Abstract
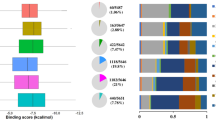
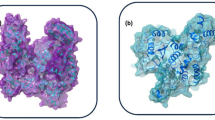
The emergence of zoonotic monkeypox (MPX) disease, caused by the double-stranded DNA monkeypox virus (MPXV), has become a global threat. Due to unavailability of a specific small molecule drug for MPX, this study investigated Moringa oleifera phytochemicals to find potent and safe inhibitors of DNA Polymerase (DNA Pol), a poxvirus drug target due to its role in the viral life cycle. For that, 146 phytochemicals were screened through drug-likeness and molecular docking analyses. Among these, 136 compounds exhibited drug-like properties, with Gossypetin showing the highest binding affinity (− 7.8 kcal/mol), followed by Riboflavin (− 7.6 kcal/mol) and Ellagic acid (− 7.6 kcal/mol). In comparison, the control drugs Cidofovir and Brincidofovir displayed lower binding affinities, with binding energies of − 6.0 kcal/mol and − 5.1 kcal/mol, respectively. Hydrogen bonds, electrostatic and hydrophobic interactions were the main non-bond interactions between inhibitors and protein active site. The identified compounds were further evaluated using molecular dynamics simulation, density functional theory analysis and ADMET analysis. Molecular dynamics simulations conducted over 200 ns revealed that DNA Pol-Gossypetin complex was not stable, however, Riboflavin and Ellagic acid complexes showed excellent stability indicating them as better DNA Pol inhibitors. The density functional theory analysis exhibited the chemical reactivity of these inhibitor compounds. The ADMET analysis suggested that the top phytochemicals were safe and showed no toxicity. In conclusion, this study has identified Riboflavin and Ellagic acid as potential DNA Pol inhibitors to control MPXV. Further experimental assays and clinical trials are needed to confirm their activity against the disease.
Graphical Abstract












Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this article.
References
Alakunle E, Moens U, Nchinda G, Okeke MI (2020) Monkeypox virus in nigeria: Infection biology, epidemiology, and evolution. Viruses. https://doi.org/10.3390/v12111257
Bunge EM, Hoet B, Chen L et al (2022) The changing epidemiology of human monkeypox—a potential threat? A systematic review. PLoS Negl Trop Dis. https://doi.org/10.1371/journal.pntd.0010141
Kugelman JR, Johnston SC, Mulembakani PM et al (2014) Genomic variability of monkeypox virus among humans, Democratic Republic of the Congo. Emerg Infect Dis 20:232–239. https://doi.org/10.3201/eid2002.130118
von Magnus P, Andersen EK, Petersen KB, Birch-Andersen A (1959) A Pox-like disease in cynomolgus monkeys. Acta Pathol Microbiol Scand 46:156–176. https://doi.org/10.1111/j.1699-0463.1959.tb00328.x
Brown K, Leggat PA (2016) Human monkeypox: current state of knowledge and implications for the future. Trop Med Infect Dis 1:8. https://doi.org/10.3390/tropicalmed1010008
Ladnyj ID, Ziegler P, Kima E (1972) A human infection caused by monkeypox virus in Basankusu territory, democratic republic of the Congo. Bull World Health Organ 46:593
Yinka-Ogunleye A, Aruna O, Dalhat M et al (2019) Outbreak of human monkeypox in Nigeria in 2017–18: a clinical and epidemiological report. Lancet Infect Dis 19:872–879. https://doi.org/10.1016/S1473-3099(19)30294-4
Isidro J, Borges V, Pinto M et al (2022) Phylogenomic characterization and signs of microevolution in the 2022 multi-country outbreak of monkeypox virus. Nat Med 2022:1–1. https://doi.org/10.1038/s41591-022-01907-y
Likos AM, Sammons SA, Olson VA et al (2005) A tale of two clades: monkeypox viruses. J Gen Virol 86:2661–2672
WHO (2022) Multi-country monkeypox outbreak in non-endemic countries. https://www.who.int/emergencies/disease-outbreak-news/item/2022-DON385. Accessed 9 Jul 2022
Alakunle EF, Okeke MI (2022) Monkeypox virus: a neglected zoonotic pathogen spreads globally. Nat Rev Microbiol 20:507–508
Chaix E, Boni M, Guillier L et al (2022) Risk of Monkeypox virus (MPXV) transmission through the handling and consumption of food. Microb Risk Anal. https://doi.org/10.1016/j.mran.2022.100237
Tiecco G, Degli Antoni M, Storti S et al (2022) Monkeypox, a literature review: what is new and where does this concerning virus come from? Viruses. https://doi.org/10.3390/v14091894
Petersen E, Kantele A, Koopmans M et al (2019) Human monkeypox: epidemiologic and clinical characteristics, diagnosis, and prevention. Infect Dis Clin North Am 33:1027–1043. https://doi.org/10.1016/j.idc.2019.03.001
Kumar N, Acharya A, Gendelman HE, Byrareddy SN (2022) The 2022 outbreak and the pathobiology of the monkeypox virus. J Autoimmun. https://doi.org/10.1016/J.JAUT.2022.102855
Mukherjee AG, Wanjari UR, Kannampuzha S et al (2023) The pathophysiological and immunological background of the monkeypox virus infection: an update. J Med Virol. https://doi.org/10.1002/jmv.28206
Okyay RA, Bayrak E, Kaya E et al (2022) Another epidemic in the shadow of Covid 19 pandemic: a review of monkeypox. Eurasian J Med Oncol 6:95–99. https://doi.org/10.14744/ejmo.2022.2022
Adler H, Gould S, Hine P et al (2022) Clinical features and management of human monkeypox: a retrospective observational study in the UK. Lancet Infect Dis. https://doi.org/10.1016/S1473-3099(22)00228-6
Farahat RA, Abdelaal A, Shah J et al (2022) Monkeypox outbreaks during COVID-19 pandemic: are we looking at an independent phenomenon or an overlapping pandemic? Ann Clin Microbiol Antimicrob 21:1–3. https://doi.org/10.1186/S12941-022-00518-2
Gessain A, Nakoune E, Yazdanpanah Y (2022) Monkeypox. N Engl J Med 387:1783–1793. https://doi.org/10.1056/NEJMra2208860
Berdis AJ (2008) DNA polymerases as therapeutic targets. Biochemistry 47:8253–8260. https://doi.org/10.1021/bi801179f
Luczkowiak J, Álvarez M, Sebastián-Martín A, Menéndez-Arias L (2018) DNA-dependent DNA polymerases as drug targets in herpesviruses and poxviruses. In: Gupta SP (ed) Viral polymerases: structures, functions and roles as antiviral drug targets. Elsevier, Amsterdam, pp 95–134
Moss B (2013) Poxvirus DNA replication. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a010199
Andrei G, De Clercq E, Snoeck R (2009) Viral DNA polymerase inhibitors. Viral genome replication. Springer, Berlin, pp 481–526
Tsai C-H, Lee P-Y, Stollar V, Li M-L (2006) Antiviral therapy targeting viral polymerase. Curr Pharm Des 12:1339–1355. https://doi.org/10.2174/138161206776361156
Antoine TE, Park PJ, Shukla D (2013) Glycoprotein targeted therapeutics: a new era of anti-herpes simplex virus-1 therapeutics. Rev Med Virol 23:194–208. https://doi.org/10.1002/rmv.1740
Cui W, Aouidate A, Wang S et al (2020) Discovering anti-cancer drugs via computational methods. Front Pharmacol. https://doi.org/10.3389/fphar.2020.00733
Li K, Du Y, Li L, Wei D-Q (2019) Bioinformatics approaches for anti-cancer drug discovery. Curr Drug Targets 21:3–17. https://doi.org/10.2174/1389450120666190923162203
Biswas D, Nandy S, Mukherjee A et al (2020) Moringa oleifera Lam. and derived phytochemicals as promising antiviral agents: a review. South African J Bot 129:272–282. https://doi.org/10.1016/J.SAJB.2019.07.049
Peng Q, Xie Y, Kuai L et al (2023) Structure of monkeypox virus DNA polymerase holoenzyme. Science 379:100–105. https://doi.org/10.1126/SCIENCE.ADE6360
Burley SK, Berman HM, Christie C et al (2018) RCSB Protein Data Bank: Sustaining a living digital data resource that enables breakthroughs in scientific research and biomedical education. Protein Sci 27:316–330. https://doi.org/10.1002/pro.3331
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Morris GM, Huey R, Olson AJ (2008) Using AutoDock for ligand-receptor docking. Curr Protoc Bioinforma. https://doi.org/10.1002/0471250953.bi0814s24
Mohanraj K, Karthikeyan BS, Vivek-Ananth RP et al (2018) IMPPAT: a curated database of Indian medicinal plants, phytochemistry and therapeutics. Sci Rep. https://doi.org/10.1038/s41598-018-22631-z
Kim S, Chen J, Cheng T et al (2019) PubChem 2019 update: improved access to chemical data. Nucleic Acids Res 47:D1102–D1109. https://doi.org/10.1093/nar/gky1033
O’Boyle NM, Banck M, James CA et al (2011) Open babel: an open chemical toolbox. J Cheminform. https://doi.org/10.1186/1758-2946-3-33
Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. Methods Mol Biol 1263:243–250. https://doi.org/10.1007/978-1-4939-2269-7_19
Jász Á, Rák Á, Ladjánszki I, Cserey G (2019) Optimized GPU implementation of Merck molecular force field and universal force field. J Mol Struct 1188:227–233. https://doi.org/10.1016/j.molstruc.2019.04.007
Rappé AK, Casewit CJ, Colwell KS et al (1992) UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J Am Chem Soc 114:10024–10035. https://doi.org/10.1021/ja00051a040
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep. https://doi.org/10.1038/srep42717
Lipinski CA (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341. https://doi.org/10.1016/j.ddtec.2004.11.007
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2012) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 64:4–17. https://doi.org/10.1016/j.addr.2012.09.019
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25. https://doi.org/10.1016/S0169-409X(96)00423-1
Trott O, Olson AJ (2010) AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461. https://doi.org/10.1002/JCC.21334
Reynolds CH, Bembenek SD, Tounge BA (2007) The role of molecular size in ligand efficiency. Bioorganic Med Chem Lett 17:4258–4261. https://doi.org/10.1016/j.bmcl.2007.05.038
Hopkins AL, Keserü GM, Leeson PD et al (2014) The role of ligand efficiency metrics in drug discovery. Nat Rev Drug Discov 13:105–121. https://doi.org/10.1038/nrd4163
Hopkins AL, Groom CR, Alex A (2004) Ligand efficiency: a useful metric for lead selection. Drug Discov Today 9:430–431. https://doi.org/10.1016/S1359-6446(04)03069-7
Edwards MP, Price DA (2010) Role of physicochemical properties and ligand lipophilicity efficiency in addressing drug safety risks. Elsevier, Amsterdam
Murray CW, Erlanson DA, Hopkins AL et al (2014) Validity of ligand efficiency metrics. ACS Med Chem Lett 5:616–618. https://doi.org/10.1021/ml500146d
BIOVIA DS (2021) BIOVIA Discovery Studio
Abraham MJ, Murtola T, Schulz R et al (2015) Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/J.SOFTX.2015.06.001
Vanommeslaeghe K, Hatcher E, Acharya C et al (2009) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem NA-NA. https://doi.org/10.1002/jcc.21367
Lindahl E, Hess B, van der Spoel D (2001) GROMACS 3.0: a package for molecular simulation and trajectory analysis. J Mol Model 7:306–317. https://doi.org/10.1007/s008940100045
Kavitha R, Karunagaran S, Chandrabose SS et al (2015) Pharmacophore modeling, virtual screening, molecular docking studies and density functional theory approaches to identify novel ketohexokinase (KHK) inhibitors. Biosystems 138:39–52. https://doi.org/10.1016/J.BIOSYSTEMS.2015.10.005
BIOVIA DS (2020) BIOVIA Materials Studio
Tsuneda T, Song JW, Suzuki S, Hirao K (2010) On Koopmans’ theorem in density functional theory. J Chem Phys 133:174101. https://doi.org/10.1063/1.3491272
Curreli F, Do KY, Belov DS et al (2017) Synthesis, antiviral potency, in vitro ADMET, and X-ray structure of potent CD4 mimics as entry inhibitors that target the Phe43 cavity of HIV-1 gp120. J Med Chem 60:3124–3153. https://doi.org/10.1021/ACS.JMEDCHEM.7B00179
Pires DEV, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58:4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Banerjee P, Eckert AO, Schrey AK, Preissner R (2018) ProTox-II: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res 46:W257–W263. https://doi.org/10.1093/NAR/GKY318
Mathialagan S, Piotrowski MA, Tess DA et al (2017) Quantitative prediction of human renal clearance and drug-drug interactions of organic anion transporter substrates using in vitro transport data: a relative activity factor approach. Drug Metab Dispos 45:409–417. https://doi.org/10.1124/DMD.116.074294/-/DC1
Velavan TP, Meyer CG, Thirumalaisamy Velavan CP (2022) Monkeypox 2022 outbreak: an update. Trop Med Int Heal 27:604–605. https://doi.org/10.1111/tmi.13785
Ren S-Y, Li J, Gao R-D (2022) 2022 Monkeypox outbreak: Why is it a public health emergency of international concern? What can we do to control it? World J Clin cases 10:10873–10881. https://doi.org/10.12998/wjcc.v10.i30.10873
Rizk JG, Lippi G, Henry BM et al (2022) Prevention and treatment of monkeypox. Drugs 82:957–963. https://doi.org/10.1007/S40265-022-01742-Y/TABLES/3
Jones EV, Moss B (1984) Mapping of the vaccinia virus DNA polymerase gene by marker rescue and cell-free translation of selected RNA. J Virol 49:72–77. https://doi.org/10.1128/jvi.49.1.72-77.1984
Prichard MN, Kern ER (2012) Orthopoxvirus targets for the development of new antiviral agents. Antiviral Res 94:111–125. https://doi.org/10.1016/j.antiviral.2012.02.012
Traktman P, Sridhar P, Condit RC, Roberts BE (1984) Transcriptional mapping of the DNA polymerase gene of vaccinia virus. J Virol 49:125–131. https://doi.org/10.1128/jvi.49.1.125-131.1984
Ollis DL, Brick P, Hamlin R et al (1985) Structure of large fragment of Escherichia coli DNA polymerase I complexed with dTMP. Nature 313:762–766. https://doi.org/10.1038/313762a0
Patel PH, Loeb LA (2001) Getting a grip on how DNA polymerases function. Nat Struct Biol 8:656–659. https://doi.org/10.1038/90344
Steitz TA (1999) DNA polymerases: structural diversity and common mechanisms. J Biol Chem 274:17395–17398. https://doi.org/10.1074/jbc.274.25.17395
Varma AK, Patil R, Das S et al (2010) Optimized hydrophobic interactions and hydrogen bonding at the target-ligand interface leads the pathways of Drug-Designing. PLoS One. https://doi.org/10.1371/journal.pone.0012029
Alex A, Millan DS, Perez M et al (2011) Intramolecular hydrogen bonding to improve membrane permeability and absorption in beyond rule of five chemical space. Medchemcomm 2:669–674. https://doi.org/10.1039/c1md00093d
Ferreira De Freitas R, Schapira M (2017) A systematic analysis of atomic protein-ligand interactions in the PDB. Medchemcomm 8:1970–1981. https://doi.org/10.1039/c7md00381a
Smith RD, Engdahl AL, Dunbar JB, Carlson HA (2012) Biophysical limits of protein-ligand binding. J Chem Inf Model 52:2098–2106. https://doi.org/10.1021/ci200612f
Nag A, Chowdhury RR (2020) Piperine, an alkaloid of black pepper seeds can effectively inhibit the antiviral enzymes of Dengue and Ebola viruses, an in silico molecular docking study. VirusDisease 31:308–315. https://doi.org/10.1007/s13337-020-00619-6
Raj U, Varadwaj PK (2016) Flavonoids as multi-target inhibitors for proteins associated with ebola virus. In silico discovery using virtual screening and molecular docking studies. Interdiscip Sci Comput Life Sci 8:132–141. https://doi.org/10.1007/s12539-015-0109-8
Eberle RJ, Olivier DS, Amaral MS et al (2022) Riboflavin, a Potent neuroprotective vitamin: focus on flavivirus and alphavirus proteases. Microorganisms. https://doi.org/10.3390/microorganisms10071331
Farah N, Chin VK, Chong PP et al (2022) Riboflavin as a promising antimicrobial agent? A multi-perspective review. Curr Res Microb Sci. https://doi.org/10.1016/j.crmicr.2022.100111
Morosetti G, Criscuolo AA, Santi F, Perno CF, Piccione E, Ciotti M (2017) Ellagic acid and Annona muricata in the chemoprevention of HPV-related pre-neoplastic lesions of the cervix. Oncol Lett 13(3):1880–1884
Alexova R, Alexandrova S, Dragomanova S et al (2023) Anti-COVID-19 potential of ellagic acid and polyphenols of Punica granatum L. Molecules 28:3772. https://doi.org/10.3390/MOLECULES28093772
Cui Q, Du R, Anantpadma M et al (2018) Identification of ellagic acid from plant rhodiola rosea l. as an anti-ebola virus entry inhibitor. Viruses. https://doi.org/10.3390/v10040152
Acquadro S, Civra A, Cagliero C et al (2020) Punica granatum Leaf ethanolic extract and ellagic acid as inhibitors of Zika virus infection. Planta Med 86:1363–1374. https://doi.org/10.1055/a-1232-5705
Sargsyan K, Grauffel C, Lim C (2017) How molecular size impacts RMSD applications in molecular dynamics simulations. J Chem Theory Comput 13:1518–1524. https://doi.org/10.1021/acs.jctc.7b00028
Kuzmanic A, Zagrovic B (2010) Determination of ensemble-average pairwise root mean-square deviation from experimental B-factors. Biophys J 98:861–871. https://doi.org/10.1016/j.bpj.2009.11.011
Bornot A, Etchebest C, De Brevern AG (2011) Predicting protein flexibility through the prediction of local structures. Proteins Struct Funct Bioinforma 79:839–852. https://doi.org/10.1002/PROT.22922
Ghasemi F, Zomorodipour A, Karkhane AA, Khorramizadeh MR (2016) In silico designing of hyper-glycosylated analogs for the human coagulation factor IX. J Mol Graph Model 68:39–47. https://doi.org/10.1016/J.JMGM.2016.05.011
Shahbaaz M, Nkaule A, Christoffels A (2019) Designing novel possible kinase inhibitor derivatives as therapeutics against Mycobacterium tuberculosis: an in silico study. Sci Rep. https://doi.org/10.1038/s41598-019-40621-7
Durham E, Dorr B, Woetzel N et al (2009) Solvent accessible surface area approximations for rapid and accurate protein structure prediction. J Mol Model 15:1093–1108. https://doi.org/10.1007/s00894-009-0454-9
Pearson RG (2005) Chemical hardness and density functional theory. J Chem Sci 117:369–377. https://doi.org/10.1007/BF02708340
Khan SA, Rizwan K, Shahid S et al (2020) Synthesis, DFT, computational exploration of chemical reactivity, molecular docking studies of novel formazan metal complexes and their biological applications. Appl Organomet Chem. https://doi.org/10.1002/aoc.5444
Mumit MA, Pal TK, Alam MA et al (2020) DFT studies on vibrational and electronic spectra, HOMO–LUMO, MEP, HOMA, NBO and molecular docking analysis of benzyl-3-N-(2,4,5-trimethoxyphenylmethylene)hydrazinecarbodithioate. J Mol Struct. https://doi.org/10.1016/j.molstruc.2020.128715
Pearson RG (1988) Electronic spectra and chemical reactivity. J Am Chem Soc 110:2092–2097. https://doi.org/10.1021/ja00215a013
Ma Y, Tao Y, Qu H et al (2022) Exploration of plant-derived natural polyphenols toward COVID-19 main protease inhibitors: DFT, molecular docking approach, and molecular dynamics simulations. RSC Adv 12:5357–5368. https://doi.org/10.1039/d1ra07364h
Radchenko EV, Dyabina AS, Palyulin VA (2016) Zefirov NS (2016) Prediction of human intestinal absorption of drug compounds. Russ Chem Bull 652(65):576–580. https://doi.org/10.1007/S11172-016-1340-0
Wadanambi PM, Mannapperuma U (2021) Computational study to discover potent phytochemical inhibitors against drug target, squalene synthase from Leishmania donovani. Heliyon 7:e07178. https://doi.org/10.1016/J.HELIYON.2021.E07178
Smith DA, Beaumont K, Maurer TS, Di L (2019) Clearance in drug design. J Med Chem 62:2245–2255. https://doi.org/10.1021/ACS.JMEDCHEM.8B01263/ASSET/IMAGES/MEDIUM/JM-2018-01263H_0010.GIF
Author information
Authors and Affiliations
Contributions
The authors' contributions to this study are as follows: MAY conceptualized and designed the research; conducted molecular docking, density functional theory and ADMET analysis; and prepared the original draft. SB and SS executed the molecular dynamics simulation.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical Approval
Not applicable.
Consent to Participate
All authors agreed to participate in this research.
Consent for Publication
All authors have approved the last version of the manuscript for its submission.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yousaf, M.A., Basheera, S. & Sivanandan, S. Inhibition of Monkeypox Virus DNA Polymerase Using Moringa oleifera Phytochemicals: Computational Studies of Drug-Likeness, Molecular Docking, Molecular Dynamics Simulation and Density Functional Theory. Indian J Microbiol (2024). https://doi.org/10.1007/s12088-024-01244-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12088-024-01244-3