Abstract
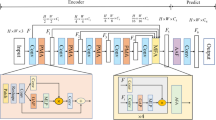
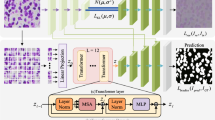
Medical image segmentation has witnessed rapid advancements with the emergence of encoder–decoder based methods. In the encoder–decoder structure, the primary goal of the decoding phase is not only to restore feature map resolution, but also to mitigate the loss of feature information incurred during the encoding phase. However, this approach gives rise to a challenge: multiple up-sampling operations in the decoder segment result in the loss of feature information. To address this challenge, we propose a novel network that removes the decoding structure to reduce feature information loss (CBL-Net). In particular, we introduce a Parallel Pooling Module (PPM) to counteract the feature information loss stemming from conventional and pooling operations during the encoding stage. Furthermore, we incorporate a Multiplexed Dilation Convolution (MDC) module to expand the network's receptive field. Also, although we have removed the decoding stage, we still need to recover the feature map resolution. Therefore, we introduced the Global Feature Recovery (GFR) module. It uses attention mechanism for the image feature map resolution recovery, which can effectively reduce the loss of feature information. We conduct extensive experimental evaluations on three publicly available medical image segmentation datasets: DRIVE, CHASEDB and MoNuSeg datasets. Experimental results show that our proposed network outperforms state-of-the-art methods in medical image segmentation. In addition, it achieves higher efficiency than the current network of coding and decoding structures by eliminating the decoding component.





Similar content being viewed by others
Data Availability
The dataset used in this paper is publicly available.
References
Zhan, B., Song, E., & Liu, H. (2023). FSA-Net: Rethinking the attention mechanisms in medical image segmentation from releasing global suppressed information. Computers in Biology and Medicine, 161, 106932. https://doi.org/10.1016/j.compbiomed.2023.106932
Shin, H.-C., Roth, H. R., Gao, M., Lu, L., Xu, Z., Nogues, I., Yao, J., Mollura, D., & Summers, R. M. (2016). Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Transactions on Medical Imaging, 35(5), 1285–1298. https://doi.org/10.1109/TMI.2016.2528162
Foruzan, A. H., Zoroofi, R. A., Sato, Y., & Hori, M. (2012). A Hessian-based filter for vascular segmentation of noisy hepatic CT scans. International Journal of Computer Assisted Radiology and Surgery, 7(2), 199–205. https://doi.org/10.1007/s11548-011-0640-y
Staal, J., Abràmoff, M. D., Niemeijer, M., Viergever, M. A., & Van Ginneken, B. (2004). Ridge-based vessel segmentation in color images of the retina. IEEE Transactions on Medical Imaging, 23(4), 501–509. https://doi.org/10.1109/TMI.2004.825627
Lecun, Y., Bengio, Y., & Hinton, G. (2015). Deep learning. Nature, 521(7553), 436–444. https://doi.org/10.1038/nature14539
Long, J., Shelhamer, E., & Darrell, T. (2015). Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA (pp. 3431–3440). https://doi.org/10.1109/cvpr.2015.7298965
Ronneberger, O., Fischer, P., & Brox, T. (2015). U-net: Convolutional networks for biomedical image segmentation. In: International Conference on Medical Image Computing and Computer Assisted Intervention, Munich, Germany (pp. 234–241). https://doi.org/10.1007/978-3-319-24574-4_28
Zhou, Z., Rahman Siddiquee, M. M., Tajbakhsh, N., & Liang, J. (2018). Unet++: A nested u-net architecture for medical image segmentation. In: Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support: 4th International Workshop, DLMIA 2018, and 8th International Workshop, ML-CDS 2018, Held in Conjunction with MICCAI 2018, Proceedings 4, 2018, Granada, Spain (pp. 3–11). https://doi.org/10.1007/978-3-030-00889-5_1
Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T., & Ronneberger, O. (2016). 3D U-Net: Learning dense volumetric segmentation from sparse annotation. In: Medical Image Computing and Computer-assisted Intervention–MICCAI 2016: 19th International Conference, Proceedings, Part II 19, 2016, Athens, Greece (pp. 424–432). https://doi.org/10.1007/978-3-319-46723-8_49
Ibtehaz, N., & Rahman, M. S. (2020). MultiResUNet: Rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural Networks, 121, 74–87. https://doi.org/10.1016/j.neunet.2019.08.025
Guan, S., Khan, A. A., Sikdar, S., & Chitnis, P. V. (2019). Fully dense UNet for 2-D sparse photoacoustic tomography artifact removal. IEEE Journal of Biomedical and Health Informatics, 24(2), 568–576. https://doi.org/10.1109/JBHI.2019.2912935
Alom, M. Z., Hasan, M., Yakopcic, C., Taha, T. M., & Asari, V. K. (2018). Recurrent residual convolutional neural network based on u-net (r2u-net) for medical image segmentation. arXiv preprint arXiv:1802.06955. https://doi.org/10.48550/arXiv.1802.06955
Zhang, J., Li, C., Kosov, S., Grzegorzek, M., Shirahama, K., Jiang, T., Sun, C., Li, Z., & Li, H. (2021). LCU-Net: A novel low-cost U-Net for environmental microorganism image segmentation. Pattern Recognition, 115, 107885. https://doi.org/10.1016/j.patcog.2021.107885
Ni, J., Liu, J., Li, X., & Chen, Z. (2022). SFA-Net: Scale and feature aggregate network for retinal vessel segmentation. Journal of Healthcare Engineering, 2022, 4695136. https://doi.org/10.1155/2022/4695136
Huang, H., Lin, L., Tong, R., Hu, H., Zhang, Q., Iwamoto, Y., Han, X., Chen, Y.-W., & Wu, J. (2020). Unet 3+: A full-scale connected unet for medical image segmentation. In: ICASSP 2020–2020 IEEE International Conference on Acoustics, Speech and Signal Processing 2020, Barcelona, Spain (pp. 1055–1059). https://doi.org/10.1109/icassp40776.2020.9053405
Chen, J., Lu, Y., Yu, Q., Luo, X., Adeli, E., Wang, Y., Lu, L., Yuille, A.L., & Zhou, Y. (2021). Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306. https://doi.org/10.48550/arXiv.2102.04306
Cao, H., Wang, Y., Chen, J., Jiang, D., Zhang, X., Tian, Q., & Wang, M. (2022). Swin-unet: Unet-like pure transformer for medical image segmentation. In European Conference on Computer Vision (pp. 205–218). Springer. https://doi.org/10.1007/978-3-031-25066-8_9
Ni, J., Sun, H., Xu, J., Liu, J., & Chen, Z. (2023). A feature aggregation and feature fusion network for retinal vessel segmentation. Biomedical Signal Processing and Control, 85, 104829. https://doi.org/10.1016/j.bspc.2023.104829
Ni, J., Wu, J., Elazab, A., Tong, J., & Chen, Z. (2022). DNL-Net: Deformed non-local neural network for blood vessel segmentation. BMC Medical Imaging, 22(1), 1–14. https://doi.org/10.1186/s12880-022-00836-z
Wu, H., Wang, W., Zhong, J., Lei, B., Wen, Z., & Qin, J. (2021). Scs-net: A scale and context sensitive network for retinal vessel segmentation. Medical Image Analysis, 70, 102025. https://doi.org/10.1016/j.media.2021.102025
Fu, J., Liu, J., Tian, H., Li, Y., Bao, Y., Fang, Z., & Lu, H. (2019). Dual attention network for scene segmentation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Los Angeles CA, United States (pp. 3146–3154). https://doi.org/10.1109/CVPR.2019.00326
Ni, J., Wu, J., Tong, J., Chen, Z., & Zhao, J. (2020). GC-Net: Global context network for medical image segmentation. Computer Methods and Programs in Biomedicine, 190, 105121. https://doi.org/10.1016/j.cmpb.2019.105121
Ni, J., Wu, J., Wang, H., Tong, J., Chen, Z., Wong, K. K., & Abbott, D. (2020). Global channel attention networks for intracranial vessel segmentation. Computers in Biology and Medicine, 118, 103639. https://doi.org/10.1016/j.compbiomed.2020.103639
Guo, C., Szemenyei, M., Yi, Y., Wang, W., Chen, B., & Fan, C. (2021). Sa-unet: Spatial attention u-net for retinal vessel segmentation. In: 2020 25th International Conference on Pattern Recognition (ICPR), Milan, Italy (pp. 1236–1242). https://doi.org/10.1109/ICPR48806.2021.9413346
Wang, C., Wang, Y., Liu, Y., He, Z., He, R., & Sun, Z. (2019). ScleraSegNet: An attention assisted U-Net model for accurate sclera segmentation. IEEE Transactions on Biometrics, Behavior, and Identity Science, 2(1), 40–54. https://doi.org/10.1109/TBIOM.2019.2962190
Woo, S., Park, J., Lee, J.-Y., & So Kweon, I. (2018). Cbam: Convolutional block attention module. In: Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany (pp. 3–19). https://doi.org/10.1007/978-3-030-01234-2_1
Oktay, O., Schlemper, J., Folgoc, L. L., Lee, M., Heinrich, M., Misawa, K., Mori, K., Mcdonagh, S., Hammerla, N. Y., & Kainz, B. (2018). Attention u-net: Learning where to look for the pancreas. arXiv preprint arXiv:1804.03999. https://doi.org/10.48550/arXiv.1804.03999
Roy, A.G., Navab, N. & Wachinger, C. (2018). Concurrent spatial and channel ‘squeeze & excitation’ in fully convolutional networks. In: International Conference on Medical Image Computing and Computer Assisted Intervention, Granada, Spain (pp. 421–429). https://doi.org/10.1007/978-3-030-00928-1_48
Qin, Y., Kamnitsas, K., Ancha, S., Nanavati, J., Cottrell, G., Criminisi, A., & Nori, A. (2018). Autofocus layer for semantic segmentation. In: International Conference on Medical Image Computing and Computer-assisted Intervention, Granada, Spain (pp. 603–611). https://doi.org/10.1007/978-3-030-00931-1_69
Wang, Y., Deng, Z., Hu, X., Zhu, L., Yang, X., Xu, X., Heng, P.-A., & Ni, D. (2018). Deep attentional features for prostate segmentation in ultrasound. In: International Conference on Medical Image Computing and Computer-assisted Intervention, Granada, Spain (pp. 523–530). https://doi.org/10.1007/978-3-030-00937-3_60
Zhu, H., Zeng, H., Liu, J., & Zhang, X. (2021). Logish: A new nonlinear nonmonotonic activation function for convolutional neural network. Neurocomputing, 458, 490–499. https://doi.org/10.1016/j.neucom.2021.06.067
Chollet, F. (2017). Xception: Deep learning with depth wise separable convolutions. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, Hawaii (pp. 1251–1258). https://doi.org/10.1109/CVPR.2017.195
Owen, C. G., Rudnicka, A. R., Mullen, R., Barman, S. A., Monekosso, D., Whincup, P. H., Ng, J., & Paterson, C. (2009). Measuring retinal vessel tortuosity in 10-year-old children: Validation of the computer-assisted image analysis of the retina (CAIAR) program. Investigative Ophthalmology and Visual Science, 50(5), 2004–2010. https://doi.org/10.1167/iovs.08-3018
Kumar, N., Verma, R., Anand, D., Zhou, Y., Onder, O. F., Tsougenis, E., Chen, H., Heng, P.-A., Li, J., & Hu, Z. (2019). A multi-organ nucleus segmentation challenge. IEEE Transactions on Medical Imaging, 39(5), 1380–1391. https://doi.org/10.1109/TMI.2019.2947628
Funding
This study was funded by the National Key Research and Development Program of China (Grant 2020YFB1708900) and the Fundamental Research Funds for the Central Universities (Grant No. B220201044).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ni, J., Mu, W., Pan, A. et al. Rethinking the Encoder–decoder Structure in Medical Image Segmentation from Releasing Decoder Structure. J Bionic Eng (2024). https://doi.org/10.1007/s42235-024-00513-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42235-024-00513-7