Abstract
Purpose
Previous observational studies have suggested an association between sleep disturbance and metabolic syndrome (MetS). However, it remains unclear whether this association is causal. This study aims to investigate the causal effects of sleep-related traits on MetS using Mendelian randomization (MR).
Methods
Single-nucleotide polymorphisms strongly associated with daytime napping, insomnia, chronotype, short sleep, and long sleep were selected as genetic instruments from the corresponding genome-wide association studies (GWAS). Summary-level data for MetS were obtained from two independent GWAS datasets. Univariable and multivariable MR analyses were conducted to investigate and verify the causal effects of sleep traits on MetS.
Results
The univariable MR analysis demonstrated that genetically predicted daytime napping and insomnia were associated with increased risk of MetS in both discovery dataset (OR daytime napping = 1.630, 95% CI 1.273, 2.086; OR insomnia = 1.155, 95% CI 1.108, 1.204) and replication dataset (OR daytime napping = 1.325, 95% CI 1.131, 1.551; OR insomnia = 1.072, 95% CI 1.046, 1.099). For components, daytime napping was positively associated with triglycerides (beta = 0.383, 95% CI 0.160, 0.607) and waist circumference (beta = 0.383, 95% CI 0.184, 0.583). Insomnia was positively associated with hypertension (OR = 1.101, 95% CI 1.042, 1.162) and waist circumference (beta = 0.067, 95% CI 0.031, 0.104). The multivariable MR analysis indicated that the adverse effect of daytime napping and insomnia on MetS persisted after adjusting for BMI, smoking, drinking, and another sleep trait.
Conclusion
Our study supported daytime napping and insomnia were potential causal factors for MetS characterized by central obesity, hypertension, or elevated triglycerides.
Similar content being viewed by others
Introduction
Metabolic syndrome (MetS), characterized by a cluster of clinical syndromes due to metabolic abnormalities including central obesity, high blood pressure, hyperglycemia, and dyslipidemia [1], is a well-established contributor to cardiovascular disease–related morbidity and mortality [2]. It is estimated that one-quarter of adults worldwide have MetS [3] and the prevalence of MetS is rising [4]. Given the detrimental health consequences and high prevalence of MetS, early identification and modification of risk factors for MetS in high-risk populations are warranted to lower the societal burden of this disease.
Some observational studies supported sleep disturbance as a risk factor for MetS, as poor sleep behaviors were significantly associated with several MetS-related cardiometabolic diseases (i.e., ischemic stroke, coronary artery disease, heart failure, or type 2 diabetes mellitus) [5]. Moreover, abnormal sleep behaviors, such as daytime napping [6], shorter or longer sleep duration [7, 8], insomnia [9], and evening chronotype [10], were found to be disproportionately higher in patients with MetS. Notably, the results of observational studies are susceptible to confounding factors such as smoking [11], alcohol consumption [12], and body mass index (BMI) [13]. Thus, whether these sleep traits have causal effects on the development of MetS remains uncertain.
Mendelian randomization (MR) is a widely utilized method for investigating causal associations. As the distribution of genetic variation at conception is random and precedes acquired exposure, utilizing genetic variants associated with risk factors as instrumental variables can mitigate bias introduced by confounding factors and reverse causation [14]. Therefore, in this study, we performed MR analyses based on large-scale genome-wide association studies (GWAS) data to investigate the causal association of sleep-related traits with the development of MetS and its components.
Method
MR design
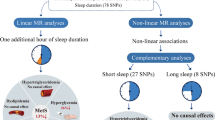
The study design is presented in Fig. 1. We performed a univariable MR analysis to explore the causal effects of sleep traits on MetS and its components. For sleep traits with significant results, the multivariable MR analysis was employed to assess the independent effect of these traits on MetS and further verify the stability of causality by adjusting for common confounders (e.g., smoking initiation, alcohol consumption, and BMI).
The overview of the study design. Assumption 1 represents that genetic instruments and exposure are strongly correlated. Assumption 2 represents that genetic instruments are unrelated to any confounders. Assumption 3 represents that genetic instruments influence the outcome only through exposure. Abbreviation: MR, Mendelian randomization
GWAS data sources
Sleep traits
Genetic instruments for daytime napping were obtained from a GWAS among 452,633 individuals of European ancestry [15]. Daytime napping was evaluated by answering “Do you have a nap during the day?”. Genetic instruments for short sleep duration (≤ 6 h, N = 411,934) and long sleep duration (≥ 9h, N = 339,926) were identified from the largest GWAS in Europe, which assessed sleep duration via questionnaires and actigraphy devices [16]. GWAS data for insomnia were derived from a GWAS meta-analysis that included up to 1,331,010 participants [17]. We selected genetic instruments for insomnia based on 248 independent lead SNPs (p < 5 × 10−8; r2 < 0.1). Chronotype refers to a person’s preference for early or late sleep timing, and the corresponding genetic instruments were extracted from the GWAS with 403,195 individuals of European ancestry [18]. Detailed information of GWAS datasets on sleep-related traits is described in Table S1.
MetS and its components
The primary outcome was MetS, and the secondary outcomes were components of MetS, including waist circumference (WC), hypertension, high-density lipoprotein cholesterol (HDL-C), triglyceride (TG), and fasting blood glucose (FBG).
Summary-level statistics for MetS were extracted from a publicly available GWAS of European ancestry, consisting of 291,107 individuals (59,677 cases and 231,430 controls) [19]. The diagnostic criteria of MetS was according to NCEP [20]. To validate our findings and mitigate potential bias from sample overlap between the exposure and outcome GWASs [21], we included another independent GWAS dataset (N = 461,920) for replication analysis [22]. This data utilized structural equation modeling to investigate the genetic clustering of MetS components.
For secondary outcomes, summary-level statistics for WC were obtained from the Genetic Investigation of ANthropometric Traits (GIANT), consisting of 232,101 participants [23]. Summary-level statistics for hypertension were obtained from Finland nationwide health registries, involving 218,754 individuals [24]. Summary-level statistics for TG and HDL-C were extracted from the global lipids genetics consortium, which included 177,861 and 187,167 individuals, respectively [25]. For FBG, we obtained GWAS data from the meta‐analyses of glucose and insulin‐related traits consortium, comprising 200,622 individuals of European ancestry [26] (Table S1).
Other traits
To reduce the interference of potential confounders, we obtained the genetic instruments for smoking initiation, alcohol consumption, and BMI from the most recent or largest GWASs. Summary-level statistics for smoking initiation and weekly alcohol intake were obtained from the GWAS and sequencing consortium of alcohol and nicotine use, which included 607,291 and 335,394 participants, respectively [27]. Summary-level statistics for BMI were extracted from GIANT, including 681,275 participants of European descent [28] (Table S1).
Genetic instrument selection criteria
Genetic instruments for sleep phenotypes, smoking initiation, alcohol consumption, and BMI were all selected at genome-wide significant threshold (p < 5 × 10−8). Subsequently, SNPs in linkage disequilibrium were removed based on the 1000 Genomes European reference panel (r2 ≥ 0.001). To satisfy the third assumption of the MR design, SNPs strongly associated with outcomes (p < 5 × 10−8) were excluded. Furthermore, we searched for each identified instrumental variable from PhenoScanner v2 to exclude SNPs associated with potential confounding factors (p < 5 × 10−8), particularly those related to components of MetS (Table S2). The F-statistic was calculated to estimate bias due to weak instrumental variables using the equation:\(F={R}^{2}(N-2)/(1-{R}^{2})\), where N is the sample size of the GWASs for exposure and R2 is the proportion of exposure explained by instrumental variables. An F-statistic > 10 was recommended for MR analysis. The MR analyses of long sleep duration with MetS (replication dataset), triglycerides, waist circumference, and HDL-C were excluded because no sufficient genetic instruments were included.
Statistical analyses
The random-effect inverse-variance-weighted (IVW) method was selected as the primary approach for all MR analyses [29]. Moreover, we performed four sensitivity analyses including MR-Egger [30], weighted median [31], MR-Pleiotropy Residual Sum and Outlier (MR-PRESSO) [32], and Robust Adjustment Profile Score (RAPS) methods. The consistency assumptions and strengths of the methods are presented in Table S3. Cochran’s Q statistic was employed to evaluate the heterogeneity of the instrumental variables. The intercept test of MR-Egger regression was utilized to assess the horizontal pleiotropy. Additionally, the presence of sample overlap between sleep traits and MetS (discovery dataset) may cause the effect of weak instrument bias to shift towards the observational correlation. To test the robustness of our observations, we employed more stringent p values (p < 5.0 × 10−9 and p < 5.0 × 10−10, respectively) for instrument construction.
Considering the modest genetic overlap between sleep traits [17], and controlling for potential confounders, we performed multivariable MR analyses to identify whether sleep traits with significant MR results had an independent causal effect on MetS after mutual adjustment with other sleep traits. Meanwhile, multivariable MR analyses were also conducted to estimate the direct effect of these traits on MetS by adjusting for confounders. The IVW and MR-Egger methods were used to estimate the causal effect in multivariable MR analysis.
We applied Bonferroni corrected p value < 0.01 (0.05/5) when analyzing the association between sleep traits and MetS, and p value < 0.002 (0.05/22) when analyzing the association between sleep traits and five MetS components to identify statistical significance. A p value between the corrected threshold and 0.05 was deemed as a suggestive association. All tests were two-sided and conducted using the “TwoSampleMR,” “MRPRESSO,” “mr.raps,” and “MendelianRandomization” packages in the R software (version 4.2.2).
Result
Strength of instrumental variables
Detailed information on IVs is provided in Tables S4–S8. According to the criteria of genetic instrument selection, 103, 22, 7, 141, and 131 SNPs were selected as IVs for daytime napping, short sleep time, long sleep time, insomnia, and chronotype, respectively. IVs used in this study account for 0.1–1.1% variance in sleep traits. The F-statistics for all IVs were greater than 10, suggesting a low likelihood of potential weak instrument bias.
Causal effect of sleep traits on MetS
The results of the MR analysis using the random-effect IVW approach are shown in Fig. 2. Genetically predicted daytime napping and insomnia were significantly associated with an increased risk of MetS in the discovery dataset, and the association remained consistent in the replication dataset. For one-unit increase in log odds of genetically predicted daytime napping, the odds ratio (OR) was 1.630 (95% confidence interval (CI) 1.273, 2.086; p = 1.070 × 10−4) for MetS in the discovery dataset and 1.325 (95% CI 1.131, 1.551; p = 4.786 × 10−4) for MetS in the replication dataset. For one-unit increase in log odds of genetically proxied insomnia, the OR was 1.155 (95% CI 1.108, 1.204; p = 7.340 × 10−12) for MetS in the discovery dataset and 1.072 (95% CI 1.046, 1.099; p = 4.943 × 10−8) for MetS in the replication dataset. No association was found between either sleep duration or chronotype and MetS.
The observed associations were overall consistent across alternative MR methods (Table S9). Cochran’s Q test suggested that there may be some heterogeneity across genetic instruments for sleep traits. However, the MR-Egger intercept test did not find evidence of pleiotropy (Table S10). Furthermore, when more stringent thresholds of instrument construction (p < 5.0 × 10−9 and p < 5.0 × 10−10) were used, the results of daytime napping and insomnia remained stable (Table S11).
Causal effect of sleep traits on components of MetS
In IVW analyses (Table 1), genetically predicted daytime napping was significantly associated with elevated triglycerides (beta = 0.383, 95% CI 0.160, 0.607; p = 7.740 × 10−4) and increased waist circumference (beta = 0.383, 95% CI 0.184, 0.583; p = 1.663 × 10−5). Insomnia increased the risk of hypertension (OR = 1.101, 95% CI 1.042, 1.162; p = 5.573 × 10−4) and was associated with increased waist circumference (beta = 0.067, 95% CI 0.031, 0.104; p = 2.955 × 10−4). Additionally, there were suggestive associations for daytime napping with the risk of hypertension and for insomnia with elevated triglycerides and fasting glucose. Direction of the causal estimates in sensitivity analyses was generally consistent (Table S12). Heterogeneity was detected for most analyses, but no directional pleiotropy was observed in the MR-Egger intercept test (Table S10).
Multivariable MR analyses
Genetically predicted insomnia was still associated with an increased risk of MetS after controlling for daytime napping (IVW: OR = 1.356, 95% CI 1.229, 1.495; p = 1.335 × 10−9). Similarly, the causal association between daytime napping and MetS was robust with adjustment for insomnia (IVW: OR = 1.376, 95% CI 1.073, 1.765; p = 0.012). Additionally, daytime napping and insomnia had an independent causal effect on the risk of MetS after separately adjusting for potential confounders such as BMI, smoking initiation, and alcohol consumption. All directions and most of the statistical significance of IVW results in multivariable MR were consistent with multivariable MR Egger results (Table 2).
Discussion
This MR study utilized two independent MetS datasets and combined multiple approaches to investigate the causal association of sleep-related traits with MetS and its components. The results from the univariable MR indicated that daytime napping and insomnia were positively associated with MetS and its components such as triglycerides, waist circumference, or hypertension. In multivariable MR analyses, the causal effect of daytime napping and insomnia on the risk of MetS remained significant even after adjusting for confounders and other sleep traits.
Most previous studies have suggested a positive association between daytime napping and MetS. A meta-analysis, including ten cross-sectional studies and a cohort study, found that daytime napping was significantly associated with increased risk of MetS in adults (pooled OR = 1.17, 95% CI 1.07, 1.28) [6]. Additionally, several observational studies also provided evidence for a potential association of daytime napping with increased waist circumference [33] and triglycerides [34]. Similarly, insomnia may be recognized as a risk factor for MetS with the available evidence. A cross-sectional study involving 830 individuals reported that insomnia increased the risk of MetS (OR = 1.97, 95% CI 1.61, 4.26) [35]. Another observational study conducted by Syauqy et.al. showed that insomnia symptoms significantly increased the odds ratios of large waist circumference and high blood pressure [36]. Consistent with previous findings, our MR study supported the causal association of daytime napping and insomnia with increased risk of MetS, these findings were validated by two independent MetS datasets and various sensitivity analyses.
Although most previous studies showed that sleep duration was associated with MetS, especially short sleep duration [8], our MR study did not observe a similar association. Several reasons may explain this situation. First, estimates from traditional observational studies are susceptible to confounders. Second, different diagnostic criteria for MetS may lead to inconsistent results. Third, only 7 SNPs were used as genetic instruments for long sleep duration in our MR study, which may explain the low phenotypic variance and restrict statistical power. Studies on chronotype were limited. Our finding suggested that neither evening nor morning chronotype had a causal effect on MetS, which aligned with a previous study utilizing a genetic risk score [37].
The exact mechanisms of daytime napping and insomnia on increased risk of MetS have not been fully determined, but some hypotheses have been proposed. Daytime napping [38] and insomnia may increase sympathetic activity and boost the renin-angiotensin system. Hyperactivation of the sympathetic nervous and renin-angiotensin system results in elevated cortisol production and systemic inflammation, contributing to glucose dysregulation, insulin resistance, and abnormal lipid distribution [39]. Moreover, frequent daytime napping may increase the risk of MetS due to obesity by reducing physical activity and lowering energy expenditure [6].
This study has several highlights. The major strength is the MR design, which minimizes the biases from confounding factors and reverse causation. Second, this study focused on individuals with European ancestry and generated estimates that were validated by two independent MetS datasets, reducing the bias due to population stratification and weak instrument bias caused by sample overlap. Moreover, multiple complementary sensitivity analyses were conducted to ensure the robustness of the results. However, our study also has limitations. First, the evaluation of five sleep traits in the original GWAS data was primarily based on self-reported questionnaires instead of objective measures. Furthermore, genetic instruments used for certain sleep traits explained a low phenotypic variance. Thus, our effect estimates may be biased due to potential exposure misclassification and low statistical power. Second, other sleep disorders such as sleep apnea might be associated with daytime napping and sleep duration, which may bias our results. Due to the limitations of the available GWAS data, we did not include it in our study. Further studies are needed to confirm and extend our findings when appropriate datasets become available. Third, sample overlap might exist in our study, causing weak instrumental bias and distortion of the results. Nevertheless, our IVs were selected at a genome-wide significant threshold and validated using a range of more stringent thresholds. Additionally, all F-statistics exceeded 10, which suggested that the bias in MR estimates was likely to be negligible. Fourth, MR assesses the causal effect of lifetime exposure, meaning that estimates are not exactly equivalent to real-world effect sizes. Finally, MR analysis relies on various assumptions which we had tried to test where possible, but residual uncertainty inevitably remains.
Conclusion
In summary, our MR study suggested that daytime napping and insomnia were potential risk factors for MetS. Moreover, daytime napping and insomnia-induced MetS primarily manifested as hypertension, increased waist circumference, or elevated triglyceride levels. However, future studies are need to confirm these association using more appropriate sleep-related traits datasets and elucidate the underlying mechanisms.
Data availability
The data underlying this article are available in the article and in its online supplementary material.
References
Eckel RH, Alberti K, Grundy SM, Zimmet PZ (2010) The metabolic syndrome. Lancet 375:181–183. https://doi.org/10.1016/S0140-6736(09)61794-3
Pammer LM, Lamina C, Schultheiss UT et al (2021) Association of the metabolic syndrome with mortality and major adverse cardiac events: a large chronic kidney disease cohort. J Intern Med 290:1219–1232. https://doi.org/10.1111/joim.13355
Saklayen MG (2018) The global epidemic of the metabolic syndrome. Curr Hypertens Rep 20:12. https://doi.org/10.1007/s11906-018-0812-z
Hirode G, Wong RJ (2020) Trends in the prevalence of metabolic syndrome in the United States, 2011–2016. JAMA 323:2526. https://doi.org/10.1001/jama.2020.4501
Jia Y, Guo D, Sun L et al (2022) Self-reported daytime napping, daytime sleepiness, and other sleep phenotypes in the development of cardiometabolic diseases: a Mendelian randomization study. Eur J Prev Cardiol 29:1982–1991. https://doi.org/10.1093/eurjpc/zwac123
Sun J, Ma C, Zhao M et al (2022) Daytime napping and cardiovascular risk factors, cardiovascular disease, and mortality: a systematic review. Sleep Med Rev 65:101682. https://doi.org/10.1016/j.smrv.2022.101682
Che T, Yan C, Tian D et al (2021) The association between sleep and metabolic syndrome: a systematic review and meta-analysis. Front Endocrinol 12:773646. https://doi.org/10.3389/fendo.2021.773646
Deng H-B, Tam T, Zee BC-Y et al (2017) Short sleep duration increases metabolic impact in healthy adults: a population-based cohort study. Sleep 40:zsx130. https://doi.org/10.1093/sleep/zsx130
Garbarino S, Magnavita N (2019) Sleep problems are a strong predictor of stress-related metabolic changes in police officers. A prospective study. PLoS ONE 14:e0224259. https://doi.org/10.1371/journal.pone.0224259
Yu JH, Yun C-H, Ahn JH et al (2015) Evening chronotype is associated with metabolic disorders and body composition in middle-aged adults. J Clin Endocrinol Metab 100:1494–1502. https://doi.org/10.1210/jc.2014-3754
Vizmanos B, Cascales AI, Rodríguez-Martín M et al (2023) Lifestyle mediators of associations among siestas, obesity, and metabolic health. Obes Silver Spring Md 31:1227–1239. https://doi.org/10.1002/oby.23765
Choi S, Kim K, Lee JK et al (2019) Association between change in alcohol consumption and metabolic syndrome: analysis from the health examinees study. Diabetes Metab J 43:615–626. https://doi.org/10.4093/dmj.2018.0128
Yao F, Bo Y, Zhao L et al (2021) Prevalence and influencing factors of metabolic syndrome among adults in China from 2015 to 2017. Nutrients 13:4475. https://doi.org/10.3390/nu13124475
Wen L, Fan J, Shi X, et al (2024) Causal association of rheumatoid arthritis with frailty and the mediation role of inflammatory cytokines: a Mendelian randomization study. Arch Gerontol Geriatr 105348. https://doi.org/10.1016/j.archger.2024.105348
Dashti HS, Daghlas I, Lane JM et al (2021) Genetic determinants of daytime napping and effects on cardiometabolic health. Nat Commun 12:900. https://doi.org/10.1038/s41467-020-20585-3
Dashti HS, Jones SE, Wood AR et al (2019) Genome-wide association study identifies genetic loci for self-reported habitual sleep duration supported by accelerometer-derived estimates. Nat Commun 10:1100. https://doi.org/10.1038/s41467-019-08917-4
Jansen PR, Watanabe K, Stringer S et al (2019) Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat Genet 51:394–403. https://doi.org/10.1038/s41588-018-0333-3
Jones SE, Lane JM, Wood AR et al (2019) Genome-wide association analyses of chronotype in 697,828 individuals provides insights into circadian rhythms. Nat Commun 10:343. https://doi.org/10.1038/s41467-018-08259-7
Lind L (2019) Genome-wide association study of the metabolic syndrome in UK Biobank. Metab Syndr Relat Disord 17:505–511. https://doi.org/10.1089/met.2019.0070
Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults (2001) Executive Summary of The Third Report of The National Cholesterol Education Program (NCEP) Expert Panel on Detection, Evaluation, And Treatment of High Blood Cholesterol In Adults (Adult Treatment Panel III). JAMA 285:2486–2497. https://doi.org/10.1001/jama.285.19.2486
Burgess S, Davies NM, Thompson SG (2016) Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol 40:597–608. https://doi.org/10.1002/gepi.21998
van Walree ES, Jansen IE, Bell NY et al (2022) Disentangling genetic risks for metabolic syndrome. Diabetes 71:2447–2457. https://doi.org/10.2337/db22-0478
Shungin D, Winkler TW, Croteau-Chonka DC et al (2015) New genetic loci link adipose and insulin biology to body fat distribution. Nature 518:187–196. https://doi.org/10.1038/nature14132
Kurki MI, Karjalainen J, Palta P et al (2023) FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613:508–518. https://doi.org/10.1038/s41586-022-05473-8
Willer CJ, Schmidt EM, Sengupta S et al (2013) Discovery and refinement of loci associated with lipid levels. Nat Genet 45:1274–1283. https://doi.org/10.1038/ng.2797
Chen J, Spracklen CN, Marenne G et al (2021) The trans-ancestral genomic architecture of glycemic traits. Nat Genet 53:840–860. https://doi.org/10.1038/s41588-021-00852-9
Liu M, Jiang Y, Wedow R et al (2019) Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet 51:237–244. https://doi.org/10.1038/s41588-018-0307-5
Yengo L, Sidorenko J, Kemper KE et al (2018) Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum Mol Genet 27:3641–3649. https://doi.org/10.1093/hmg/ddy271
Burgess S, Butterworth A, Thompson SG (2013) Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol 37:658–665. https://doi.org/10.1002/gepi.21758
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44:512–525. https://doi.org/10.1093/ije/dyv080
Bowden J, Davey Smith G, Haycock PC, Burgess S (2016) consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol 40:304–314. https://doi.org/10.1002/gepi.21965
Verbanck M, Chen C-Y, Neale B, Do R (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50:693–698. https://doi.org/10.1038/s41588-018-0099-7
Tse LA, Wang C, Rangarajan S et al (2021) Timing and length of nocturnal sleep and daytime napping and associations with obesity types in high-, middle-, and low-income countries. JAMA Netw Open 4:e2113775. https://doi.org/10.1001/jamanetworkopen.2021.13775
Cai H, Su N, Li W et al (2021) Relationship between afternoon napping and cognitive function in the ageing Chinese population. Gen Psychiatry 34:e100361. https://doi.org/10.1136/gpsych-2020-100361
Zou D, Wennman H, Hedner J et al (2021) Insomnia is associated with metabolic syndrome in a middle-aged population: the SCAPIS pilot cohort. Eur J Prev Cardiol 28:e26–e28. https://doi.org/10.1177/2047487320940862
Syauqy A, Hsu C-Y, Rau H-H et al (2019) Association of sleep duration and insomnia symptoms with components of metabolic syndrome and inflammation in middle-aged and older adults with metabolic syndrome in Taiwan. Nutrients 11:1848. https://doi.org/10.3390/nu11081848
Vera B, Dashti HS, Gómez-Abellán P, et al (2018) Modifiable lifestyle behaviors, but not a genetic risk score, associate with metabolic syndrome in evening chronotypes. Sci Rep 8. https://doi.org/10.1038/s41598-017-18268-z
Devine JK, Wolf JM (2016) Determinants of cortisol awakening responses to naps and nighttime sleep. Psychoneuroendocrinology 63:128–134. https://doi.org/10.1016/j.psyneuen.2015.09.016
Anagnostis P, Athyros VG, Tziomalos K et al (2009) The pathogenetic role of cortisol in the metabolic syndrome: a hypothesis. J Clin Endocrinol Metab 94:2692–2701. https://doi.org/10.1210/jc.2009-0370
Funding
This research was supported by the National Natural Science Foundation of China (No.82304251).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
We used publicly available GWAS summary statistics and each GWAS was approved by its corresponding ethics committee and followed the tenants of the Declaration of Helsinki.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, Y., Wen, L., Shi, X. et al. Causal effects of sleep traits on metabolic syndrome and its components: a Mendelian randomization study. Sleep Breath (2024). https://doi.org/10.1007/s11325-024-03020-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11325-024-03020-5