Abstract
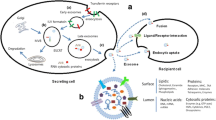
Breast cancer is one of the most prevalent malignancies affecting women globally and poses a significant public health challenge. Early clinical detection plays a pivotal role in providing optimal treatment opportunities and favorable prognoses, crucial for reducing breast cancer mortality and enhancing patients’ quality of life. Therefore, the timely identification and diagnosis of breast cancer are imperative. Conventional tumor markers, such as carcinoembryonic antigen (CEA) and carbohydrate antigen 15-3 (CA15-3), serve as reliable methods for actively monitoring disease progression and have become a routine auxiliary diagnostic approach in clinical settings. However, these biomarkers exhibit limitations in sensitivity and specificity, particularly in the early screening and diagnosis of tumors, often yielding results inconsistent with clinical manifestations. In recent years, research has increasingly focused on exosomes released by tumor cells as potential new biomarkers for early stage breast cancer screening. Exosomes carry various components, including tumor-derived proteins, nucleic acids, and lipids. This paper delves into the specific utilization of serum exosomal microRNA-21 (miR-21) as a biomarker for early detection and diagnosis of breast cancer, evaluating its efficacy within this framework.
Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- CEA:
-
Carcinoembryonic antigen
- CA15-3:
-
Carbohydrate antigen 15-3
- miR-21 (microRNA-21):
-
Microribonucleic acid-21
- mRNAs:
-
Messenger RNAs
- miRNAs:
-
MicroRNAs
- ncRNAs:
-
Non-coding RNAs
- lncRNAs:
-
Long non-coding RNAs
- Bcl-2:
-
B-cell lymphoma 2
- BRCA1:
-
Breast cancer susceptibility gene 1
- BRCA2:
-
Breast cancer susceptibility gene 2
- P53:
-
Tumor suppressor protein p53
- DOR:
-
Diagnostic odds ratio
- NLR:
-
Negative likelihood of detection results ratio
- PLR:
-
Positive likelihood ratio
- CI:
-
Confidence interval
- PTEN:
-
Phosphatase and tensin homolog deleted on chromosome 10
- PI3K:
-
The phosphoinositide 3-kinase
- PI3K/AKT/mTOR:
-
Phosphoinositide 3-kinase/v-akt murine thymoma viral oncogene/mammalian target of rapamycin
- MFC-7:
-
Michigan Cancer Foundation-7
- PDCD4:
-
Programmed cell death protein 4
- EGF:
-
Epithelial growth factor
- TGF-β:
-
Transforming growth factor β
- TPM1:
-
Tumor suppressor gene tropomyosin 1
- Maspin:
-
Mammary serine protease inhibitor
- LC/MS/MS:
-
Liquid chromatography/mass spectrometry
- RPM1:
-
Receptor protein against Pseudomonas MACULICOLA1
- HER2:
-
Human epidermal growth receptor 2
- BCSC:
-
Breast cancer stem cells
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in185 countries. CA Cancer J Clin. 2021;71(3):209–49.
Rabbani F, Yazdiniapour Z, Ghanadian M, Zolfaghari B, Maleki M, Shafiee F. Cytotoxicity and apoptosis assay of novel cyclomyrsinol diterpenes against breast cancer cell lines. World J Tradit Chin Med. 2022;8:273–7.
Tang Z, Xu L, Sun Z, Chen W, Li R. Differential diagnosis of benign and malignant breast lesions using mammography and 3.0T magnetic resonance imaging. Chin Med Equip. 2022;19(10):48–52.
Huang W-H, Chen I-W, Yang C-C, Wang T-Y. The sonography is a valuable tool for diagnosis of microcalcification in screening mammography. Ultrasound Med Biol. 2017;43:S28.
Wang H, Song Q, Zheng G, Cao g. Study on the diagnosis of early breast cancer with molybdenum target and ultrasound Doppler combined with serum tumor markers. China Med Equip. 2024;21(1):82–7.
Li X, Xu Y, Zhang L. Serum CA153 as biomarker for cancer and noncancer diseases. Prog Mol Biol Transl Sci. 2019;162:265–76.
Jia L, Li G, Ma N, Zhang A, Zhou Y, Ren L, et al. Soluble POSTN is a novel biomarker complementing CA153 and CEA for breast cancer diagnosis and metastasis prediction. BMC Cancer. 2022;22(1):760.
Lin Z. Clinical significance of CA15-3, CA125 and CEA joint detection in the diagnosis of breast cancer. Exp Lab Med. 2009;27(3):261–2.
Tanman Ü, Yangın S, Cansaran-Duman D. Determination of dysregulated miRNA expression levels by qRT-PCR after the application of usnic acid to breast cancer. J Anticancer Agents Med Chem. 2020;20(5):548–58.
Ortega MA, Fraile-Martínez O, Guijarro LG, Casanova C, Coca S, Álvarez-Mon M, et al. The regulatory role of mitochondrial microRNAs (MitomiRs) in breast cancer: translational implications present and future. Cancers (Basel). 2020;12(9):2443.
Ren FL, Li P, Wang CM, Chen QH. Application of exosomes in cancer diagnosis and treatment and progress in isolation technology. J Chin Oncol. 2020;26(6):475–80.
Boriachek K, Islam MN, Möller A, Salomon C, Nguyen NT, Hossain MSA, et al. Biological functions and current advances in isolation and detection strategies for exosome nanovesicles. Small. 2018;14(6).
Lauretti E, Dabrowski K, Praticò D. The neurobiology of non-coding RNAs and Alzheimer’s disease pathogenesis: pathways, mechanisms and translational opportunities. Ageing Res Rev. 2021;71:101425.
Hill M, Tran N. miRNA interplay: mechanisms and consequences in cancer. Dis Model Mech. 2021;14(4):dmm047662.
Kim MW, Park S, Lee H, Gwak H, Hyun KA, Kim JY, et al. Multi-miRNA panel of tumor-derived extracellular vesicles as promising diagnostic biomarkers of early-stage breast cancer. Cancer Sci. 2021;112(12):5078–87.
Abdel-Hamid NR, Mohammed EA, Abbas AH, Badr FM. MicroRNA-21 expression in primary breast cancer tissue among Egyptian female patients and its correlation with chromosome 17 aneusomy. Mol Diagn Ther. 2015;19(6):365–73.
Shaaban NZ, Ibrahim NK, Saada HN, El-Rashidy FH, Shaaban HM, Elbakary NM, et al. The implication of microRNAs as non-invasive biomarkers in 179 Egyptian breast cancer female patients. Oncol Res. 2023;30(6):269–76.
Wang M, Wang Y, Tian X, Wang Q, Huang H, Lu X, et al. Diagnostic and predictive value of liquid biopsy-derived exosome miR-21 for breast cancer: a systematic review and meta-analysis. Expert Rev Mol Diagn. 2023;23(4):315–24.
Yu X, Liang J, Xu J, Li X, Xing S, Li H, et al. Identification and validation of circulating microRNA signatures for breast cancer early detection based on large scale tissue-derived data. J Breast Cancer. 2018;21(4):363–70.
Liang Y, Hao M, Guo R, Li X, Li Y, Yu C, et al. Biomarkers for early screening and diagnosis of breast cancer: a review. Sheng Wu Gong Cheng Xue Bao. 2023;39(4):1425–44.
García Z, Kumar A, Marqués M, Cortés I, Carrera AC. Phosphoinositide 3-kinase controls early and late events in mammalian cell division. EMBO J. 2006;25(4):655–61.
Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011;57(1):84–91.
Frankel LB, Christoffersen NR, Jacobsen A, Lindow M, Krogh A, Lund AH. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem. 2008;283(2):1026–33.
Han M, Wang F, Gu Y, Pei X, Guo G, Yu C, et al. MicroRNA- 21 induces breast cancer cell invasion and migration by suppressing smad7 via EGF and TGF-β pathways. Oncol Rep. 2016;35(1):73–80.
Wu MF, Yang J, Xiang T, Shi YY, Liu LJ. miR-21 targets Fas ligand-mediated apoptosis in breast cancer cell line MCF-7. J Huazhong Univ Sci Technol Med Sci. 2014;34(2):190–4.
Wang W, Li J, Zhu W, Gao C, Jiang R, Li W, et al. MicroRNA-21 and the clinical outcomes of various carcinomas: a systematic review and meta-analysis. BMC Cancer. 2014;14:819. https://doi.org/10.1186/1471-2407-14-819.
Zhu W, Xu B. MicroRNA-21 identified as predictor of cancer outcome: a meta-analysis. PLoS One. 2014;9(8):103373.
Desouky EM, Khaliefa AK, Hozayen WG, Shaaban SM, Hasona NA. Signature of miR-21 and MEG-2 and their correlation with TGF-β signaling in breast cancer. Hum Exp Toxicol. 2023;42:9603271231159800.
El-Toukhy SE, El-Daly SM, Kamel MM, Nabih HK. The diagnostic significance of circulating miRNAs and metabolite profiling in early prediction of breast cancer in Egyptian women. J Cancer Res Clin Oncol. 2023;149(8):5437–51.
Acknowledgements
We would like to acknowledge the hard and dedicated work of all the staff who implemented the intervention and evaluation components of the study.
Funding
No external funding received to conduct this study.
Author information
Authors and Affiliations
Contributions
Conception and design of the research: HL, XT. Acquisition of data: HL, XT. Analysis and interpretation of the data: HL. Statistical analysis: HL. Obtaining financing: None. Writing of the manuscript: HL. Critical revision of the manuscript for intellectual content: HL. All authors read and approved the final draft.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, H., Tie, Xj. Exploring research progress in studying serum exosomal miRNA-21 as a molecular diagnostic marker for breast cancer. Clin Transl Oncol (2024). https://doi.org/10.1007/s12094-024-03454-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12094-024-03454-z