Abstract
Background
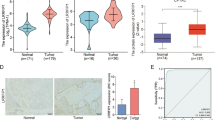
Esophageal cancer (ESCA) is a common malignant tumor of the digestive tract, and its poor prognosis is mainly attributed to the occurrence of invasion and metastasis. Z-DNA binding protein 1 (ZBP1), as a mRNA regulatory factor, plays an important role in the occurrence and development of various tumors. However, the role of ZBP1 in ESCA is not yet understood.
Aims
This study aims to explore the expression of ZBP1 in ESCA and its role in the development of ESCA.
Methods
Using bioinformatics analysis and immunohistochemistry staining, we detected the expression of ZBP1 in ESCA and normal tissues. The potential mechanism of ZBP1 in ESCA was analyzed from the aspects of genetic mutations, protein interaction networks, and pathway enrichment. We performed functional experiments in vitro to elucidate the effect of ZBP1 on ESCA cells.
Results
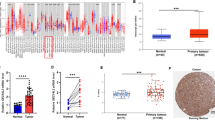
ZBP1 was found to be significantly upregulated in ESCA compared to adjacent noncancerous tissues, and its expression is closely related to gender, age, and lymph node metastasis. In ESCA, the genetic variation rate of ZBP1 is 8%, and its expression is positively correlated with immune cell infiltration. The ZBP1 co-expressed gene is mainly involved in processes such as lymph node proliferation and intercellular adhesion. In vitro experiments have confirmed that downregulation of ZBP1 significantly inhibited the proliferation, migration, and invasion of ESCA cells.
Conclusion
This research proves that downregulation of ZBP1 can inhibit the progression of ESCA. This finding indicates that ZBP1 may be a novel biomarker to improve the diagnosis and treatment of ESCA.
Graphical Abstract










Similar content being viewed by others
Data Availability
Public datasets were obtained from UALCAN (https://ualcan.path.uab.edu/), TIMER2.0 (http://timer.cistrome.org/), GEPIA2 (http://gepia2.cancer-pku.cn/#index), cBioportal (http://www.cbioportal.org/), STRING (https://string-db.org/) and LinkedOmics (http://www.Linkedomics.org).
Consent for publication
All authors agreed to publish the article.
Abbreviations
- ZBP1:
-
Z-DNA binding protein 1
- IAV:
-
Influenza A virus
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ECSA:
-
Esophageal cancer
- DAI:
-
DNA-dependent activator of IFN regulatory factors
- TCGA:
-
The Cancer Genome Atlas
- ANOVA:
-
One-way analysis of variance
- BLCA:
-
Bladder cancer
- BRCA:
-
Breast invasive Carcinoma
- CESC:
-
Cervical Squamous Cell Carcinoma
- CHOL:
-
Cholangio Carcinoma
- GBM:
-
Glioblastoma multiform
- HNSC:
-
Head and neck squamous cell carcinoma
- KIRC:
-
Kidney renal clear cell Carcinoma
- KIRP:
-
Kidney Renal Papillary Cell Carcinoma
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- PRAD:
-
Prostate adenocarcinoma
- SARC:
-
Sarcoma
- UCEC:
-
Uterine corpus endometrial carcinoma
- READ:
-
Rectum adenocarcinoma
- THCA:
-
Thyroid cancer
- COAD:
-
Colon adenocarcinoma
- IFN-I:
-
Type I interferon
- MLKL:
-
Mixed linear kinase domain-like protein
- OS:
-
Overall survival
- DFS:
-
Disease-free survival
- BP:
-
Biological processes
- CC:
-
Cellular components
- MF:
-
Molecular functions
References
Liu CQ, Ma YL, Qin Q, et al. Epidemiology of esophageal cancer in 2020 and projections to 2030 and 2040. Thoracic Cancer 2023; 14: 3–11.
Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021; 71: 209–249.
Yu C, Xi Y, Zhang P, et al. Dissecting the molecular profiling and tumor immune microenvironment of three subtypes of esophageal cancer. J Gene Med 2023; 25: e3482.
Chen Z, Wang Q, Huang L, et al. LncRNA PVT1 confers cisplatin resistance of esophageal cancer cells through modulating the miR-181a-5p-glutaminase (GLS) axis. Nut Cancer 2023; 75: 1646–1657.
Zhang L, Zong L, Li W, et al. Construction of lncRNA prognostic model related to cuproptosis in esophageal carcinoma. Front Genet 2023; 14: 1120827.
Gu W, Pan F, Singer RH. Blocking beta-catenin binding to the ZBP1 promoter represses ZBP1 expression, leading to increased proliferation and migration of metastatic breast-cancer cells. J Cell Sci 2009; 122: 1895–1905.
Hao Y, Yang B, Yang J, et al. ZBP1: a powerful innate immune sensor and double-edged sword in host immunity. Int J Mol Sci 2022; 23: 10224.
Baik JY, Liu Z, Jiao D, et al. ZBP1 not RIPK1 mediates tumor necroptosis in breast cancer. Nat Commun 2021; 12: 2666.
Liu Y, Cao H, Zhao Y, et al. Fisetin-induced cell death in human ovarian cancer cell lines via zbp1-mediated necroptosis. J Ovarian Res 2022; 15: 57.
Cheng R, Liu X, Wang Z, et al. ABT-737, a Bcl-2 family inhibitor, has a synergistic effect with apoptosis by inducing urothelial carcinoma cell necroptosis. Mol Med Rep 2021; 23: 412.
Ponnusamy K, Tzioni MM, Begum M, et al. The innate sensor ZBP1-IRF3 axis regulates cell proliferation in multiple myeloma. Haematologica 2022; 107: 721–732.
Chen D, Ermine K, Wang YJ, et al. PUMA/RIP3 mediates chemotherapy response via necroptosis and local immune activation in colorectal cancer. Mol Cancer Ther 2023; 23: 354–367.
Li J, Tang M, Ke RX, et al. The anti-cancer drug candidate CBL0137 induced necroptosis via forming left-handed Z-DNA and its binding protein ZBP1 in liver cells. Toxicol Appl Pharmacol 2024; 482: 116765.
Chandrashekar DS, Karthikeyan SK, Korla PK, et al. UALCAN: an update to the integrated cancer data analysis platform. Neoplasia (New York NY) 2022; 25: 18–27.
Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res 2017; 77: e108–e110.
Li B, Severson E, Pignon JC, et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol 2016; 17: 174.
Tang Z, Kang B, Li C, et al. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res 2019; 47: W556–W560.
Renner C, Reimer N, Christoph J, et al. Extending cBioPortal for therapy recommendation documentation in molecular tumor boards: development and usability study. JMIR Med Inform 2023; 11: e50017.
Szklarczyk D, Gable AL, Nastou KC, et al. The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 2021; 49: D605–D612.
Vasaikar SV, Straub P, Wang J, et al. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res 2018; 46: D956–D963.
He H, Zhang P, Li F, et al. Predicting the prognosis of esophageal cancer based on extensive analysis of new inflammatory response-related signature. J Biochem Mol Toxicol 2023; 37: e23291.
Xiong Y. The emerging role of PANoptosis in cancer treatment. Biomed Pharmacother 2023; 168: 115696.
Huang QF, Fang DL, Nong BB, et al. Focal pyroptosis-related genes AIM2 and ZBP1 are prognostic markers for triple-negative breast cancer with brain metastases. Transl Cancer Res 2021; 10: 4845–4858.
Zeng R, Huang S, Qiu X, et al. Predicting the prognosis of esophageal adenocarcinoma by a pyroptosis-related gene signature. Front Pharmacol 2021; 12: 767187.
Yang Y, Wu M, Cao D, et al. ZBP1-MLKL necroptotic signaling potentiates radiation-induced antitumor immunity via intratumoral STING pathway activation. Sci Adv 2021; 7: eabf6290.
Mirzaei S, Saghari S, Bassiri F, et al. NF-κB as a regulator of cancer metastasis and therapy response: a focus on epithelial–mesenchymal transition. J Cell Physiol 2022; 237: 2770–2795.
Ye J, Deng W, Zhong Y, et al. MELK predicts poor prognosis and promotes metastasis in esophageal squamous cell carcinoma via activating the NF-κB pathway. Int J Oncol 2022; 61: 94.
Yang X, Wang J, Liu W. Molecular markers of type II alveolar epithelial cells in acute lung injury by bioinformatics analysis. Sci Rep 2023; 13: 17797.
Zhang Z, Wang D, Wang P, et al. OTUD1 negatively regulates Type I IFN induction by disrupting noncanonical ubiquitination of IRF3. J Immunol 2020; 204: 1904–1918.
Tian M, Wang X, Sun J, et al. IRF3 prevents colorectal tumorigenesis via inhibiting the nuclear translocation of β-catenin. Nat Commun 2020; 11: 5762.
Wei Z, Wang Y, Ma W, et al. Serine-arginine splicing factor 2 promotes oesophageal cancer progression by regulating alternative splicing of interferon regulatory factor 3. RNA Biol 2023; 20: 359–367.
Zhang CF, Cao BW, Lu ZM, et al. Relationship between polymorphism of IRF-3 gene codon 427 and esophageal cancer in Anyang population of China. Beijing Da Xue Xue Bao Yi Xue Ban 2004; 36: 345–347.
Li Y, Hu X, Song Y, et al. Identification of novel alternative splicing variants of interferon regulatory factor 3. Biochim Biophys Acta 2011; 1809: 166–175.
Zhuang Y, Ortega-Ribera M, Thevkar Nagesh P, et al. Bile acid-induced IRF3 phosphorylation mediates cell death, inflammatory responses and fibrosis in cholestasis-induced liver and kidney injury via regulation of ZBP1. Hepatology (Baltim Md). 2023. https://doi.org/10.1097/HEP.0000000000000611.
Takaoka A, Wang Z, Choi MK, et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 2007; 448: 501–505.
Oh S, Kim E, Lee H. Comparative impact of PD-1 and PD-L1 inhibitors on advanced esophageal or gastric/gastroesophageal junction cancer treatment: a systematic review and meta-analysis. J Clin Med 2021; 10: 3612.
Ricciuti B, Wang X, Alessi JV, et al. Association of high tumor mutation burden in non-small cell lung cancers with increased immune infiltration and improved clinical outcomes of PD-L1 blockade across PD-L1 expression levels. JAMA Oncol 2022; 8: 1160–1168.
Gohil SH, Iorgulescu JB, Braun DA, et al. Applying high-dimensional single-cell technologies to the analysis of cancer immunotherapy. Nat Rev Clin Oncol 2021; 18: 244–256.
Walcher L, Kistenmacher AK, Suo H, et al. Cancer stem cells—origins and biomarkers: perspectives for targeted personalized therapies. Front Immunol 2020; 11: 1280.
Pan C, Wang Y, Liu Q, et al. Phenotypic profiling and prognostic significance of immune infiltrates in esophageal squamous cell carcinoma. Oncoimmunology 2021; 10: 1883890.
Acknowledgments
We acknowledge TCGA databases for providing their platforms and data, and we thank the contributors who uploaded their datasets for public access and use.
Funding
This work was supported by the Special Research Project for Health and Young Medical Science and Technology Talents in Xinjiang Uygur Autonomous Region (WJWY-202345).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study's conception and design. Shabahaiti Wusiman, Yining Liu, Hui Li and Ling Liu conceived and supervised the study; Shabahaiti Wusiman, Hui Li and Ling Liu designed experiments; Shabahaiti Wusiman, Yining Liu, Ximing Qu, Hainisayimu Tuerxun and Yuhan Deng performed experiments; Shabahaiti Wusiman, Ling Liu and Yining Liu analyzed data; Ling Liu, Yining Liu, and Shabahaiti Wusiman wrote the manuscript; Yining Liu, Hui Li, Ximing Qu, Hainisayimu Tuerxun and Yuhan Deng made manuscript revisions. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The study was approved by the Ethics Committee of Shanghai Xinchao Biotechnology Co., Ltd. (ethics approval No. YB M-05-02).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wusiman, S., Liu, Y., Li, H. et al. Highly Expressed Z-DNA Binding Protein 1 in Esophageal Cancer Promotes Tumor Growth. Dig Dis Sci (2024). https://doi.org/10.1007/s10620-024-08375-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10620-024-08375-z