Abstract
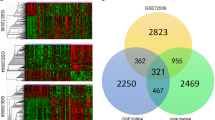
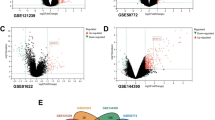
Discoid lupus erythematosus (DLE) is a disorder of the immune system commonly seen in women of childbearing age. The pathophysiology and aetiology are still poorly understood, and no cure is presently available. Therefore, there is an urgent need to explore the underlying molecular mechanisms, as well as search for new therapeutic targets. Gene expression data from skin biopsies samples of DLE patients and healthy controls were downloaded from the Gene Expression Omnibus database. The differentially expressed genes (DEGs) between DLE and healthy control samples were identified by differential expression analysis. Samples were analysed using CIBERSORT to examine the proportion of immune infiltration. Weighted gene co-expression network analysis was used to screen for the module most relevant to immune infiltration. Candidate genes were uploaded to the TRRUST database to obtain the potential transcription factors regulating these genes. Protein–protein interaction (PPI) analysis was performed to obtain the hub genes most associated with immune infiltration among the candidate genes. A total of 273 DEGs were identified between the DLE and healthy control samples. The results of immunoinfiltration analysis showed that the abundances of resting memory CD4 T cells, activated memory CD4 T cells and M1 macrophages were significantly higher, while those of resting infiltration of plasma cells, regulatory T cells and dendritic cells were lower in DLE samples than in healthy control samples. Correlation analysis showed that ISG15, TRIM22, XAF1, IFIT2, OAS2, OAS3, OAS1, IFI44, IFI6, BST2, IFIT1 and MX2 were negatively correlated with the abundances of plasma cells, T-cell regulatory cells and resting dendritic cells and positively correlated with activated memory CD4 T cells and M1 macrophages. Our study shows that these hub genes may regulate DLE via immune-related pathways mediated by the infiltration of these immune cells.







Similar content being viewed by others
Data Availability
The datasets generated during and/or analysed during the current study are available in the Gene Expression Omnibus repository, https://www.ncbi.nlm.nih.gov/geo/.
References
Cervera R, Khamashta MA, Font J et al (1993) Systemic lupus erythematosus: clinical and immunologic patterns of disease expression in a cohort of 1,000 patients: the European working party on systemic lupus erythematosus. Medicine 72:113–124
Chen KL, Krain RL, Werth VP (2019) Advancing understanding, diagnosis, and therapies for cutaneous lupus erythematosus within the broader context of systemic lupus erythematosus. F1000Research. https://doi.org/10.12688/f1000research.17787.1
Chen YC, Chou YC, Hsieh YT et al (2021) Targeting intra-pulmonary P53-dependent long non-coding RNA expression as a therapeutic intervention for systemic lupus erythematosus-associated diffuse alveolar hemorrhage. Int J Mol Sci. https://doi.org/10.3390/ijms22136948
Coit P, Jeffries M, Altorok N et al (2013) Genome-wide DNA methylation study suggests epigenetic accessibility and transcriptional poising of interferon-regulated genes in naïve CD4+ T cells from lupus patients. J Autoimmun 43:78–84. https://doi.org/10.1016/j.jaut.2013.04.003
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57. https://doi.org/10.1038/nprot.2008.211
De S, Barnes BJ (2014) B cell transcription factors: Potential new therapeutic targets for SLE. Clin Immunol 152:140–151. https://doi.org/10.1016/j.clim.2014.03.009
Diamond MS, Farzan M (2013) The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nat Rev Immunol 13:46–57. https://doi.org/10.1038/nri3344
Durosaro O, Davis MD, Reed KB, Rohlinger AL (2009) Incidence of cutaneous lupus erythematosus, 1965–2005: a population-based study. Arch Dermatol 145:249–253. https://doi.org/10.1001/archdermatol.2009.21
Gambichler T, Pätzholz J, Schmitz L, Lahner N, Kreuter A (2015) FOXP3+ and CD39+ regulatory T cells in subtypes of cutaneous lupus erythematosus. J Eur Acad Dermatol Venereol 29:1972–1977. https://doi.org/10.1111/jdv.13123
Garza-Mayers AC, McClurkin M, Smith GP (2016) Review of treatment for discoid lupus erythematosus. Dermatol Ther 29:274–283. https://doi.org/10.1111/dth.12358
Hikami K, Kawasaki A, Ito I et al (2011) Association of a functional polymorphism in the 3′-untranslated region of SPI1 with systemic lupus erythematosus. Arthritis Rheum 63:755–763. https://doi.org/10.1002/art.30188
Jeffries MA, Dozmorov M, Tang Y, Merrill JT, Wren JD, Sawalha AH (2011) Genome-wide DNA methylation patterns in CD4+ T cells from patients with systemic lupus erythematosus. Epigenetics 6:593–601. https://doi.org/10.4161/epi.6.5.15374
Klarquist J, Zhou Z, Shen N, Janssen EM (2016) Dendritic cells in systemic lupus erythematosus: from pathogenic players to therapeutic tools. Mediators Inflamm 2016:5045248. https://doi.org/10.1155/2016/5045248
Ko WC, Li L, Young TR et al (2021) Gene expression profiling in the skin reveals strong similarities between subacute and chronic cutaneous lupus that are distinct from lupus nephritis. J Invest Dermatol 141:2808–2819. https://doi.org/10.1016/j.jid.2021.04.030
Koelsch KA, Webb R, Jeffries M et al (2013) Functional characterization of the MECP2/IRAK1 lupus risk haplotype in human T cells and a human MECP2 transgenic mouse. J Autoimmun 41:168–174. https://doi.org/10.1016/j.jaut.2012.12.012
Kuhn A, Landmann A, Wenzel J (2016) Advances in the treatment of cutaneous lupus erythematosus. Lupus 25:830–837. https://doi.org/10.1177/0961203316641771
Lazear HM, Schoggins JW, Diamond MS (2019) Shared and distinct functions of type I and type III interferons. Immunity 50:907–923. https://doi.org/10.1016/j.immuni.2019.03.025
Li Z, Guo J, Bi L (2020) Role of the NLRP3 inflammasome in autoimmune diseases. Biomed Pharmacother 130:110542. https://doi.org/10.1016/j.biopha.2020.110542
Little AJ, Vesely MD (2020) Cutaneous lupus erythematosus: current and future pathogenesis-directed therapies. Yale J Biol Med 93:81–95
Liu X, Xu S, Li Y, Chen Q, Zhang Y, Peng L (2021) Identification of CALU and PALLD as potential biomarkers associated with immune infiltration in heart failure. Front Cardiovasc Med 8:774755. https://doi.org/10.3389/fcvm.2021.774755
Means TK, Latz E, Hayashi F, Murali MR, Golenbock DT, Luster AD (2005) Human lupus autoantibody-DNA complexes activate DCs through cooperation of CD32 and TLR9. J Clin Investig 115:407–417. https://doi.org/10.1172/jci23025
Méndez-Flores S, Hernández-Molina G, Azamar-Llamas D, Zúñiga J, Romero-Díaz J, Furuzawa-Carballeda J (2019) Inflammatory chemokine profiles and their correlations with effector CD4 T cell and regulatory cell subpopulations in cutaneous lupus erythematosus. Cytokine 119:95–112. https://doi.org/10.1016/j.cyto.2019.03.010
Newman AM, Liu CL, Green MR et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12:453–457. https://doi.org/10.1038/nmeth.3337
Okon LG, Werth VP (2013) Cutaneous lupus erythematosus: diagnosis and treatment. Best Pract Res Clin Rheumatol 27:391–404. https://doi.org/10.1016/j.berh.2013.07.008
Postal M, Vivaldo JF, Fernandez-Ruiz R, Paredes JL, Appenzeller S, Niewold TB (2020) Type I interferon in the pathogenesis of systemic lupus erythematosus. Curr Opin Immunol 67:87–94. https://doi.org/10.1016/j.coi.2020.10.014
Qi YY, Zhao XY, Liu XR, et al (2021) Lupus susceptibility region containing CTLA4 rs17268364 functionally reduces CTLA4 expression by binding EWSR1 and correlates IFN-α signature. Arthritis Res Ther 23:279. https://doi.org/10.1186/s13075-021-02664-y
Rönnblom L (2011) The type I interferon system in the etiopathogenesis of autoimmune diseases. Upsala J Med Sci 116:227–237. https://doi.org/10.3109/03009734.2011.624649
Rönnblom L, Eloranta ML, Alm GV (2003) Role of natural interferon-alpha producing cells (plasmacytoid dendritic cells) in autoimmunity. Autoimmunity 36:463–472. https://doi.org/10.1080/08916930310001602128
Saadeh D, Kurban M, Abbas O (2016) Update on the role of plasmacytoid dendritic cells in inflammatory/autoimmune skin diseases. Exp Dermatol 25:415–421. https://doi.org/10.1111/exd.12957
Salah E (2018) Clinical and dermoscopic spectrum of discoid lupus erythematosus: novel observations from lips and oral mucosa. Int J Dermatol 57:830–836. https://doi.org/10.1111/ijd.14015
Sarkar SN, Bandyopadhyay S, Ghosh A, Sen GC (1999) Enzymatic characteristics of recombinant medium isozyme of 2′-5′ oligoadenylate synthetase. J Biol Chem 274:1848–1855. https://doi.org/10.1074/jbc.274.3.1848
Shapouri-Moghaddam A, Mohammadian S, Vazini H et al (2018) Macrophage plasticity, polarization, and function in health and disease. J Cell Physiol 233:6425–6440. https://doi.org/10.1002/jcp.26429
Sherman BT, Hao M, Qiu J et al (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 50:W216-w221. https://doi.org/10.1093/nar/gkac194
Smith S, Fernando T, Wu PW et al (2017) MicroRNA-302d targets IRF9 to regulate the IFN-induced gene expression in SLE. J Autoimmun 79:105–111. https://doi.org/10.1016/j.jaut.2017.03.003
Teng Z, Lin X, Luan C, Sun Y, Li X (2021) The high expression of miR-564 in patients with systemic lupus erythematosus promotes differentiation and maturation of DC cells by negatively regulating TP53 expression in vitro. Lupus 30:1469–1480. https://doi.org/10.1177/09612033211020367
Wenzel J (2019) Cutaneous lupus erythematosus: new insights into pathogenesis and therapeutic strategies. Nat Rev Rheumatol 15:519–532. https://doi.org/10.1038/s41584-019-0272-0
Yuan Y, Ma H, Ye Z, Jing W, Jiang Z (2018) Interferon-stimulated gene 15 expression in systemic lupus erythematosus : diagnostic value and association with lymphocytopenia. Z Rheumatol 77:256–262. https://doi.org/10.1007/s00393-017-0274-8
Zhang Z, Shi L, Song L, Ephrem E, Petri M, Sullivan KE (2015) Interferon regulatory factor 1 marks activated genes and can induce target gene expression in systemic lupus erythematosus. Arthritis Rheumatol 67:785–796. https://doi.org/10.1002/art.38964
Zhao X, Zhang L, Wang J et al (2021) Identification of key biomarkers and immune infiltration in systemic lupus erythematosus by integrated bioinformatics analysis. J Transl Med 19:35. https://doi.org/10.1186/s12967-020-02698-x
Zheng F, Xu H, Zhang C et al (2021) Immune cell and TCR/BCR repertoire profiling in systemic lupus erythematosus patients by single-cell sequencing. Aging 13:24432–24448. https://doi.org/10.18632/aging.203695
Funding
The authors declare that no funds, grants or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
YH contributed towards conceptualization, methodology and writing- original draft preparation. SL contributed towards conceptualization, methodology and software. YS contributed towards methodology and software. BD contributed towards writing- reviewing and editing, and supervision. XG contributed towards writing- reviewing and editing, and supervision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, Y., Liu, S., Shi, S. et al. Screening of Genes Associated with Immune Infiltration of Discoid Lupus Erythematosus Based on Weighted Gene Co-expression Network Analysis. Biochem Genet (2024). https://doi.org/10.1007/s10528-023-10603-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-023-10603-6