Abstract
Purpose
The standard of care for prostate cancer (PCa) diagnosis is the histopathological analysis of tissue samples obtained via transrectal ultrasound (TRUS) guided biopsy. Models built with deep neural networks (DNNs) hold the potential for direct PCa detection from TRUS, which allows targeted biopsy and subsequently enhances outcomes. Yet, there are ongoing challenges with training robust models, stemming from issues such as noisy labels, out-of-distribution (OOD) data, and limited labeled data.
Methods
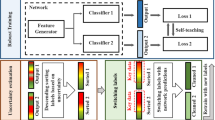
This study presents LensePro, a unified method that not only excels in label efficiency but also demonstrates robustness against label noise and OOD data. LensePro comprises two key stages: first, self-supervised learning to extract high-quality feature representations from abundant unlabeled TRUS data and, second, label noise-tolerant prototype-based learning to classify the extracted features.
Results
Using data from 124 patients who underwent systematic prostate biopsy, LensePro achieves an AUROC, sensitivity, and specificity of 77.9%, 85.9%, and 57.5%, respectively, for detecting PCa in ultrasound. Our model shows it is effective for detecting OOD data in test time, critical for clinical deployment. Ablation studies demonstrate that each component of our method improves PCa detection by addressing one of the three challenges, reinforcing the benefits of a unified approach.
Conclusion
Through comprehensive experiments, LensePro demonstrates its state-of-the-art performance for TRUS-based PCa detection. Although further research is necessary to confirm its clinical applicability, LensePro marks a notable advancement in enhancing automated computer-aided systems for detecting prostate cancer in ultrasound.




Similar content being viewed by others
Notes
Excessive hand motion is detected by checking the B-mode videos recorded during the biopsy procedure.
We used a composition of random rotation (\(\pm 20\deg \)), contrast fluctuation, and crops.
References
Smeenge M, de la Rosette JJMCH, Wijkstra H (2012) Current status of transrectal ultrasound techniques in prostate cancer. Curr Opin Urol 22(4):297–302
Ahmed HU, El-Shater Bosaily A, Brown LC, Gabe R, Kaplan R, Parmar MK, Collaco-Moraes Y, Ward K, Hindley RG, Freeman A et al (2017) Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. The Lancet 389(10071):815–822
Madej A, Wilkosz J, Różański W, Lipiński M (2012) Complication rates after prostate biopsy according to the number of sampled cores. Cent Eur J Urol 65(3):116
Feng Y, Yang F, Zhou X, Guo Y, Tang F, Ren F, Guo J, Ji S (2018) A deep learning approach for targeted contrast-enhanced ultrasound based prostate cancer detection. IEEE/ACM Trans Comput Biol Bioinf 16(6):1794–1801
Gilany M, Wilson P, Jamzad A, Fooladgar F, To MNN, Wodlinger B, Abolmaesumi P, Mousavi P (2022) Towards confident detection of PCa using high resolution micro-ultrasound. In: Medical image computing and computer assisted interventions, pp 411–420
Fooladgar F, To MNN, Javadi G, Samadi S, Bayat S, Sojoudi S, Eshumani W, Hurtado A, Chang S, Black P, et al (2022) Uncertainty-aware deep ensemble model for targeted ultrasound-guided prostate biopsy. In: 2022 IEEE 19th international symposium on biomedical imaging (ISBI), pp 1–5
Javadi G, Samadi S, Bayat S, Sojoudi S, Hurtado A, Chang S, Black P, Mousavi P, Abolmaesumi P (2021) Training deep networks for prostate cancer diagnosis using coarse histopathological labels. In: International conference on medical image computing and computer-assisted intervention, pp 680–689
Le H, Samaras D, Kurc T, Gupta R, Shroyer K, Saltz J (2019) Pancreatic cancer detection in whole slide images using noisy label annotations. In: Medical image computing and computer assisted intervention. Springer, pp 541–549
Ashraf M, Robles WRQ, Kim M, Ko YS, Yi MY (2022) A loss-based patch label denoising method for improving whole-slide image analysis using a convolutional neural network. Sci Rep 12(1):1392
Javadi G, Samadi S, Bayat S, Pesteie M, Jafari MH, Sojoudi S, Kesch C, Hurtado A, Chang S, Mousavi P et al (2020) Multiple instance learning combined with label invariant synthetic data for guiding systematic prostate biopsy: a feasibility study. Int J Comput Assist Radiol Surg 15(6):1023–1031
Linmans J, Elfwing S, van der Laak J, Litjens G (2023) Predictive uncertainty estimation for out-of-distribution detection in digital pathology. Med Image Anal 83:102655
Karimi D, Gholipour A (2022) Improving calibration and out-of-distribution detection in deep models for medical image segmentation. IEEE Trans Artif Intell 4:383–397
Azizi S, Mustafa B, Ryan F, Beaver Z, Freyberg J, Deaton J, Loh A, Karthikesalingam A, Kornblith S, Chen T, et al (2021) Big self-supervised models advance medical image classification. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 3478–3488
Wilson PFR, Gilany M, Jamzad A, Fooladgar F, To MNN, Wodlinger B, Abolmaesumi P, Mousavi P (2023) Self-supervised learning with limited labeled data for prostate cancer detection in high frequency ultrasound. IEEE Trans Ultrason Ferroelectr Freq Control 70:1073–1083
Jiang Y, Sui X, Ding Y, Xiao W, Zheng Y, Zhang Y (2022) A semi-supervised learning approach with consistency regularization for tumor histopathological images analysis. Front Oncol 12:7200
Macêdo D, Ren TI, Zanchettin C, Oliveira ALI, Ludermir T (2021) Entropic out-of-distribution detection. In: 2021 international joint conference on neural networks. IEEE, pp 1–8
To MNN, Fooladgar F, Javadi G, Bayat S, Sojoudi S, Hurtado A, Chang S, Black P, Mousavi P, Abolmaesumi P (2022) Coarse label refinement for improving prostate cancer detection in ultrasound imaging. Int J Comput Assist Radiol Surg 17:1–7
Macêdo D, Ludermir T (2021) Enhanced isotropy maximization loss: seamless and high-performance out-of-distribution detection simply replacing the softmax loss. arXiv preprint, arXiv:2105.14399
Bardes A, Ponce J, Lecun Y (2022) Vicreg: variance-invariance-covariance regularization for self-supervised learning. In: ICLR 2022-international conference on learning representations
Karim N, Rizve MN, Rahnavard N, Mian A, Shah M (2022) Unicon: combating label noise through uniform selection and contrastive learning. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 9676–9686
Li J, Socher R, Hoi SCH (2019) Dividemix: learning with noisy labels as semi-supervised learning. In: International conference on learning representations
Han B, Yao Q, Yu X, Niu G, Xu M, Hu W, Tsang I, Sugiyama M (2018) Co-teaching: robust training of deep neural networks with extremely noisy labels. In: Advances in neural information processing systems, vol 31
Zhou X, Liu X, Jiang J, Gao X, Ji X (2021) Asymmetric loss functions for learning with noisy labels. In: International conference on machine learning. PMLR, pp 12846–12856
Zhang Z, Sabuncu M (2018) Generalized cross entropy loss for training deep neural networks with noisy labels. In: Advances in neural information processing systems, vol 31
Xia X, Han B, Zhan Y, Yu J, Gong M, Gong C, Liu T (2023) Combating noisy labels with sample selection by mining high-discrepancy examples. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 1833–1843
Acknowledgements
This research is supported by Natural Sciences and Engineering Research Council of Canada (NSERC) and the Canadian Institutes of Health Research (CIHR). Parvin Mousavi is supported by Canada CIFAR AI Chair and the Vector Institute. We acknowledge the staff at Vancouver General Hospital who assisted with data acquisition for our study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors confirm that there are no known conflicts of interest with this publication.
Ethical approval
All studies involving human participants were in accordance with the ethical standards of the institutional research board and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
To, M.N.N., Fooladgar, F., Wilson, P. et al. LensePro: label noise-tolerant prototype-based network for improving cancer detection in prostate ultrasound with limited annotations. Int J CARS (2024). https://doi.org/10.1007/s11548-024-03104-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11548-024-03104-3