for A. thaliana, O. sativa, G. max, P. vulgari. The phylogenetic tree was constructed using MEGA 11 software with the Neighbor-Joining tree method with 1000 bootstrap values. (PDF 1408 kb)
Supplementary file 2
Fig. S2 Transmembrane topology models applied to PvAAT superfamily. DeepTMHMM analyses of the protein sequences. Pink regions indicate putative transmembrane domains with the relative probability of each indicated on the Y-axis. Yellow and blue regions correspond to predicted extracellular and intracellular segments respectively (PDF 2900 kb)
Supplementary file 3
Table S1 The subcellular localization of AAT genes segregated according to the subfamilies using in silico tool Plant-mPLoc (version 2.0) (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/). Table S2 In silico promoter analyses of PvAAT genes by PlantPAN3.0 (XLSX 30 kb)
Supplementary file 4
Table S3 The Phaseolus vulgaris tissues selected for expression analysis (PDF 77 kb)
Supplementary file 5
Table S4 Primer sequences of Phaseolus vulgaris genes used to perform quantitative RT-PCR (PDF 29 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nanjareddy, K., Guerrero-Carrillo, M.F., Lara, M. et al. Genome-wide identification and comparative analysis of the Amino Acid Transporter (AAT) gene family and their roles during Phaseolus vulgaris symbioses. Funct Integr Genomics 24, 47 (2024). https://doi.org/10.1007/s10142-024-01331-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10142-024-01331-0