Abstract
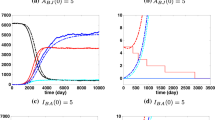
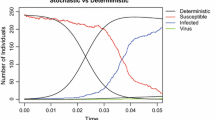
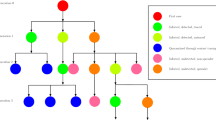
The potential of zoonotic diseases to cross-species physical boundaries makes them a persistent threat to global public health. Of all the zoonoses, the bubonic plague serves as a historical and modern model. In this work, we investigate a bubonic plague stochastic model using a continuous-time Markov chain (CTMC) model to study the disease dynamics in rats and humans. Using a Galton–Watson multi-type (GWbp) branching process, we have derived an analytical expression for disease extinction probability at the beginning of the epidemic. There is a variation in disease extinction probability calculated via the branching process and numerical simulations, which is the consequence of the discrete assumption of an infected flea instead of being considered in terms of an infected rat in the numerical simulation. An analytical expression for the distribution of first passage time (FPT) to spillover is also obtained in this work using the probability generating function (PGF) technique, and it agrees well with the FPT distribution obtained numerically. This distribution is defective since infected fleas can die before infecting human. Furthermore, the effect of the initial rat population on the FPT of spillover is also shown, which suggest that as the initial rat population increases, the likelihood of spillover decreases, and it increases as the rat population decreases. Additionally, we have derived an expression for the mean and variance of the first passage time to spillover, and we incorporate the impact of various parameters on the mean first passage time of spillover. Lastly, we have performed numerical simulations to estimate the peak size of each infected class and the related time to attain peak infection. The peak of the infected flea, which is a disease carrier, is much higher as compared to that of rat and human. Also, there is a delay in attaining peak size for the human population, which is the consequence of the human population being a secondary host for fleas to survive.







Similar content being viewed by others
Data Availability Statement
Data sharing is not applicable to this article as no new data were created or analysed in this study.
References
W.B. Karesh, A. Dobson, J.O. Lloyd-Smith, J. Lubroth, M.A. Dixon, M. Bennett, S. Aldrich, T. Harrington, P. Formenty, E.H. Loh et al., Ecology of zoonoses: natural and unnatural histories. The Lancet 380(9857), 1936–1945 (2012)
J.O. Lloyd-Smith, D. George, K.M. Pepin, V.E. Pitzer, J.R.C. Pulliam, A.P. Dobson, P.J. Hudson, B.T. Grenfell, Epidemic dynamics at the human–animal interface. science 326(5958), 1362–1367 (2009)
S. Singh, D.J. Schneider, C.R. Myers, Using multitype branching processes to quantify statistics of disease outbreaks in zoonotic epidemics. Phys. Rev. E 89(3), 032702 (2014)
R. Coker, J. Rushton, S. Mounier-Jack, E. Karimuribo, P. Lutumba, D. Kambarage, D.U. Pfeiffer, K. Stärk, M. Rweyemamu, Towards a conceptual framework to support one-health research for policy on emerging zoonoses. Lancet. Infect. Dis 11(4), 326–331 (2011)
W.B. Karesh, R.A. Cook, The human-animal link. Foreign Aff. 84, 38 (2005)
L.J.S. Allen, V.L. Brown, C.B. Jonsson, S.L. Klein, S.M. Laverty, K. Magwedere, J.C. Owen, P. Van Den Driessche, Mathematical modeling of viral zoonoses in wildlife. Nat. Resour. Model. 25(1), 5–51 (2012)
M.J. Keeling, C.A. Gilligan, Metapopulation dynamics of bubonic plague. Nature 407(6806), 903–906 (2000)
F. Gascuel, M. Choisy, J.-M. Duplantier, F. Débarre, C. Brouat, Host resistance, population structure and the long-term persistence of bubonic plague: contributions of a modelling approach in the Malagasy focus. PLoS Comput. Biol. 9(5), e1003039 (2013)
World Health Organization et al. Human plague: review of regional morbidity and mortality, 2004–2009: introduction. Weekly Epidemiological Record= Relevé épidémiologique Hebdomadaire 85(06), 40–45 (2010)
S. Chanteau, Atlas de la peste à Madagascar. IRD editions (2006)
R.C. Ngeleja, L.S. Luboobi, Y. Nkansah-Gyekye, Modelling the dynamics of bubonic plague with yersinia pestis in the environment. Commun. Math. Biol. Neurosci., Article-ID (2016)
R.R. Parmenter, E.P. Yadav, C.A. Parmenter, P. Ettestad, K.L. Gage, Incidence of plague associated with increased winter–spring precipitation in new Mexico. Am. J. Trop. Med. Hyg. 61(5), 814–821 (1999)
W.E. Wingfield, S.B. Palmer, Veterinary Disaster Response (Wiley, New York, 2009)
C. McEvedy, The bubonic plague. Sci. Am. 258(2), 118–123 (1988)
A.J. Nicholson, V.A. Bailey, The balance of animal populations—part I. In: Proceedings of the Zoological Society of London, vol. 105, pp. 551–598. Wiley Online Library (1935)
K.R. Dean, F. Krauer, L. Walløe, O.C. Lingjærde, B. Bramanti, N.C. Stenseth, B.V. Schmid, Human ectoparasites and the spread of plague in Europe during the second pandemic. Proc. Natl. Acad. Sci. 115(6), 1304–1309 (2018)
M.J. Keeling, C.A. Gilligan, Bubonic plague: a metapopulation model of a zoonosis. Proc. R. Soc. Lond. B 267(1458), 2219–2230 (2000)
M. Buzby, D. Neckels, M.F. Antolin, D. Estep, Analysis of the sensitivity properties of a model of vector-borne bubonic plague. J. R. Soc. Interface 5(26), 1099–1107 (2008)
J.E. Foley, J. Zipser, B. Chomel, E. Girvetz, P. Foley, Modeling plague persistence in host-vector communities in California. J. Wildl. Dis. 43(3), 408–424 (2007)
J.A. Mwasunda, J.I. Irunde, D. Kajunguri, D. Kuznetsov, Outbreak or extinction of bovine cysticercosis and human taeniasis: a stochastic modelling approach. Appl. Math. Model. 106, 73–85 (2022)
T. Antal, P.L. Krapivsky, Exact solution of a two-type branching process: models of tumor progression. J. Stat. Mech.: Theory Exp. 2011(08), P08018 (2011)
K.B. Athreya, P.E. Ney, K.B. Athreya, P.E. Ney, One Dimensional Continuous Time Markov Branching Processes. Branching Processes (Springer, Berlin, 1972), pp.102–136
N.T.J. Bailey, The Elements of Stochastic Processes with Applications to the Natural Sciences, vol. 25 (Wiley, New York, 1991)
J. Curran-Sebastian, L. Pellis, I. Hall, T. House, Calculation of Epidemic First Passage and Peak Time Probability Distributions. arXiv preprintarXiv:2301.07058 (2023)
J.C. Miller, A primer on the use of probability generating functions in infectious disease modeling. Infect. Dis. Model. 3, 192–248 (2018)
B.-S. Chen, Z.-Y. Wu, Y.-J. Chen, J.-L. Chern, Data forecasts of the epidemic Covid-19 by deterministic and stochastic time-dependent models. arXiv preprint arXiv:2209.02880 (2022)
Z.G. İşlier, R. Güllü, W. Hörmann, An exact and implementable computation of the final outbreak size distribution under Erlang distributed infectious period. Math. Biosci. 325, 108363 (2020)
H.E. Daniels, The maximum size of a closed epidemic. Adv. Appl. Probab. 6(4), 607–621 (1974)
J.R. Artalejo, A. Economou, M.J. Lopez-Herrero, The maximum number of infected individuals in sis epidemic models: computational techniques and quasi-stationary distributions. J. Comput. Appl. Math. 233(10), 2563–2574 (2010)
J. Amador, M. Lopez-Herrero, Cumulative and maximum epidemic sizes for a nonlinear seir stochastic model with limited resources. Discret. Contin. Dyn. Syst. Ser. B 23(8), 3137 (2018)
J. Amador Pacheco, D. Armesto, A. Gómez-Corral, Extreme values in sir epidemic models with two strains and cross-immunity. Math. Biosci. & Engg. 16(4), 1992–2022 (2019). https://doi.org/10.3934/mbe.2019098
C.J. Edholm, B.O. Emerenini, A.L Murillo, O. Saucedo, N. Shakiba, X. Wang, L.J.S. Allen, A. Peace, Searching for superspreaders: Identifying epidemic patterns associated with superspreading events in stochastic models. Understanding complex biological systems with mathematics, p. 1–29 (2018)
O. Diekmann, J.A.P. Heesterbeek, M.G. Roberts, The construction of next-generation matrices for compartmental epidemic models. J. R. Soc. Interface 7(47), 873–885 (2010)
P. Van den Driessche, J. Watmough, Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math. Biosci. 180(1–2), 29–48 (2002)
P. Sarathi Mandal, S. Maity, Impact of demographic variability on the disease dynamics for honeybee model. Chaos Interdiscip. J. Nonlinear Sci. 32(8), 083120 (2022)
L.J.S. Allen, A primer on stochastic epidemic models: formulation, numerical simulation, and analysis. Infect. Dis. Model. 2(2), 128–142 (2017)
M. Maliyoni, Probability of disease extinction or outbreak in a stochastic epidemic model for West Nile virus dynamics in birds. Acta. Biotheor. 69(2), 91–116 (2021)
M. Maliyoni, F. Chirove, H.D. Gaff, K.S. Govinder, A stochastic tick-borne disease model: exploring the probability of pathogen persistence. Bull. Math. Biol. 79, 1999–2021 (2017)
M. Maliyoni, F. Chirove, H.D. Gaff, K.S. Govinder, A stochastic epidemic model for the dynamics of two pathogens in a single tick population. Theor. Popul. Biol. 127, 75–90 (2019)
S. Maity, P.S. Mandal, A comparison of deterministic and stochastic plant–vector–virus models based on probability of disease extinction and outbreak. Bull. Math. Biol. 84(3), 41 (2022)
R.J. Eisen, A.P. Wilder, S.W. Bearden, J.A. Montenieri, K.L. Gage, Early-phase transmission of Yersinia pestis by Unblocked Xenopsylla cheopis (Siphonaptera: Pulicidae) is as efficient as transmission by blocked fleas. J. Med. Entomol. 44(4), 678–682 (2007)
L.A. White, L. Mordechai, Modeling the justinianic plague: comparing hypothesized transmission routes. PLoS (30 Apr. 2020) (2020)
S. Hautphenne, Extinction probabilities of supercritical decomposable branching processes. J. Appl. Probab. 49(3), 639–651 (2012)
S. Hautphenne, G. Latouche, G. Nguyen, Extinction probabilities of branching processes with countably infinitely many types. Adv. Appl. Probab. 45(4), 1068–1082 (2013)
Acknowledgements
Suman Kumari’s research is supported by research fellowship from MHRD, Government of India, Partha Sarathi Mandal’s research is supported by CSIR Project [File No: 25(0303)/19/EMR-II].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts to disclose.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumari, S., Mandal, P.S. & Sen, M. First passage time and peak size probability distributions for a complex epidemic model. Eur. Phys. J. Plus 139, 324 (2024). https://doi.org/10.1140/epjp/s13360-024-05108-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjp/s13360-024-05108-z