Abstract
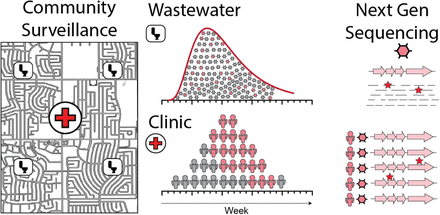
Areas of dense population congregation are prone to experience respiratory virus outbreaks. We monitored wastewater and clinic patients for the presence of respiratory viruses on a large, public university campus. Campus sewer systems were monitored in 16 locations for the presence of viruses using next generation sequencing over 22 weeks in 2023. During this period, we detected a surge in human adenovirus (HAdV) levels in wastewater. Hence, we initiated clinical surveillance at an on-campus clinic from patients presenting with acute respiratory infection. From whole genome sequencing of 123 throat and/or nasal swabs collected, we identified an outbreak of HAdV, specifically of HAdV-E4 and HAdV-B7 genotypes overlapping in time. The temporal dynamics and proportions of HAdV genotypes found in wastewater were corroborated in clinical infections. We tracked specific single nucleotide polymorphisms (SNPs) found in clinical virus sequences and showed that they arose in wastewater signals concordant with the time of clinical presentation, linking community transmission of HAdV to the outbreak. This study demonstrates how wastewater-based epidemiology can be integrated with surveillance at ambulatory healthcare settings to monitor areas prone to respiratory virus outbreaks and provide public health guidance.
Competing Interest Statement
The authors have declared no competing interest.
Funding Statement
This research was funded by Arizona State University, Tohono OOdham Nation (2020-01 ASU) and the Centers for Disease Control and Prevention (CDC U01 IP001180).
Author Declarations
I confirm all relevant ethical guidelines have been followed, and any necessary IRB and/or ethics committee approvals have been obtained.
Yes
The details of the IRB/oversight body that provided approval or exemption for the research described are given below:
This study was approved by site reliance on Duke University and Arizona State University Institutional Review Boards.
I confirm that all necessary patient/participant consent has been obtained and the appropriate institutional forms have been archived, and that any patient/participant/sample identifiers included were not known to anyone (e.g., hospital staff, patients or participants themselves) outside the research group so cannot be used to identify individuals.
Yes
I understand that all clinical trials and any other prospective interventional studies must be registered with an ICMJE-approved registry, such as ClinicalTrials.gov. I confirm that any such study reported in the manuscript has been registered and the trial registration ID is provided (note: if posting a prospective study registered retrospectively, please provide a statement in the trial ID field explaining why the study was not registered in advance).
Yes
I have followed all appropriate research reporting guidelines, such as any relevant EQUATOR Network research reporting checklist(s) and other pertinent material, if applicable.
Yes
Footnotes
Abbreviations: human adenovirus (HAdV), human coronavirus (HCoV), next generation sequencing (NGS), real-time quantitative reverse transcription PCR (qRT-PCR), human metapneumovirus (HMPV), human parainfluenza virus (HPIV), human rhinovirus (HRV)
Data availability
Sequencing data has been deposited to NCBI SRA (BioProject ID PRJNA1090887). Consensus genome sequences for human adenovirus B3, human adenovirus E4, human adenovirus B7, rhinovirus A, human metapneumovirus, human parainfluenza virus 1, human parainfluenza virus 2, human parainfluenza virus 3, human parainfluenza virus 4, human coronavirus NL63, human coronavirus HKU1 have been submitted to NCBI GenBank (accession numbers pending). SARS-CoV-2 genome consensus sequences have been deposited in the GISAID database with Accession IDs: EPI_ISL_17162425, EPI_ISL_17162426, EPI_ISL_17162427, EPI_ISL_17162428, EPI_ISL_17162429, EPI_ISL_17194616, EPI_ISL_17253217, EPI_ISL_17253218, EPI_ISL_17253219, EPI_ISL_17253220, EPI_ISL_17323764, EPI_ISL_17416692, EPI_ISL_17416693, EPI_ISL_17552873, EPI_ISL_17552874, EPI_ISL_17590094, EPI_ISL_17658673, EPI_ISL_17658674, EPI_ISL_17738172, EPI_ISL_17738173, EPI_ISL_17783646, EPI_ISL_17789350. Influenza genome consensus sequences have been deposited in the GISAID database with Accession IDs: EPI_ISL_17648695 and EPI_ISL_17648783.