Abstract
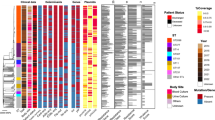
Guangdong Province, China’s largest economy, has a high incidence of tuberculosis (TB). At present, there are few reports on the distribution, transmission and drug resistance of Mycobacterium tuberculosis (Mtb) strains in this region. In this study, we performed minimum inhibitory concentration testing for 14 anti-TB drugs and whole-genome sequencing of 713 clinical Mtb isolates from 20,662 sputum culture-positive tuberculosis patients registered at 31 tuberculosis drug resistance surveillance sites covering 20 cities in Guangdong Province from 2016 to 2018. Moreover, we evaluated genome-wide associations between mutations and drug resistance, and further investigated the differences in the MICs of mutations. The epidemiology, drug-resistant phenotypes and whole genome sequencing data of 713 clinical Mtb isolates were analyzed, revealing the lineage distribution and drug-resistant gene profiles in Guangdong Province. WGS combined with quantitative MIC measurements identified several novel loci associated with resistance, of which 16 loci were found to be related to resistance to more than one drug. This study analyzed the lineage distribution, prevalence characteristics and resistance-corresponding gene profiles of Mtb isolates in Guangdong province, and provided a theoretical basis for the formulation of tuberculosis prevention and control policy in the province.






Similar content being viewed by others
Availability of Data and Materials
The raw data of WGS has already been submitted to the Sequence Read Archive (SRA) database. The BioProject accession numbers are PRJNA866200.
Abbreviations
- TB:
-
Tuberculosis
- Mtb:
-
Mycobacterium tuberculosis
- MDR/RR–TB:
-
Isoniazid and rifampicin TB or rifampicin-resistant TB
- WGS:
-
Whole-genome sequencing
- MIC:
-
Minimum inhibitory concentration
- INH:
-
Isoniazid
- RIF:
-
Rifampicin
- SM:
-
Streptomycin
- PZA:
-
Pyrazinamide
- EMB:
-
Ethambutol
- OFX:
-
Ofloxacin
- RBU:
-
Rifabutin
- MFX:
-
Moxifloxacin
- PAS:
-
Para-aminosalicylic acid
- LFX:
-
Levofloxacin
- CAP:
-
Capreomycin
- AKM:
-
Amikacin
- KM:
-
Kanamycin
- PTO:
-
Prothionamide
- ETH:
-
Ethionamide
- GWAS:
-
Genome-wide association study
- QQ:
-
Quantile-quantile
- LJ:
-
Löwenstein–Jensen
- NTM:
-
Non-tuberculosis mycobacteria
- DST:
-
Drug susceptibility testing
References
World Health Organization. Global tuberculosis report 2022. Available at: https://www.who.int/publications/i/item/9789240061729. Accessed 27 Oct 2022
Satta G, Lipman M, Smith GP, Arnold C, Kon OM, McHugh TD (2018) Mycobacterium tuberculosis and whole-genome sequencing: how close are we to unleashing its full potential? Clin Microbiol Infect 24:604–609
de Paiva J, Magalhaes M, Leal TC, Da SL, Da SL, Do CR, de Souza C (2022) Time trend, social vulnerability, and identification of risk areas for tuberculosis in Brazil: an ecological study. PLoS ONE 17:e247894
Wang L, Xu C, Hu M, Qiao J, Chen W, Li T, Qian S, Yan M (2021) Spatio-temporal variation in tuberculosis incidence and risk factors for the disease in a region of unbalanced socio-economic development. BMC Public Health 21:1–1817
Najafizada M, Rahman A, Taufique Q, Sarkar A (2021) Social determinants of multidrug-resistant tuberculosis: a scoping review and research gaps. Indian J Tuberc 68:99–105
Hota SR, Padhi SK, Pahari A, Behera BK, Panda B, Mor SK, Singh VK, Goyal SM, Sahoo N (2022) Characterization and whole genome sequencing of chromobacterium violaceum OUAT_2017: a zoonotic pathogen found fatal to a wild asiatic elephant. Indian J Microbiol 62:627–633
Liu D, Huang F, Zhang G, He W, Ou X, He P, Zhao B, Zhu B, Liu F, Li Z et al (2022) Whole-genome sequencing for surveillance of tuberculosis drug resistance and determination of resistance level in China. Clin Microbiol Infect 28:731–739
van Beek J, Haanperä M, Smit PW, Mentula S, Soini H (2019) Evaluation of whole genome sequencing and software tools for drug susceptibility testing of Mycobacterium tuberculosis. Clin Microbiol Infec 25:82–86
Chen X, He G, Wang S, Lin S, Chen J, Zhang W (2019) Evaluation of whole-genome sequence method to diagnose resistance of 13 anti-tuberculosis drugs and characterize resistance genes in clinical multi-drug resistance mycobacterium tuberculosis isolates from China. Front Microbiol 2019, 10
Schleusener V, Koser CU, Beckert P, Niemann S, Feuerriegel S (2017) Mycobacterium tuberculosis resistance prediction and lineage classification from genome sequencing: comparison of automated analysis tools. Sci Rep 7:46327
Ngo TM, Teo YY (2019) Genomic prediction of tuberculosis drug-resistance: benchmarking existing databases and prediction algorithms. BMC Bioinform 20:68
Andersson DI (2003) Persistence of antibiotic resistant bacteria. Curr Opin Microbiol 6:452–456
Chopra I, O Neill AJ, Miller K (2003) The role of mutators in the emergence of antibiotic-resistant bacteria. Drug Resist Update 6:137–145
Davies J (1994) Inactivation of antibiotics and the dissemination of resistance genes. Science (American Association for the Advancement of Science) 264:375–382
Smith T, Wolff KA, Nguyen L (2012) molecular biology of drug resistance in mycobacterium tuberculosis. In vol. 374. Springer, Berlin, 53–80
Huang H, Ding N, Yang T, Li C, Jia X, Wang G, Zhong J, Zhang J, Jiang G, Wang S et al (2019) Cross-sectional whole-genome sequencing and epidemiological study of multidrug-resistant mycobacterium tuberculosis in China. Clin Infect Dis 69:405–413
Jiang Q, Liu Q, Ji L, Li J, Zeng Y, Meng L, Luo G, Yang C, Takiff HE, Yang Z et al (2020) Citywide transmission of multidrug-resistant tuberculosis under China’s rapid urbanization: a retrospective population-based genomic spatial epidemiological study. Clin Infect Dis 71:142–151
Coll F, McNerney R, Preston MD, Guerra-Assuncao JA, Warry A, Hill-Cawthorne G, Mallard K, Nair M, Miranda A, Alves A et al (2015) Rapid determination of anti-tuberculosis drug resistance from whole-genome sequences. Genome Med 7:51
Wei W, Zhao Y, Zhang C, Yu M, Wu Z, Xu L, Peng K, Wu Z, Li Y, Wang X (2023) Whole-genome sequencing and transcriptome-characterized in vitro evolution of aminoglycoside resistance in Mycobacterium tuberculosis. Microb Genom 9(5)
Lees JA, Galardini M, Bentley SD, Weiser JN, Corander J (2018) Pyseer: a comprehensive tool for microbial pangenome-wide association studies. Bioinformatics 34:4310–4312
Liu Z, Dong H, Wu B, Zhang M, Zhu Y, Pang Y, Wang X (2019) Is rifampin resistance a reliable predictive marker of multidrug-resistant tuberculosis in China: a meta-analysis of findings. J Infection 79:349–356
Riccardi N, Saderi L, Borroni E, Tagliani E, Cirillo DM, Marchese V, Matteelli A, Piana A, Castellotti P, Ferrarese M et al (2021) Therapeutic strategies and outcomes of MDR and pre-XDR-TB in Italy: a nationwide study. Int J Tuberc Lung Dis 25:395–399
Kurbatova EV, Cavanaugh JS, Dalton T, Click ES, Cegielski JP (2013) Epidemiology of pyrazinamide-resistant tuberculosis in the United States, 1999–2009. Clin Infect Dis 57:1081–1093
Xu P, Wu J, Yang C, Luo T, Shen X, Zhang Y, Nsofor CA, Zhu G, Gicquel B, Gao Q (2016) Prevalence and transmission of pyrazinamide resistant Mycobacterium tuberculosis in China. Tuberculosis (Edinb) 98:56–61
Walker TM, Miotto P, Koser CU, Fowler PW, Knaggs J, Iqbal Z, Hunt M, Chindelevitch L, Farhat M, Cirillo DM et al (2022) The 2021 WHO catalogue of Mycobacterium tuberculosis complex mutations associated with drug resistance: a genotypic analysis. Lancet Microbe 3:e265–e273
Li J, Yang T, Hong C, Yang Z, Wu L, Gao Q, Yang H, Tan W, Carvalho-Assef APDA (2022) Whole-genome sequencing for resistance level prediction in multidrug-resistant tuberculosis. Microbiol Spectrum 10:e271421
Crook DW, Rodrigues C, Ismail NA, Mistry N, Iqbal Z, Merker M, Moore D, Walker AS, Thwaites G, Niemann S et al (2022) Genome-wide association studies of global Mycobacterium tuberculosis resistance to 13 antimicrobials in 10,228 genomes identify new resistance mechanisms. PLOS Biol 20:e3001755
Mustafa AS (2022) Early secreted antigenic target of 6 kda-like proteins of mycobacterium tuberculosis: diagnostic and vaccine relevance. Int J Mycobacteriol 11:10–15
Li S, Poulton NC, Chang JS, Azadian ZA, DeJesus MA, Ruecker N, Zimmerman MD, Eckartt KA, Bosch B, Engelhart CA et al (2022) CRISPRi chemical genetics and comparative genomics identify genes mediating drug potency in Mycobacterium tuberculosis. Nat Microbiol 7:766–779
Meneguello JE, Arita GS, Silva JVDO, Ghiraldi-Lopes LD, Caleffi-Ferracioli KR, Siqueira VLD, Scodro RBDL, Pilau EJ, Campanerut-Sá PAZ, Cardoso RF (2020) Insight about cell wall remodulation triggered by rifampicin in Mycobacterium tuberculosis. Tuberculosis (Edinb) 120:101903
Acknowledgements
The authors thank the partners of TB drug resistance monitoring stations in Guangdong province for their hard work in sorting out patient information and collecting Mtb strains. The authors thank Tao Chen, Zhenyan Li and Li Yu, formerly working in Center for Tuberculosis Control of Guangdong Province, for their hard work in the establishment of biobank. The authors express appreciation of Zhichao Chen from Shanghai Gene-Optimal Science &Technology Co., Ltd for help with WGS analysis.
Funding
This work was supported by a Major Infectious Disease Prevention and Control of the National Science and Technique Major Project [2018ZX10715004], the Science and Technology Planning Project of Guangzhou [202201010785] and Medical Science Foundation of Guangdong Province [B2021012].
Author information
Authors and Affiliations
Contributions
Wenjing Wei, Yuhui Chen and Meiling Yu conceived the study. Wenjing Wei and Yuhui Chen acquired funding, supervised and administered the project. Chenchen Zhang, Zhuhua Wu, Xinchun Huang, Yuchuan Zhao, Meiling Yu and Qi Sun processed the samples and performed the experiments. Chenchen Zhang, Meiling Yu, Yanmei Chen, Qi Sun, Huixin Guo and Wenya Dong were responsible for collecting clinical strains and conducting drug susceptibility tests. Chenchen Zhang, Yuchuan Zhao and Wenjing Wei analyzed the data. Qinghua Liao, Huizhong Wu and Xunxun Chen were responsible for providing guidance on clinical TB drug use. Anqi Liang and Wenya Dong were responsible for clinical strain information collection and collation. Chenchen Zhang, Wenjing Wei and Yuhui Chen prepared the manuscript. All the authors have read the final version of the manuscript and have approved it.
Corresponding authors
Ethics declarations
Coonflict of interests
The authors declare no competing interests.
Ethics Approval and Consent to Participate
The clinical Mtb isolates used in this study were collected from the Major Infectious Disease Prevention and Control of the National Science and Technique Major Project and preserved by the Biobank of Center for Tuberculosis Control of Guangdong Province. Written informed consent was also obtained from each patient in face-to-face interviews by trained clinicians. The study was performed in accordance with the Declaration of Helsinki and approved by the Medical Ethics Committee of Center for Tuberculosis Control of Guangdong Province, China.
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, C., Wu, Z., Huang, X. et al. A Profile of Drug-Resistant Mutations in Mycobacterium tuberculosis Isolates from Guangdong Province, China. Indian J Microbiol (2024). https://doi.org/10.1007/s12088-024-01236-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12088-024-01236-3